Abstract
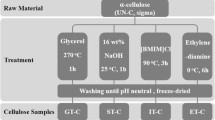
This study evaluates how bacterial cellulose (BC) properties influence the efficiency of enzymatic hydrolysis. BC was produced by the Medusomyces gisevii Sa-12 symbiotic culture in an enzymatic hydrolyzate obtained from oat hulls and was characterized by Fourier-transform infrared spectroscopy, scanning electron microscopy, thermogravimetric analysis, and X-ray diffraction. The enzymatic hydrolysis was examined with dried BCs unwashed and washed of culture medium components and cell debris, as well as with wet BC washed of culture medium components, at initial solid loadings of 10 and 30 g/L. The enzymatic hydrolysis of the BC sample unwashed of culture medium components and cell debris exhibited a substrate conversion degree of 56.3–66.6%. The conversion degree of the BC samples washed of culture medium components and cell debris was 89.4–99.5%. The removal of culture medium components and cell debris increased the conversion degree by 1.5 times. The drying of wet BC was found to decrease the enzymatic hydrolysis rate but it did not affect the conversion degree: the maximum yield of reducing sugars of 99.5% was achieved in 56 h for dried BC and in 16 h for wet BC. The substrate’s impurity content (growth medium components and cells) and moisture had the greatest effect on the performance of enzymatic hydrolysis of BC. The high BC crystallinity index of 90% was found to be not a determinant for the enzymatic hydrolysis efficiency.
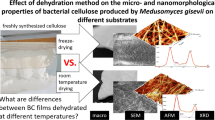
Graphical abstract







Similar content being viewed by others
References
Adepu S, Khandelwal M (2018) Broad-spectrum antimicrobial activity of bacterial cellulose silver nanocomposites with sustained release. J Mater Sci 53(3):1596–1906. https://doi.org/10.1007/s10853-017-1638-9
Ahvenainen P, Kontro I, Svedstrom K (2016) Comparison of sample crystallinity determination methods by X-ray diffraction for challenging cellulose I materials. Cellulose 23(2):1073–1086. https://doi.org/10.1007/s10570-016-0881-6
Akduman B, Uygun M, Çoban EP, Uygun DA, Bıyık H, Akgöl S (2013) Reversible immobilization of urease by using bacterial cellulose nanofibers. Appl Biochem Biotechnol 171(8):2285–2294. https://doi.org/10.1007/s12010-013-0541-3
Aleshina LA, Glazkova SV, Lugovskaya LA, Podoynikova MV, Fofanov AD, Silina EV (2001) A contemporary view of cellulose structure (review). Chem Plant Raw Mater 1:5–36 (rus)
Aleshina LA, Gladysheva EK, Budaeva VV, Skiba EA, Arkharova NA, Sakovich GV (2018) X-ray diffraction study of bacterial nanocellulose produced by the Medusomyces Gisevii Sa-12 culture in enzymatic hydrolysates of oat hulls. Crystallogr Rep 63(6):955–960. https://doi.org/10.1134/S1063774518050024
Amano Y, Nozaki K, Araki T, Shibasaki H, Kuga S, Kanda T (2002) Reactivities of cellulases from fungi towards ribbon-type bacterial cellulose and band-shaped bacterial cellulose. Cellulose 8(4):267–274. https://doi.org/10.1023/A:1015167304364
Anwar B, Bundjali B, Arcana IM (2015) Isolation of cellulose nanocrystals from bacterial cellulose produced from pineapple peel waste juice as culture medium. Proc Chem 16:279–284. https://doi.org/10.1016/j.proche.2015.12.051
Barth A (2007) Infrared spectroscopy of proteins: review. Biochim Biophys Acta 1767(9):1073–1101. https://doi.org/10.1016/j.bbabio.2007.06.004
Barud HS, Araújo Júnior AM, Santos DB, Assunção RMN, Meireles CS, Cerqueira DA, Filho GR, Ribeiro CA, Messaddeq Y, Ribeiro SJL (2008) Thermal behavior of cellulose acetate produced from homogeneous acetylation of bacterial cellulose. Thermochim Acta 471(1–2):61–69. https://doi.org/10.1016/j.tca.2008.02.009
Belgacem MN, Gandini A (2008) Monomers, polymers and composites from renewable resources. Elsevier, Amsterdam
Boisset C, Fraschini C, Schulein M, Henrissat B, Chanzy H (2000) Imaging the enzymatic digestion of bacterial cellulose ribbons reveals the endo character of the cellobiohydrolase Cel6A from Humicola insolens and its mode of synergy with cellobiohydrolase Cel7A. J Appl Environ Microbiol 66(4):1444–1452. https://doi.org/10.1128/AEM.66.4.1444-1452.2000%5d
