Abstract
Purpose
We investigated the occurrence and the prognostic and predictive relationship of a selected number of somatic mutations in triple-negative breast cancer (TNBC) patients having known clinical outcomes treated within the IBCSG Trial 22-00.
Methods
A matched case–control sampling selected patients enrolled in the IBCSG Trial 22-00 who had TNBC tumors, based on local assessment. Cases had invasive breast cancer recurrence (at local, regional, or distant site) according to the protocol definition. Matched controls had not recurred. Mutational analysis was performed with OncoCarta panel v1.0 using Mass Array System. The panel includes 19 genes belonging to different functional pathways as PI3K pathway, receptor tyrosine kinase, and cell cycle-metabolic group. Conditional logistic regression assessed the association of mutation status with breast cancer recurrence.
Results
Mutation assessment was successful for 135 patients (49 cases, 86 controls). A total of 37 (27.4%) of the 135 patients had at least one mutation in the selected genes. PIK3CA was the most common mutated gene (18/135; 13.3%), followed by BRAF, KIT and PDGFRA (each 4/135, 3.0%) and AKT1 (3/135; 2.2%). TNBC patients with at least one mutation had increased odds of recurrence compared with those with wild-type tumors (odds ratio (OR) 2.28; 95% CI 0.88–5.92), though this difference was not statistically significant (p = 0.09). We found no evidence that these mutations were predictive for the value of maintenance metronomic chemotherapy.
Conclusions
Mutations in the tested oncogenes were not associated with breast cancer recurrence in this TNBC subset of patients. The question of whether any of these mutated genes (e.g., PIK3CA) may represent a useful therapeutic target in TNBC may be answered by ongoing clinical trials and/or larger dataset analysis.
Similar content being viewed by others
Introduction
Recent studies of gene expression profiling led to the identification of breast cancer subtypes based on common molecular features [1, 2]. Triple-negative breast cancers (TNBC) do not express estrogen and progesterone receptors (ER/PgR) and HER2. They show typical morphological features and have been classified into intrinsic basal-like or claudin-low subtypes [3, 4].
TNBC represents the group of tumors with a worse prognosis. They are typically high grade, although low-grade tumors can rarely occur and these may have a better prognosis [5, 6]. Therefore, TNBC can be considered a heterogeneous entity including different subtypes [7] and several attempts have been made to establish a molecular classification of TNBC based on the results of both transcriptomic and genomic studies [8,9,10]. Recent independent studies have demonstrated that TNBC can be sub-classified either into six subtypes (basal-like I, basal-like II, mesenchymal, mesenchymal stem-like, immunomodulatory, and luminal androgen receptor) [9] or four subtypes (luminal androgen receptor, mesenchymal, basal-like immune-suppressed, and basal-like immune-activated) [10]. Following the integrative analysis of gene expression and genome-wide copy number alterations (CNAs) by the METABRIC consortium [11], TNBC were characterized by complex patterns of copy number gains and losses throughout the genome.
Genomic profiles of TNBC have recently been defined using whole exome sequencing. TNBC has a predominance of TP53 alterations that can be present in up to 80% of the cases, and a large set of mutated genes occurring with minor mutation frequencies [11,12,13]. TNBC shows mutations in PIK3CA (7%) [12], mutation or loss of phosphatase and tensin homolog (PTEN) (35%), and loss of inositol polyphosphate-4-phosphatase type II (INPP4B) (30%) [14]. Although several potentially actionable molecular alterations (e.g., PI3K/mTOR or RAS/RAF/MEK) have been found in TNBC, none have been confirmed as a ‘driver alteration.’
MALDI (Matrix-Assisted Laser Desorption/ionization)—TOF (Time of Flight) mass spectrometry is a versatile tool for high-performance DNA analysis and was selected to perform mutation analysis [15]. Pre-designed assays are commercially available for comprehensive mutation screening of solid tumors; in particular the OncoCarta Panel v1.0 used in the present study is able to screen more than 230 potentially druggable mutations across 19 common oncogenes (ABL1, AKT1, AKT2, BRAF, CDK4, EGFR, ERBB2, FGFR1, FGFR3, FLT3, JAK2, KIT, MET, HRAS, KRAS, NRAS, PDGFRA, PIK3CA, RET). These genes belong to different functional pathways including PI3K pathway, receptor tyrosine kinase, and cell cycle-metabolic group. Moreover, FGFR1 and FGFR3 codify for mitogenic signaling molecules that play a role in angiogenesis and cell migration and the mutational status of these genes may offer information about the response to antiangiogenic therapy.
