Abstract
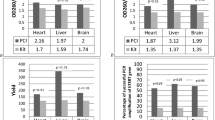
Many factors affect the integrity of messenger RNA from human autopsy tissues including postmortem interval (PMI) between death and tissue preservation and the pre-mortem agonal and disease states. In this communication, we describe RNA isolation and characterization of 389 samples from 18 different tissues from elderly donors who were participants in a rapid whole-body autopsy program located in Sun City, Arizona (www.brainandbodydonationprogram.org). Most tissues were collected within a PMI of 2–6 h (median 3.15 h; N = 455), but for this study, tissue from cases with longer PMIs (1.25–29.25 h) were included. RNA quality was assessed by RNA integrity number (RIN) and total yield (ng RNA/mg tissue). RIN correlated with PMI for heart (r = −0.531, p = 0.009) and liver (r = −558, p = 0.0017), while RNA yield correlated with PMI for colon (r = −485, p = 0.016) and skin (r = −0.460, p = 0.031). RNAs with the lowest integrity were from skin and cervix where 22.7 and 31.4 % of samples respectively failed to produce intact RNA; by contrast all samples from esophagus, lymph node, jejunum, lung, stomach, submandibular gland and kidney produced RNA with measurable RINs. Expression levels in heart RNA of 4 common housekeeping normalization genes showed significant correlations of Ct values with RIN, but only one gene, glyceraldehyde-3 phosphate dehydrogenase, showed a correlation of Ct with PMI. There were no correlations between RIN values obtained for liver, adrenal, cervix, esophagus and lymph node and those obtained from corresponding brain samples. We show that high quality RNA can be produced from most human autopsy tissues, though with significant differences between tissues and donors. The RNA stability and yield did not depend solely on PMI; other undetermined factors are involved, but these do not include the age of the donor.



Similar content being viewed by others
References
Abasolo N, Torrell H, Roig B, Moyano S, Vilella E, Martorell L (2011) RT-qPCR study on post-mortem brain samples from patients with major psychiatric disorders: reference genes and specimen characteristics. J Psychiatr Res 45:1411–1418
Ahmed S, Shaffer A, Geddes T, Studzinski D, Mitton K, Pruetz B, Long G, Shanley C (2015) Evaluation of optimal RNA extraction method from human carotid atherosclerotic plaque. Cardiovasc Pathol 24:187–190
Andreasson A, Kiss NB, Juhlin CC, Hoog A (2013) Long-term storage of endocrine tissues at −80 °C does not adversely affect RNA quality or overall histomorphology. Biopreserv Biobank 11:366–370
Atz M, Walsh D, Cartagena P, Li J, Evans S, Choudary P, Overman K, Stein R, Tomita H, Potkin S, Myers R, Watson SJ, Jones EG, Akil H, Bunney WE Jr, Vawter MP (2007) Methodological considerations for gene expression profiling of human brain. J Neurosci Methods 163:295–309
Auer H, Mobley JA, Ayers LW, Bowen J, Chuaqui RF, Johnson LA, Livolsi VA, Lubensky IA, McGarvey D, Monovich LC, Moskaluk CA, Rumpel CA, Sexton KC, Washington MK, Wiles KR, Grizzle WE, Ramirez NC (2014) The effects of frozen tissue storage conditions on the integrity of RNA and protein. Biotech Histochem 89:518–528
Azevedo-Pouly AC, Elgamal OA, Schmittgen TD (2014) RNA isolation from mouse pancreas: a ribonuclease-rich tissue. J Vis Exp 90:e51779
Barber RD, Harmer DW, Coleman RA, Clark BJ (2005) GAPDH as a housekeeping gene: analysis of GAPDH mRNA expression in a panel of 72 human tissues. Physiol Genomics 21:389–395
Beach TG (2013) Alzheimer’s disease and the “Valley Of Death”: not enough guidance from human brain tissue? J Alzheimers Dis 33(Suppl 1):S219–S233. doi:10.3233/JAD-2012-129020.:S219-S233