Bowling AJ, Amano Y, Lindstrom R, Brown RM Jr. (2001) Rotation of cellulose ribbons during degradation with fungal cellulose. Cellulose 8(1):91–97. https://doi.org/10.1023/A:1016660621440
Budaeva VV, Makarova EI, Gismatulina Y (2016) Integrated flowsheet for conversion of nonwoody biomass into polyfunctional materials. Key Eng Mater 670:202–206. https://doi.org/10.4028/www.scientific.net/KEM.670.202
Chawla PR, Bajaj IB, Survase SA, Singhal RS (2009) Microbial cellulose: fermentative production and applications. Food Technol Biotechnol 47(2):107–124
Chen Y, Stipanovic AJ, Winter WT, Wilson DB, Kim YJ (2007) Effect of digestion by pure cellulases on crystallinity and average chain length for bacterial and microcrystalline celluloses. Cellulose 14:283–293. https://doi.org/10.1007/s10570-007-9115-2
Cheng K, Catchmark J, Demirci A (2009) Enhanced production of bacterial cellulose by using a biofilm reactor and its material property analysis. J Biol Eng 3:1–10. https://doi.org/10.1186/1754-1611-3-12
Czaja W, Krystynowicz A, Bielecki S, Brown RM Jr. (2006) Microbial cellulose: the natural power to heal wounds. Biomaterials 27(2):145–151. https://doi.org/10.1016/j.biomaterials.2005.07.035
Fontana JD, De Souza AM, Fontana CK, Torriani IL, Moreschi JC, Gallotti BJ, De Souza SJ, Narcisco GP, Bichara JA, Farah LFX (1990) Acetobacter cellulose pellicle as a temporary skin substitute. Appl Biochem Biotechnol 24(1):253–264. https://doi.org/10.1007/BF02920250
Foong CY, Hamzah MSA, Razak SIA, Saidin S, Nayan NHM (2018) Influence of poly(lactic acid) layer on the physical and antibacterial properties of dry bacterial cellulose sheet for potential acute wound healing materials. Fiber Polym 19(2):263–271. https://doi.org/10.1007/s12221-018-7850-7
French AD (2014) Idealized powder diffraction patterns for cellulose polymorphs. Cellulose 21(2):885–896. https://doi.org/10.1007/s10570-013-0030-4
French AD, Howley PS (1989) Comparisons of structures proposed for cellulose. In: Schuerch C (ed) Cellulose and wood—chemistry and technology. Wiley, New York, pp 159–167
Gladysheva EK, Skiba EA, Zolotukhin VN, Sakovich GV (2018) Study of the conditions for the biosynthesis of bacterial cellulose by the producer Medusomyces gisevii Sa-12. Appl Biochem Microbiol 54(2):179–187. https://doi.org/10.1134/S0003683818020035
Goh WN, Rosma A, Kaur B, Fazilah A, Karim AA, Rajeev B (2012a) Fermentation of black tea broth (Kombucha): I. Effects of sucrose concentration and fermentation time on the yield of microbial cellulose. Int Food Res J 19(1):109–117
Goh WN, Rosma A, Kaur B, Fazilah A, Karim AA, Rajeev B (2012b) Microstructure and physical properties of microbial cellulose produced during fermentation of black tea broth (Kombucha). II. Int Food Res J 19(1):153–158
Hall M, Bansal P, Lee JH, Realff MJ, Bommarius AS (2010) Cellulose crystallinity: a key predictor of the enzymatic hydrolysis rate. FEBS J 277(6):1571–1582. https://doi.org/10.1111/j.1742-4658.2010.07585.x
Horii F, Hirai A, Kitamura R (1987) CP/MAS 13C NMR spectra of the crystalline components of native celluloses. Macromolecules 20(9):2117–2120. https://doi.org/10.1021/ma00175a012
Hu F, Ragauskas A (2012) Pretreatment and lignocellulosic chemistry. Bioenerg Res 5(4):1043–1066. https://doi.org/10.1007/s12155-012-9208-0
Ioelovich M (2014) Study of enzymatic hydrolysis of bacterial nanocellulose. Am J BioSci 2(6–1):13–16. https://doi.org/10.11648/j.ajbio.s.2014020601.13
Ioelovich M, Morag E (2011) Effect of cellulose structure on enzymatic hydrolysis. BioResources 6(3):2818–2835. https://doi.org/10.15376/biores.6.3.2818_2835
Jung H, Yoon HG, Park W, Choi C, Wilson DB, Shin DH, Kim YJ (2008) Effect of sodium hydroxide treatment of bacterial cellulose on cellulase activity. Cellulose 15(3):465–471. https://doi.org/10.1007/s10570-007-9190-4
Keshk SM (2014) Bacterial cellulose production and its industrial applications. J Bioprocess Biotechnol 4(2):150–160. https://doi.org/10.4172/2155-9821.1000150
Khandelwal M, Windle AH, Hessler N (2016) In situ tunability of bacteria produced cellulose by additives in the culture media. J Mater Sci 51(10):4839–4844. https://doi.org/10.1007/s10853-016-9783-0