The aim of this study was to explore the prognostic and predictive relationship between the presence of a selected set of somatic mutations in TNBC and the clinical outcome within the International Breast Cancer Study Group (IBCSG) Trial 22-00, to obtain a comprehensive description of the features of this important subset of tumors.
Patients and methods
IBCSG trial 22-00 population
IBCSG Trial 22-00 is a multicenter, randomized, adjuvant phase III clinical trial assessing the efficacy of low-dose metronomic CM maintenance chemotherapy (CM: cyclophosphamide 50 mg/day orally continuously for 1 year and methotrexate 2.5 mg/twice a day orally, days 1 and 2 of every week for 1 year) to no maintenance chemotherapy (No-CM), following breast cancer surgery and standard adjuvant chemotherapy [16]. The trial enrolled 1086 patients with estrogen (ER) and progesterone (PgR) receptor-negative tumors, and any HER2 and nodal status. The IBCSG Ethics Committee and ethics committees at each center approved the study, and all patients provided written informed consent. The trial is registered at clinicaltrials.gov (NCT00022516).
Assay/genotype methods
From a representative formalin-fixed and paraffin-embedded tissues (FFPE) block, 5-micron-thick histological sections were obtained for DNA extraction (QIAamp DNA FFPE Tissue Kit Qiagen, Hilden, D). Mutational analysis was performed with OncoCarta Panle v1.0 using the MassARRAY System based on MALDI-TOF mass spectrometry (Sequenom, San Diego, CA, USA), according to the manufacturer’s protocols. Genotyping by this method relies on the principle that mutant and wild-type alleles for a given point mutation produce single-allele base extension reaction products of different mass resolved by mass spectrometry. The OncoCarta Panel v1.0 includes Amplification and extension primers for the analysis of 230 mutations in 19 genes (ABL1, AKT1, AKT2, BRAF, CDK4, EGFR, ERBB2, FGFR1, FGFR3, FLT3, JAK2, KIT, MET, HRAS, KRAS, NRAS, PDGFRA, PIK3CA, RET). Assays were conducted without knowledge of clinical outcome.
Sampling design
A matched case–control sampling selected from 570 patients enrolled in IBCSG Trial 22-00 who were known to have TNBC according to the local assessment of ER, PgR, and HER2. The cases were defined as having an invasive local, regional, or distant recurrence of breast cancer. Controls without breast cancer recurrence were matched 2:1 using the following criteria: (i) nodal status (N0, 1–3 N+, 4–9 N+, 10 N+); (ii) tumor size (≤ 2 cm, > 2 cm); (iii) age (< 40, 40–55, 56–69, ≥ 70); (iv) menopausal status (premenopausal; postmenopausal); (v) year of randomization, by using a publically available macro (%gmatch; http://www.mayo.edu/research/departments-divisions/department-health-sciences-research/division-biomedical-statistics-informatics/software/locally-written-sas-macros). Of the selected 191 patients (67 cases, 124 controls), 135 had mutational assessment (49 cases and 86 controls), after excluding 56 patients with unavailable FFPE tumor blocks or insufficient material for assessment (Fig. 1). Patients with mutational assessment consented to use of their tumor tissue for research purposes, and the project was approved by the IBCSG biological protocols working group.
Statistical analysis
Conditional logistic regression was used to assess the association of breast cancer recurrence (case) with mutation status (at least one mutation vs. wild-type (WT)); the strata variable was the matching indicator for the case and controls. Treatment assignment (CM maintenance vs. No-CM)-by-mutation status interaction was included in the model to first assess the potentially differential treatment effect (i.e., predictive effect). If there was no evidence of interaction, the model without interaction term was used to assess the prognostic mutation effect on clinical outcome. Similarly, an exploratory analysis was also performed to assess the association of recurrence with PIK3CA pathway mutation status (at least one mutation from PIK3CA, AKT1 and AKT2: yes vs. no).