Beach TG, Sue LI, Walker DG, Roher AE, Lue L, Vedders L, Connor DJ, Sabbagh MN, Rogers J (2008) The sun health research institute brain donation program: description and experience, 1987–2007. Cell Tissue Bank 9:229–245
Beach TG, Adler CH, Sue LI, Serrano G, Shill HA, Walker DG, Lue L, Roher AE, Dugger BN, Maarouf C, Birdsill AC, Intorcia A, Saxon-Labelle M, Pullen J, Scroggins A, Filon J, Scott S, Hoffman B, Garcia A, Caviness JN, Hentz JG, Driver-Dunckley E, Jacobson SA, Davis KJ, Belden CM, Long KE, Malek-Ahmadi M, Powell JJ, Gale LD, Nicholson LR, Caselli RJ, Woodruff BK, Rapscak SZ, Ahern GL, Shi J, Burke AD, Reiman EM, Sabbagh MN (2015) Arizona study of aging and neurodegenerative disorders and brain and body donation program. Neuropathology 35(4):354–389
Birdsill AC, Walker DG, Lue L, Sue LI, Beach TG (2011) Postmortem interval effect on RNA and gene expression in human brain tissue. Cell Tissue Bank 12:311–318
Brisco MJ, Morley AA (2012) Quantification of RNA integrity and its use for measurement of transcript number. Nucleic Acids Res 40:e144
Broniscer A, Baker JN, Baker SJ, Chi SN, Geyer JR, Morris EB, Gajjar A (2010) Prospective collection of tissue samples at autopsy in children with diffuse intrinsic pontine glioma. Cancer 116:4632–4637
Buesa C, Maes T, Subirada F, Barrachina M, Ferrer I (2004) DNA chip technology in brain banks: confronting a degrading world. J Neuropathol Exp Neurol 63:1003–1014
Burke WJ, O’Malley KL, Chung HD, Harmon SK, Miller JP, Berg L (1991) Effect of pre- and postmortem variables on specific mRNA levels in human brain. Brain Res Mol Brain Res 11:37–41
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Condelli V, Lettini G, Patitucci G, D’Auria F, D’Amico M, Vita G, Musto P, Cuomo C, Landriscina M (2014) Validation of vacuum-based refrigerated system for biobanking tissue preservation: analysis of cellular morphology, protein stability, and RNA quality. Biopreserv Biobank 12:35–45
Cummings TJ, Strum JC, Yoon LW, Szymanski MH, Hulette CM (2001) Recovery and expression of messenger RNA from postmortem human brain tissue. Mod Pathol 14:1157–1161
Dawany NB, Dampier WN, Tozeren A (2011) Large-scale integration of microarray data reveals genes and pathways common to multiple cancer types. Int J Cancer 128:2881–2891
Dheda K, Huggett JF, Bustin SA, Johnson MA, Rook G, Zumla A (2004) Validation of housekeeping genes for normalizing RNA expression in real-time PCR. Biotechniques 37:112–119
Ervin JF, Heinzen EL, Cronin KD, Goldstein D, Szymanski MH, Burke JR, Welsh-Bohmer KA, Hulette CM (2007) Postmortem delay has minimal effect on brain RNA integrity. J Neuropathol Exp Neurol 66:1093–1099
Fajardy I, Moitrot E, Vambergue A, Vandersippe-Millot M, Deruelle P, Rousseaux J (2009) Time course analysis of RNA stability in human placenta. BMC Mol Biol 10:21. doi:10.1186/1471-2199-10-21.:21-10
Fasold M, Binder H (2012) Estimating RNA-quality using GeneChip microarrays. BMC Genom 13:186. doi:10.1186/1471-2164-13-186.:186-13
Ferrer I, Santpere G, Arzberger T, Bell J, Blanco R, Boluda S, Budka H, Carmona M, Giaccone G, Krebs B, Limido L, Parchi P, Puig B, Strammiello R, Ströbel T, Kretzschmar H (2007) Brain protein preservation largely depends on the postmortem storage temperature: implications for study of proteins in human neurologic diseases and management of brain banks: a BrainNet Europe Study. J Neuropathol Exp Neurol 66:35–46
Gonzalez-Herrera L, Valenzuela A, Marchal JA, Lorente JA, Villanueva E (2013) Studies on RNA integrity and gene expression in human myocardial tissue, pericardial fluid and blood, and its postmortem stability. Forensic Sci Int 232:218–228
Grabmuller M, Madea B, Courts C (2015) Comparative evaluation of different extraction and quantification methods for forensic RNA analysis. Forensic Sci Int Genet 16C:195–202. doi:10.1016/j.fsigen.2015.01.006.:195-202
