Abstract
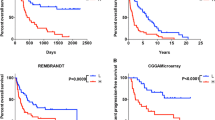
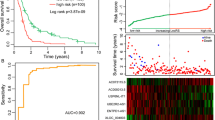
Glioma is the most common and fatal primary brain tumor in human. Long non-coding RNA (lncRNA), which are characterized by regulation of gene expression and chromatin recombination play an important role in glioma, and immunotherapy is a promising cancer treatment. Therefore, it is necessary to identify Immune-related lncRNAs in glioma. In this study,we collected and evaluated the RNA-seq data of The Cancer Genome Atlas (TCGA, https://www.ncbi.nlm.nih.gov/) and Chinese Glioma Genome Atlas (CGGA, https://www.cgga.org.cn/) glioma patients and immune-related lncRNAs were screened. Cox regression and LASSO analysis were performed to construct a risk score formula to explor the different overall survival between high- and low-risk groups in TCGA and verified with CGGA. Gene ontology (GO) and pathway-enrichment analysis (KEGG) were performed to identify the function of screened genes. Co-expression network were performed of these genes for further analysis. Eleven immune-related lncRNAs were concerned to be involved in survival and adopted to construct the risk score formula. Patients with high-risk score held poor survival both in TCGA and CGGA. Compared with current clinical data, the Area Under Curve (AUC) of different years and Principal components analysis (PCA) suggested that the formula had better predictive power. Functional Annotation of immune-related lncRNAs showed that the differences overall survival of high and low RS group might be caused by the cell differentiation, microtubule polymerization, etc. We successfully constructed an immune-related lncRNAs formula with powerful predictive function, which provides certain guidance value to the analysis of glioma pathogenesis and clinical treatment, and potential therapeutic targets for glioma treatment.







Similar content being viewed by others
References
Barsyte-Lovejoy D, Lau SK et al (2006) The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res 66(10):5330–5337
Bhan A, Soleimani M et al (2017) Long noncoding RNA and Cancer: a new paradigm. Cancer Res 77(15):3965–3981
Brandes AA, Tosoni A et al (2008) Glioblastoma in adults. Crit Rev Oncol Hematol 67(2):139–152
Cheng Z, Li Z et al (2017) Long Non-coding RNA XIST promotes glioma tumorigenicity and angiogenesis by acting as a molecular sponge of miR-429. J Cancer 8(19):4106–4116
Chernova OB, Hunyadi A et al (2001) A novel member of the WD-repeat gene family, WDR11, maps to the 10q26 region and is disrupted by a chromosome translocation in human glioblastoma cells. Oncogene 20(38):5378–5392
Davis ME (2018) Epidemiology and overview of gliomas. Semin Oncol Nurs 34(5):420–429
Fischer GM, Vashisht GY et al (2018) Metabolic strategies of melanoma cells: mechanisms, interactions with the tumor microenvironment, and therapeutic implications. Pigment Cell Melanoma Res 31(1):11–30
Giavina-Bianchi MH, Giavina-Bianchi PJ et al (2017) Melanoma: tumor microenvironment and new treatments. An Bras Dermatol 92(2):156–166
Gong X, Huang M (2017) Long non-coding RNA MEG3 promotes the proliferation of glioma cells through targeting Wnt/beta-catenin signal pathway. Cancer Gene Ther 24(9):381–385
Gousias K, Markou M et al (2009) Descriptive epidemiology of cerebral gliomas in northwest Greece and study of potential predisposing factors, 2005–2007. Neuroepidemiology 33(2):89–95
Kiang KM, Zhang XQ et al (2017) CRNDE expression positively correlates with EGFR activation and modulates glioma cell growth. Target Oncol 12(3):353–363
Larjavaara S, Mantyla R et al (2007) Incidence of gliomas by anatomic location. Neuro Oncol 9(3):319–325
Long GV, Atkinson V et al (2018) Combination nivolumab and ipilimumab or nivolumab alone in melanoma brain metastases: a multicentre randomised phase 2 study. Lancet Oncol 19(5):672–681
Matsui M, Corey DR (2017) Non-coding RNAs as drug targets. Nat Rev Drug Discov 16(3):167–179
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 15(Spec No 1):R17–29
McGranahan T, Therkelsen KE et al (2019) Current State of Immunotherapy for Treatment of Glioblastoma. Curr Treat Options Oncol 20(3):24
Ohgaki H, Kleihues P (2005) Epidemiology and etiology of gliomas. Acta Neuropathol 109(1):93–108
Omuro A, DeAngelis LM (2013) Glioblastoma and other malignant gliomas: a clinical review. JAMA 310(17):1842–1850
Ostrom QT, Bauchet L et al (2014) The epidemiology of glioma in adults: a "state of the science" review. Neuro Oncol 16(7):896–913
Shi Y, Wang Y et al (2014) Long non-coding RNA H19 promotes glioma cell invasion by deriving miR-675. PLoS ONE 9(1):e86295
Wan MT, Ming ME (2018) Nivolumab versus ipilimumab in the treatment of advanced melanoma: a critical appraisal: ORIGINAL ARTICLE: Wolchok JD, Chiarion-Sileni V, Gonzalez R et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med 2017; 377:1345-56. Br J Dermatol 179(2):296–300
Wirsching HG, Galanis E et al (2016) Glioblastoma. Handb Clin Neurol 134:381–397
Funding
No funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have read and approved to submit it to your journal. There are no conflicts of interest of any authors in relation to the submission.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xia, P., Li, Q., Wu, G. et al. An Immune-Related lncRNA Signature to Predict Survival In Glioma Patients. Cell Mol Neurobiol 41, 365–375 (2021). https://doi.org/10.1007/s10571-020-00857-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-020-00857-8