Because the statistical analysis of this type of case–control study was based on a conditional logistic regression, the only patients from the assessable population contributing to the analysis were those for which there was at least one control matched to the case. The analysis included 109 patients, comprising 44 cases and 65 controls (23 cases had one matched control; 21 cases 2 matched controls; 5 cases did not have any matched controls). Thus, 65 case–control pairs were informative strata in the conditional logistic regression analysis (Fig. 1). For a matched case–control design with 49 cases and 2 controls per case, if the frequency of at least one mutation was 22% in the control group and the correlation of mutation between matched pairs was 0.2, the study had 80% power to detect an odds ratio of 3.2 (two-sided α = 0.05) [17].
Results
The characteristics of the 570-patient cohort with locally assessed TNBC are summarized in Table 1, separately for the 135 assessable patients with mutation status and the 435 patients without mutation status assessed. The characteristics were mostly consistent, although assessable patients more frequently were node-positive as compared to those not assessed (57.0% vs. 38.4%, respectively) and more frequently had tumor size > 2 cm (63.0% vs. 52.4%, respectively).
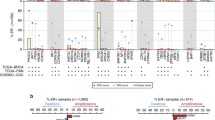
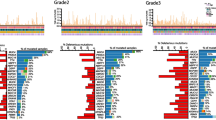
Among the 135 assessable patients, 37 (27.4%) patients had at least one mutation, affecting 12 of the 19 targeted genes, and 98 (72.6%) patients had wild-type status. Five patients had more than one mutation (Table 2). Among the specific mutations, PIK3CA was the most common (18/135; 13.3%), two of which co-occurred either with mutation in BRAF or with mutation in both AKT2 and PDGFRA. Mutations in BRAF, KIT, and PDGFRA each occurred in 4 patients (3.0%), while mutations in AKT1 occurred in 3 patients. When looking at pathway mutations (specifically: PIK3CA + AKT1 + AKT2), 22 (16%) patients had at least one of the PIK3CA, AKT1, or AKT2 mutations. The remaining mutations were detected in a single patient (0.7%) in each of 3 genes (ABL1, FGFR3, FLT3), or detected in 2 patients (1.5%) in each of 4 genes (AKT2, EGFR, ERBB2, MET). The occurrence of each mutation is shown in Fig. 2.
Using conditional logistic regression, a subset of 65 matched pairs (109 patients) formed the informative strata and contributed to the analysis of assessing the association of mutational status with breast cancer recurrence (Table 3). Patients with at least one mutation in the 19 targeted genes had increased odds of recurrence versus those with wild-type status (odds ratio (OR) 2.28; 95% CI 0.88–5.92; p = 0.09). Similar results were observed when assessing the association of at least one of PIK3CA, AKT1, and AKT2 mutation with the recurrence (ORR 2.60; 95% CI 0.62–6.41). There was no evidence of a predictive relevance of the mutation status with use or non-use of CM maintenance therapy (mutation-by-treatment interaction p = 0.73 or 0.48, respectively). For patients assigned to CM maintenance, having a targeted mutation was associated with increased odds of breast cancer recurrence (OR 1.98; 95% CI 0.57–6.92), as it was for patients assigned to No-CM (OR 2.62; 95% CI 0.76–9.07).
Discussion
The mutational analysis performed with OncoCarta panel v1.0 in 135 patients with locally assessed TNBC within trial IBCSG 22-00 identified 27.4% patients with at least one mutation in the 19 selected genes. Among patients with a breast cancer recurrence (cases), 36.7% had at least one mutation identified. Since this is a matched case–control study, the patients selected for the analysis were guaranteed to have higher-risk features (and be enrolled earlier in the trial) than the other patients, as by definition this design enriched for patients who had a recurrence (cases) and patients with matched disease characteristics.