Griffin M, Abu-El-Haija M, Abu-El-Haija M, Rokhlina T, Uc A (2012) Simplified and versatile method for isolation of high-quality RNA from pancreas. Biotechniques 52:332–334
Hatzis C, Sun H, Yao H, Hubbard RE, Meric-Bernstam F, Babiera GV, Wu Y, Pusztai L, Symmans WF (2011) Effects of tissue handling on RNA integrity and microarray measurements from resected breast cancers. J Natl Cancer Inst 103:1871–1883
Hedegaard J, Thorsen K, Lund MK, Hein AM, Hamilton-Dutoit SJ, Vang S, Nordentoft I, Birkenkamp-Demtroder K, Kruhoffer M, Hager H, Knudsen B, Andersen CL, Sorensen KD, Pedersen JS, Orntoft TF, Dyrskjot L (2014) Next-generation sequencing of RNA and DNA isolated from paired fresh-frozen and formalin-fixed paraffin-embedded samples of human cancer and normal tissue. PLoS One 9:e98187
Heinrich M, Matt K, Lutz-Bonengel S, Schmidt U (2007) Successful RNA extraction from various human postmortem tissues. Int J Legal Med 121:136–142
Hellemans J, Vandesompele J (2014) Selection of reliable reference genes for RT-qPCR analysis. Methods Mol Biol 1160:19–26. doi:10.1007/978-1-4939-0733-5_3.:19-26
Huggett J, Dheda K, Bustin S, Zumla A (2005) Real-time RT-PCR normalisation; strategies and considerations. Genes Immun 6:279–284
Kampf C, Mardinoglu A, Fagerberg L, Hallstrom BM, Edlund K, Lundberg E, Ponten F, Nielsen J, Uhlen M (2014) The human liver-specific proteome defined by transcriptomics and antibody-based profiling. FASEB J 28:2901–2914
Kap M, Oomen M, Arshad S, de JB, Riegman P (2014) Fit for purpose frozen tissue collections by RNA integrity number-based quality control assurance at the Erasmus MC tissue bank. Biopreserv Biobank 12:81–90
Kiewe P, Gueller S, Komor M, Stroux A, Thiel E, Hofmann WK (2009) Prediction of qualitative outcome of oligonucleotide microarray hybridization by measurement of RNA integrity using the 2100 Bioanalyzer capillary electrophoresis system. Ann Hematol 88:1177–1183
Kim S, Webster MJ (2009) Postmortem brain tissue for drug discovery in psychiatric research. Schizophr Bull 35:1031–1033
Koppelkamm A, Vennemann B, Lutz-Bonengel S, Fracasso T, Vennemann M (2011) RNA integrity in post-mortem samples: influencing parameters and implications on RT-qPCR assays. Int J Legal Med 125:573–580
Li JZ, Vawter MP, Walsh DM, Tomita H, Evans SJ, Choudary PV, Lopez JF, Avelar A, Shokoohi V, Chung T, Mesarwi O, Jones EG, Watson SJ, Akil H, Bunney WE Jr, Myers RM (2004) Systematic changes in gene expression in postmortem human brains associated with tissue pH and terminal medical conditions. Hum Mol Genet 13:609–616
Lukiw WJ, Bazan NG (1997) Cyclooxygenase 2 RNA message abundance, stability, and hypervariability in sporadic Alzheimer neocortex. J Neurosci Res 50:937–945
Micke P, Ohshima M, Tahmasebpoor S, Ren ZP, Ostman A, Ponten F, Botling J (2006) Biobanking of fresh frozen tissue: RNA is stable in nonfixed surgical specimens. Lab Invest 86:202–211
Mora M, Angelini C, Bignami F, Bodin AM, Crimi M, Di Donato JH, Felice A, Jaeger C, Karcagi V, LeCam Y, Lynn S, Meznaric M, Moggio M, Monaco L, Politano L, de la Paz MP, Saker S, Schneiderat P, Ensini M, Garavaglia B, Gurwitz D, Johnson D, Muntoni F, Puymirat J, Reza M, Voit T, Baldo C, Bricarelli FD, Goldwurm S, Merla G, Pegoraro E, Renieri A, Zatloukal K, Filocamo M, Lochmuller H (2014) The EuroBioBank Network: 10 years of hands-on experience of collaborative, transnational biobanking for rare diseases. Eur J Hum Genet 23:1116–1123
Nagy C, Maheu M, Lopez JP, Vaillancourt K, Cruceanu C, Gross JA, Arnovitz M, Mechawar N, Turecki G (2015) Effects of postmortem interval on biomolecule integrity in the brain. J Neuropathol Exp Neurol 74:459–469