Lee SE, Park YS (2017) The role of bacterial cellulose in artificial blood vessels. Mol Cell Toxicol 13(3):257–261. https://doi.org/10.1007/s13273-017-0028-3
Lv P, Zhou H, Zhao M, Li D, Lu K, Wang D, Huang J, Cai Y, Lucia LA (2018) Highly flexible, transparent, and conductive silver nanowire-attached bacterial cellulose conductors. Cellulose 25(6):3189–3196. https://doi.org/10.1007/s10570-018-1773-8
Meftahi A, Khajavi R, Rashidi A, Sattari M, Yazdanshenas ME, Torabi M (2010) The effects of cotton gauze coating with microbial cellulose. Cellulose 17(1):199–204. https://doi.org/10.1007/s10570-009-9377-y
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428. https://doi.org/10.1021/ac60147a030
Nishiyama Y, Langan P, Chanzy H (2002) Crystal structure and hydrogen-bonding system in cellulose Iβ from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 124(31):9074–9082. https://doi.org/10.1021/ja0257319
Nishiyama Y, Chanzy H, Langan P (2003) Crystal structure and hydrogen bonding system in cellulose Iα from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 125(47):14300–14306. https://doi.org/10.1021/ja037055w
Santos SM, Carbajo JM, Gómez N, Ladero M, Villar JC (2017) Paper reinforcing by in situ growth of bacterial cellulose. J Mater Sci 52(10):5882–5893. https://doi.org/10.1007/s10853-017-0824-0
Schumann AD, Wippermann J, Klemm OD, Kramer F, Koth D, Kosmehl H, Wahlers T, Salehi-Gelani S (2009) Artificial vascular implants from bacterial cellulose: preliminary results of small arterial substitutes. Cellulose 16(5):877–885. https://doi.org/10.1007/s10570-008-9264-y
Silva TM, Santiago PO, Purcena LLA, Fernandes KF (2010) Study of the cashew gum polysaccharide for the horseradish peroxidase immobilization: structural characteristics, stability and recovery. Mater Sci Eng 30(4):526–530. https://doi.org/10.1016/j.msec.2010.01.016
Terinte N, Ibbett R, Schuster KC (2011) Overview on native cellulose and microcrystalline cellulose I structure studied by X-ray diffraction (WAXD): comparison between measurement techniques. Lenzing Ber 89:118–131
Tiboni M, Grzybowski A, Passos M, Barison A, Lião LM, Campos FR, Pontarolo R, Fontana JD (2012) The use of dyed bacterial cellulose to monitor cellulose complex activity. Cellulose 19(6):1867–1877. https://doi.org/10.1007/s10570-012-9787-0
Torlopov MA, Mikhaylov VI, Udoratina EV, Aleshina LA, Prusskii AI, Tsvetkov NV, Krivoshapkin PV (2018) Cellulose nanocrystals with different length-to-diameter ratios extracted from various plants using novel system acetic acid/phosphotungstic acid/octanol-1. Cellulose 25(2):1031–1046. https://doi.org/10.1007/s10570-017-1624-z
Vazquez A, Foresti ML, Cerrutti P, Galvagno M (2013) Bacterial cellulose from simple and low cost production media by Gluconacetobacter xylinus. J Polym Environ 21(2):545–554. https://doi.org/10.1007/s10924-012-0541-3
Wang Y, Wang Y, Jiang L (2018) Freestanding carbon aerogels produced from bacterial cellulose and its Ni/MnO2/Ni(OH)2 decoration for supercapacitor electrodes. J Appl Electrochem 48(5):495–507. https://doi.org/10.1007/s10800-018-1183-5
Yin X, Zhang X, Yang J, Lin Q, Wang J, Zhu Q (2011) Comparison of succinylation methods for bacterial cellulose and adsorption capacities of bacterial cellulose derivatives for Cu2+ ion. Polym Bull 67(3):401–412. https://doi.org/10.1007/s00289-010-0388-5
Żywicka A, Peitler D, Rakoczy R, Junka AF, Fijałkowski K (2016) Wet and dry forms of bacterial cellulose synthetized by different strains of Gluconacetobacter xylinus as carriers for yeast immobilization. Appl Biochem Biotechnol 180(4):805–816. https://doi.org/10.1007/s12010-016-2134-4
Funding
This research was supported by the Russian Science Foundation (Project # 17-19-01054).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kashcheyeva, E.I., Gladysheva, E.K., Skiba, E.A. et al. A study of properties and enzymatic hydrolysis of bacterial cellulose. Cellulose 26, 2255–2265 (2019). https://doi.org/10.1007/s10570-018-02242-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-018-02242-7