In similar studies, the presence of a potentially targetable mutation was found in about 36% of cases with TNBC [12], although a different method for identifying mutations was used (high-throughput RNA sequencing—RNA-seq). About 40% of breast cancers contained single rare mutations that could be potential therapeutic targets [18]. Key driver mutations are thought to provide a selective advantage to a number of cells, facilitating their clonal expansion. A key finding from recent next-generation sequencing (NGS) studies was that each tumor contains a dominant clone (> 50% of cancer cells) that has a mutational profile different from the other sub-clones [19].
In our population of patients with TNBC, PIK3CA mutation was the most frequent mutation, being identified in 13.3% of selected patients’ tumors. These results were comparable to those reported in the literature showing PIK3CA mutations in 10–25% of breast cancers [12] (http://cancer.sanger.ac.uk/cancergenome/projects/cosmic/). Other groups reported that this mutation could be present in 8.3% of breast cancers classified as basal-like [20] and that patients with TNBC had a rate of PIK3CA mutation of approximately 13–15% [21] (http://cancer.sanger.ac.uk/cancergenome/projects/cosmic/) as compared with a rate of about 30% in ER/PgR-positive breast cancers.
In addition, the frequency of mutation of the BRAF gene (3.0%) observed in the present study population was similar to other TNBC series as well as in all subtypes (http://cancer.sanger.ac.uk/cancergenome/projects/cosmic/). The frequency of mutation of KIT and PDGFRA genes (3% each) observed was higher than reported in other TNBC series (0–1.4%) (http://cancer.sanger.ac.uk/cancergenome/projects/cosmic/) though the numbers are small.
A recent trial by Zhu et al. showed that although 42.1% of TNBC expressed KIT protein, only 1 activating mutation was detected in the series; the expression of KIT in TNBC lacked correlation with activating mutations in both KIT and PDGFRA genes [22]. Nevertheless, it might be worth investigating the potential effect of dasatinib on TNBCs expressing C-kit and PDGFRA in clinical trials.
Mutation of the AKT1 gene was 2.2% in the present series, about 1% in other series and 4% in all subtypes (http://cancer.sanger.ac.uk/cancergenome/projects/cosmic/). Likewise, point mutations have been reported in multiple components of a single signaling pathway (i.e., PIK3CA, PTEN, AKT1, as molecular components of the PI3K signaling pathway) indicating the potential relevance of specific signaling pathways as a therapeutic target in breast cancer [23]. In the present series, PIK3CA pathway mutations (specifically: PIK3CA + AKT1 + AKT2) were found in 22 (16%) patients. Inhibitors of the PI3K/AKT/mTOR pathway, frequently de-regulated in TNBC, are acquiring a growing interest and several inhibitors are in preclinical development or already in early phase clinical trials [24, 25].
Mutations in the tested oncogenes were not associated with breast cancer recurrence in this TNBC subset of patients. However, our analysis suggested that the presence of at least one targeted mutation may be prognostic in patients with TNBC, having observed a numerically increased odds of recurrence (OR 2.28; 95% CI 0.88–5.92), and similar results were evident when assessing the association of at least one of PIK3CA, AKT1, and AKT2 mutation with the recurrence (ORR 2.60; 95% CI 0.62–6.41). Nevertheless, evaluation in larger datasets is warranted. Evidence from the present trial showed that mutational status either single or pathway mutation, failed to be predictive for benefit of CM maintenance, as the test of interaction was not statistically significant (p = 0.73 and p = 0.48, respectively). This may be in part attributed to the restricted sample size that represents one of the limits of the study. The prognostic role of mutation status was extensively investigated either for PIK3CA mutations [26] or for TP53 status as a prognostic or predictive marker in TNBC [23]. PIK3CA mutations role of cell-free DNA (cfDNA) in early-stage TNBC patients was recently associated with a more favorable prognosis. After a median follow-up of 54.4 months, the presence of PIK3CA mutations of cfDNA was significantly associated with relapse-free survival (p = 0.0072) and breast cancer-specific survival (p = 0.016) [27]. These discordant results—if compared with the data of the present study—may be due to the differences between PIK3CA mutation detection in plasma and in tumor tissue DNA. In fact, it has been hypothesized that this discordance might be a result of the problem of tumor heterogeneity [28].
Indeed, a recent pooled analysis confirmed that the presence of a PIK3CA mutation represents an independent negative prognostic factor [29]. However, to make clarity on this subject, more studies focusing on specific exons mutations would be useful [30].