Opitz L, Salinas-Riester G, Grade M, Jung K, Jo P, Emons G, Ghadimi BM, Beissbarth T, Gaedcke J (2010) Impact of RNA degradation on gene expression profiling. BMC Med Genomics 3:36. doi:10.1186/1755-8794-3-36.:36-3
Popova T, Mennerich D, Weith A, Quast K (2008) Effect of RNA quality on transcript intensity levels in microarray analysis of human post-mortem brain tissues. BMC Genom 9:91. doi:10.1186/1471-2164-9-91.:91-99
Preece P, Cairns NJ (2003) Quantifying mRNA in postmortem human brain: influence of gender, age at death, postmortem interval, brain pH, agonal state and inter-lobe mRNA variance. Brain Res Mol Brain Res 118:60–71
Ravid R, Ferrer I (2012) Brain banks as key part of biochemical and molecular studies on cerebral cortex involvement in Parkinson’s disease. FEBS J 279:1167–1176
Rudloff U, Bhanot U, Gerald W, Klimstra DS, Jarnagin WR, Brennan MF, Allen PJ (2010) Biobanking of human pancreas cancer tissue: impact of ex vivo procurement times on RNA quality. Ann Surg Oncol 17:2229–2236
Ruggeri BA, Camp F, Miknyoczki S (2014) Animal models of disease: pre-clinical animal models of cancer and their applications and utility in drug discovery. Biochem Pharmacol 87:150–161
Schroeder A, Mueller O, Stocker S, Salowsky R, Leiber M, Gassmann M, Lightfoot S, Menzel W, Granzow M, Ragg T (2006) The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol 7:3
Shabihkhani M, Lucey GM, Wei B, Mareninov S, Lou JJ, Vinters HV, Singer EJ, Cloughesy TF, Yong WH (2014) The procurement, storage, and quality assurance of frozen blood and tissue biospecimens in pathology, biorepository, and biobank settings. Clin Biochem 47:258–266
Stan AD, Ghose S, Gao XM, Roberts RC, Lewis-Amezcua K, Hatanpaa KJ, Tamminga CA (2006) Human postmortem tissue: what quality markers matter? Brain Res 1123:1–11
Sun H, Sun R, Hao M, Wang Y, Zhang X, Liu Y, Cong X (2016) Effect of duration of ex vivo ischemia time and storage period on RNA quality in biobanked human renal cell carcinoma tissue. Ann Surg Oncol 23:297–304
Tarvin KA, Sandusky GE (2014) Using molecular profiled human tissue to accelerate drug discovery. Expert Opin Drug Discov 9:1383–1387
Tomita H, Vawter MP, Walsh DM, Evans SJ, Choudary PV, Li J, Overman KM, Atz ME, Myers RM, Jones EG, Watson SJ, Akil H, Bunney WE Jr (2004) Effect of agonal and postmortem factors on gene expression profile: quality control in microarray analyses of postmortem human brain. Biol Psychiatry 55:346–352
Vandesompele J, De PK, Pattyn F, Poppe B, Van RN, De PA, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:RESEARCH0034
Vermeulen J, De PK, Lefever S, Nuytens J, De VF, Derveaux S, Hellemans J, Speleman F, Vandesompele J (2011) Measurable impact of RNA quality on gene expression results from quantitative PCR. Nucleic Acids Res 39:e63
Acknowledgments
The Brain and Body Donation Program is supported by the National Institute of Neurological Disorders and Stroke (U24 NS072026 National Brain and Tissue Resource for Parkinson’s Disease and Related Disorders), the National Institute on Aging (P30 AG19610 Arizona Alzheimer’s Disease Core Center), the Arizona Department of Health Services (contract 211002, Arizona Alzheimer’s Research Center), the Arizona Biomedical Research Commission (contracts 4001, 0011, 05-901 and 1001 to the Arizona Parkinson’s Disease Consortium) and the Michael J. Fox Foundation for Parkinson’s Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Walker, D.G., Whetzel, A.M., Serrano, G. et al. Characterization of RNA isolated from eighteen different human tissues: results from a rapid human autopsy program. Cell Tissue Bank 17, 361–375 (2016). https://doi.org/10.1007/s10561-016-9555-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10561-016-9555-8