Although TP53 mutation/expression status did not show any significant implications in terms of prognosis for patients with TNBC, the discrepancy between mutation and expression of TP53 may indicate poor prognosis in TNBC patients [31].
Another limit of the study was that TP53 as well as other suppressor genes commonly mutated in TNBC including PTEN, PIK3R1, TSC1/2, or BRCA1/2 are not included in the panel of genes evaluated, so we were unable to assess the prognostic value of these genes in our series. However, we selected a pre-validated assay (OncoCarta Panel v1.0) that is enriched in oncogenes to prioritize the investigation of mutations that might also be clinically actionable, although less frequently reported in TNBCs. Recent results from the SAFIR-01 study (NCT01414933), which performed a comparative genomic hybridization (CGH) array and Sanger sequencing of PIK3CA and AKT1 in patients with metastatic breast cancer, reported a targetable genomic alteration in 195 (46%) patients. Therapy was therefore personalized in 55 (13%) patients [32] and of those only 9% had an objective response. These results do not appear encouraging, but the study population was heavily pretreated. Very low response rates are frequently seen in patients with breast cancer previously treated with targeted agents [33]. Response rates are similar when pretreated patients receive treatment of physician choice in 3rd or more line of treatment [34]. However, the clinical relevance of many of these alterations remains to be established. In fact, it is increasingly recognized that molecular profiling of advanced disease could help elucidate the biological basis of distant recurrence and resistance to therapy [35]. Notably, the AURORA (Aiming to Understand the Molecular Aberrations in Metastatic Breast Cancer; NCT02102165) study is another trial going in this direction, recruiting breast cancer patients across Europe. Both primary and metastatic sites are analyzed for genomic alterations, and patients are offered participation in clinical trials utilizing targeted therapeutic agents.
The AURORA pilot study was recently published reporting results in 41 patients with the primary objective of investigating the feasibility in four European recruitment sites and central pathological and sequencing facilities. Results were encouraging and at least one clinically actionable mutation was identified in 73% of patients. This pilot study demonstrated that next-generation genomic techniques are ready for international molecular screening programs in routine clinical settings, although some technical challenges remain to be addressed [36].
In the present trial, the mutational profile was performed only in the primary tumor and not repeated in the recurrence, as this analysis was retrospectively done in a subset of patients participating in the IBCSG 22-00 trial and the matching tissue for recurrence was available only for a limited number of patients. Nevertheless, the identification of critical pathways involved in carcinogenesis, metastasis, and drug resistance, as well as actionable mutations, that could emerge from these trials fuels the dream of “personalized medicine,” in which the molecular landscape of an individual’s cancer will inform clinical decision making, particularly the selection of “tailored” targeted therapies.
Finally, the results of the present study have identified a consistent subgroup of patients with TNBC and PIK3CA mutations. Since there is contradictory information about the prognostic role of PIK3CA mutation in this subset of patients, it may be worth contributing to a potential pooled analysis to elucidate the role of PI3KCA mutations/alterations in TNBC.
References
Sotiriou C, Pusztai L (2009) Gene-expression signatures in breast cancer. N Engl J Med 360(8):790–800. https://doi.org/10.1056/NEJMra0801289
Coates AS, Winer EP, Goldhirsch A et al (2015) Tailoring therapies—improving the management of early breast cancer: St Gallen international expert consensus on the primary therapy of early breast cancer 2015. Ann Oncol 26(8):1533–1546. https://doi.org/10.1093/annonc/mdv221
Carey L, Winer E, Viale G, Cameron D, Gianni L (2010) Triple-negative breast cancer: disease entity or title of convenience? Nat Rev Clin Oncol 7(12):683–692. https://doi.org/10.1038/nrclinonc.2010.154
Guiu S, Michiels S, André F et al (2012) Molecular subclasses of breast cancer: how do we define them? The IMPAKT 2012 working group statement. Ann Oncol 23:2997–3006. https://doi.org/10.1093/annonc/mds586
Haffty BG, Yang Q, Reiss M et al (2006) Locoregional relapse and distant metastasis in conservatively managed triple negative early-stage breast cancer. J Clin Oncol 24(36):5652–5657. https://doi.org/10.1200/JCO.2006.06.5664
Dent R, Trudeau M, Pritchard KI et al (2007) Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res 13(15 Pt 1):4429–4434. https://doi.org/10.1158/1078-0432.CCR-06-3045
Gluz O, Liedtke C, Gottschalk N, Pusztai L, Nitz U, Harbeck N (2009) Triple-negative breast cancer–current status and future directions. Ann Oncol 20(12):1913–1927. https://doi.org/10.1093/annonc/mdp492
Prat A, Perou CM (2011) Deconstructing the molecular portraits of breast cancer. Mol Oncol 5(1):5–23. https://doi.org/10.1016/j.molonc.2010.11.003
Lehmann BD, Bauer JA, Chen X et al (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121(7):2750–2767. https://doi.org/10.1172/JCI45014
Burstein MD, Tsimelzon A, Poage GM et al (2015) Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin Cancer Res 21(7):1688–1698. https://doi.org/10.1158/1078-0432.CCR-14-0432
Curtis C, Shah SP, Chin S-F et al (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486(7403):346–352. https://doi.org/10.1038/nature10983
Shah SP, Roth A, Goya R et al (2012) The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486(7403):395–399. https://doi.org/10.1038/nature10933
Stephens PJ, Tarpey PS, Davies H et al (2012) The landscape of cancer genes and mutational processes in breast cancer. Nature 486(7403):400–404. https://doi.org/10.1038/nature11017
Kalimutho M, Parsons K, Mittal D, López JA, Srihari S, Khanna KK (2015) Targeted therapies for triple-negative breast cancer: combating a stubborn disease. Trends Pharmacol Sci 36(12):822–846. https://doi.org/10.1016/j.tips.2015.08.009
Jurinke C, Oeth P, van den Boom D (2004) MALDI-TOF mass spectrometry: a versatile tool for high-performance DNA analysis. Mol Biotechnol 26(2):147–164. https://doi.org/10.1385/MB:26:2:147
Colleoni M, Gray KP, Gelber S et al (2016) Low-dose oral cyclophosphamide and methotrexate maintenance for hormone receptor-negative early breast cancer: International Breast Cancer Study Group Trial 22-00. J Clin Oncol. https://doi.org/10.1200/JCO.2015.65.6595
Dupont WD. (1988) Power calculations for matched case-control studies. Biometrics. 44(4):1157–1168. http://www.ncbi.nlm.nih.gov/pubmed/3233252
Santarpia L, Qi Y, Stemke-Hale K et al (2012) Mutation profiling identifies numerous rare drug targets and distinct mutation patterns in different clinical subtypes of breast cancers. Breast Cancer Res Treat 134(1):333–343. https://doi.org/10.1007/s10549-012-2035-3
Nowell PC. (1976) The clonal evolution of tumor cell populations. Science. 194(4260):23–28. http://www.ncbi.nlm.nih.gov/pubmed/959840
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A et al (2008) An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res 68(15):6084–6091. https://doi.org/10.1158/0008-5472.CAN-07-6854
Millis SZ, Gatalica Z, Winkler J et al (2015) Predictive biomarker profiling of < 6000 breast cancer patients shows heterogeneity in TNBC, with treatment implications. Clin Breast Cancer. 15(6):473–481. https://doi.org/10.1016/j.clbc.2015.04.008
Zhu Y, Wang Y, Guan B, et al. (2014) C-kit and PDGFRA gene mutations in triple negative breast cancer. Int J Clin Exp Pathol. 7(7):4280–4285. http://www.ncbi.nlm.nih.gov/pubmed/25120810
Network Cancer Genome Atlas (2012) Comprehensive molecular portraits of human breast tumours. Nature 490(7418):61–70. https://doi.org/10.1038/nature11412
Fouqué A, Jean M, van de Weghe P, Legembre P. (2016) Review of PI3K/mTOR inhibitors entering clinical trials to treat triple negative breast cancers. Recent Pat Anticancer Drug Discov. http://www.ncbi.nlm.nih.gov/pubmed/27194555
Massihnia D, Galvano A, Fanale D et al (2016) Triple negative breast cancer: shedding light onto the role of pi3k/akt/mtor pathway. Oncotarget 5:60712
Pang B, Cheng S, Sun S-P et al (2014) Prognostic role of PIK3CA mutations and their association with hormone receptor expression in breast cancer: a meta-analysis. Sci Rep 4:6255. https://doi.org/10.1038/srep06255
Takeshita T, Yamamoto Y, Yamamoto-Ibusuki M et al (2015) Prognostic role of PIK3CA mutations of cell-free DNA in early-stage triple negative breast cancer. Cancer Sci 106(11):1582–1589. https://doi.org/10.1111/cas.12813
Kurose K, Ohue Y, Wada H et al (2015) Phase Ia study of FoxP3 + CD4 treg depletion by infusion of a humanized Anti-CCR4 antibody, KW-0761, in Cancer Patients. Clin Cancer Res 21(19):4327–4336. https://doi.org/10.1158/1078-0432.CCR-15-0357
Sobhani N, Roviello G, Corona SP et al (2018) The prognostic value of PI3 K mutational status in breast cancer: a meta-analysis. J Cell Biochem. https://doi.org/10.1002/jcb.26687
Pang B, Cheng S, Sun SP et al (2014) Prognostic role of PIK3CA mutations and their association with hormone receptor expression in breast cancer: a meta-analysis. Sci Rep. 1(4):6255. https://doi.org/10.1038/srep06255
Kim J-Y, Park K, Jung HH et al (2016) Association between mutation and expression of TP53 as a potential prognostic marker of triple-negative breast cancer. Cancer Res Treat. https://doi.org/10.4143/crt.2015.430
André F, Bachelot T, Commo F et al (2014) Comparative genomic hybridisation array and DNA sequencing to direct treatment of metastatic breast cancer: a multicentre, prospective trial (SAFIR01/UNICANCER). Lancet Oncol 15(3):267–274. https://doi.org/10.1016/S1470-2045(13)70611-9
Ellard SL, Clemons M, Gelmon KA et al (2009) Randomized phase II study comparing two schedules of everolimus in patients with recurrent/metastatic breast cancer: nCIC Clinical Trials Group IND.163. J Clin Oncol 27(27):4536–4541. https://doi.org/10.1200/JCO.2008.21.3033
Cortes J, O’Shaughnessy J, Loesch D et al (2011) Eribulin monotherapy versus treatment of physician’s choice in patients with metastatic breast cancer (EMBRACE): a phase 3 open-label randomised study. Lancet 377(9769):914–923. https://doi.org/10.1016/S0140-6736(11)60070-6
de Dueñas EM, Hernández AL, Zotano AG et al (2014) Prospective evaluation of the conversion rate in the receptor status between primary breast cancer and metastasis: results from the GEICAM 2009-03 ConvertHER study. Breast Cancer Res Treat 143(3):507–515
Maetens M, Brown D, Irrthum A et al (2017) The AURORA pilot study for molecular screening of patients with advanced breast cancer-a study of the breast international group. NPJ Breast Cancer. 29(3):23. https://doi.org/10.1038/s41523-017-0026-6
Acknowledgements
We thank the patients, physicians, nurses, and data managers who participated in the International Breast Cancer Study Group (IBCSG) Trial 22-00, which was supported by the IBCSG and participating centers. Support for Trial 22-00 central coordination, data management and statistics was provided by the Swedish Cancer League; The Cancer Council Australia; Australia & New Zealand Breast Cancer Trials Group; the Frontier Science and Technology Research Foundation; the Swiss Group for Clinical Cancer Research; the Swiss Cancer League/Oncosuisse. Funding for this study was provided by Italian Ministry of Health, Ricerca Finalizzata, RF-2009-1536545.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest related to this study.
Rights and permissions
About this article
Cite this article
Munzone, E., Gray, K.P., Fumagalli, C. et al. Mutational analysis of triple-negative breast cancers within the International Breast Cancer Study Group (IBCSG) Trial 22-00. Breast Cancer Res Treat 170, 351–360 (2018). https://doi.org/10.1007/s10549-018-4767-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-018-4767-1