Abstract
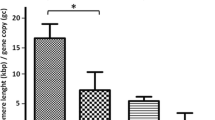
Telomere repeat binding factor 2 (TRF2) binds directly to telomeres and preserves the structural integrity of chromosome ends. In vitro models suggest that expression of TRF2 protein increases during mammary cancer progression. However, a recent study has reported that TRF2 mRNA levels tend to be lower in clinical specimens of malignant breast tissue. Here, we conduct the first large-scale investigation to assess the levels and cellular localization of the TRF2 protein in normal, pre-malignant and malignant breast tissues. Breast tissue arrays, containing normal, ductal carcinoma in situ (DCIS) and invasive carcinoma specimens, were used to assess the expression and localization of TRF2 protein. Telomere lengths were semi-quantitatively measured using a pantelomeric peptide nucleic acid probe. A mixed effects modeling approach was used to assess the relationship between TRF2 expression and telomeric signal scores across disease states or clinical staging. We demonstrate that TRF2 is exclusively nuclear with a trend toward lower expression with increased malignancy. More case-to-case variability of TRF2 immunostaining intensity was noted amongst the invasive carcinomas than the other disease groups. Invasive carcinomas also displayed variable telomere lengths while telomeres in normal mammary epithelium were generally longer. Statistical analyses revealed that increased TRF2 immunostaining intensity in invasive carcinomas is associated with shorter telomeres and shorter telomeres correlate with a higher TNM stage. All immortalized and cancer cell lines within the array displayed strong, nuclear TRF2 expression. Our data indicate that elevated expression of TRF2 is not a frequent occurrence during the transformation of breast cancer cells in vivo, but higher levels of this telomere-binding protein may be important for protecting advanced cancer cells with critically short telomeres. Our findings also reinforce the concept that serially propagated cancer cells, although tumor-derived, may not model all types of authentic tumors especially those demonstrating genetic heterogeneity.




Similar content being viewed by others
References
Blackburn EH (1991) Structure and function of telomeres. Nature 350:569–573
Holt SE, Shay JW (1999) Role of telomerase in cellular proliferation and cancer. J Cell Physiol 180:10–18
Shay JK, Bacchetti S (1997) A survey of telomerase activity in human cancer. Eur J Cancer 33:787–791
Meeker AK, Hicks JL, Gabrielson E, Strauss WM, De Marzo AM, Argani P (2004) Telomere shortening occurs in subsets of normal breast epithelium as well as in situ and invasive carcinoma. Am J Pathol 164:925–935
Meeker AK, Hicks JL, Iacobuzio-Donahue CA, Montgomery EA, Westra WH, Chan TY, Ronnett BM, De Marzo AM (2004) Telomere length abnormalities occur early in the initiation of epithelial carcinogenesis. Clin Cancer Res 10:3317–3326
Deng Y, Chang S (2007) Role of telomeres and telomerase in genomic instability, senescence, and cancer. Lab Invest 87:1071–1076
de Lange T (2002) Protection of mammalian telomeres. Oncogene 21:532540
de Lange T (2005) Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev 19:2100–2110
Karlseder J, Smogorsewska A, de Lange T (2002) Senescence induced by altered telomere state, not telomere loss. Science 295:2446–2449
Karlseder J (2003) Telomere repeat binding factors: keeping the ends in check. Cancer Lett 194:189–197
Baykal A, Rosen D, Zhou C, Liu J, Sahin AA (2004) Telomerase in breast cancer. Adv Anat Pathol 11:262–268
Karlseder J, Broccoli D, Dai Y, Hardy S, de Lange T (1999) p53- and ATM-dependent apoptosis induced by telomeres lacking TRF2. Science 283:1321–1325
Nijjar T, Bassett E, Garbe J, Takenaka Y, Stampfer MR, Gilley D, Yaswen P (2005) Accumulation and altered localization of telomere-associated protein TRF2 in immortally transformed and tumor-derived human breast cells. Oncogene 24:3369–3376
Blanco R, Munoz P, Flores JM, Klatt P, Blasco MA (2007) Telomerase abrogation dramatically accelerates TRF2-induced epithelial carcinogenesis. Genes Dev 21:206–220
Salhab M, Jiang WG, Newbold RF, Mokbel K (2008) The expression of gene transcripts of telomere-associated proteins in human breast cancer: correlation with clinic-pathological parameters and clinical outcome. Breast Cancer Res Treat 109:35–46
Di X, Bright AT, Gaskins, Robert J, Holt S, Gewirtz DA, Elmore LW (2008) A chemotherapy-associated senescence bystander effect in breast cancer cells. Cancer Biol Ther 6:864–872
Shay JW, Tomlinson G, Piatyszek MA, Gollahon LS (1995) Spontaneous in vitro immortalization of breast epithelial cells from a patient with Li-Fraumeni syndrome. Mol Cell Biol 15:425–432
Soule HD, Maloney TM, Wolams SR, Peterson WD et al (1990) Isolation and characterization of a spontaneously immortalized human breast epithelial cell line, MCF-10. Cancer Res 18:6075–6086
Elmore LW, Rehder CW, Di X, McChesney PA, Jackson-Cook CK, Gewirtz DA, Holt SE (2002) Adriamycin induces telomere length-dependent senescence in breast tumor cells. J Biol Chem 277:35509–35515
Deng Z, Atanasiu C, Burg JS, Broccoli D, Lieberman PM (2003) Telomere repeat binding factors TRF1, TRF2, and hRAP1 modulate replication of Epstein-Barr virus OriP. J Virol 77:1992–2001
Huda N, Tanaka H, Mendonca MS, Gilley D (2009) DNA damage-induced phosphorylation of TRF2 is required for the fast pathways of DNA double-strand break repair. Mol Cell Biol 13:3597–3604
Diehl MC, Idowu MO, Kimmelshue K, York TP, Elmore LW, Holt SE (2009) Elevated expression of nuclear Hsp90 in invasive breast tumors. Cancer Biol Ther 8:1952–1961
Jenkins RB, Qian J, Lee HK, Huang H et al (1998) A molecular cytogenetic analysis of 7q31 in prostate cancer. Cancer Res 58:759–766
R Development Core Team (2009) R: a language and environment for the statistical computing. R. Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL:http://www.R-project.org
Takai KK, Hooper S, Blackwood S, Gandhi R, de Lange T (2010) In vivo stoichiometry of shelterin components. J Biol Chem 285:1457–1467
Lin K-W, Yan J (2005) The telomere length dynamic and methods of its assessment. J Cell Mol Med 9:977–989
Odagiri E, Kanada N, Jibiki K, Demura R, Aikawa E, Demura H (1994) Reduction of telomeric length and c-erbB2 gene amplification in human breast cancer, fibroadenoma, and gynecomastia. Relationship to histologic grade and clinical parameters. Cancer 73:2978–2984
Poonepalli A, Banerjee B, Ramnarayanan K, Palanisamy N, Putti TC, Hande MP (2008) Telomere-mediated genomic instability and the clinico-pathological parameters in breast cancer. Genes Chromosomes Cancer 47:1098–1109
Raynaud CM, Hernandez J, Llorca FP, Nuciforon P et al (2009) DNA damage repair and telomere length in normal breast, preneoplastic lesions, and invasive cancer. Am J Clin Oncol Oct 30. [Epub ahead of print]
Bradshaw PS, Stavropoulous DJ, Meyn MS (2005) Human telomeric protein TRF2 associates with genomic double-strand breaks as an early response to DNA damage. Nat Genet 37:193–197
Escobar PF, Patrick RJ, Rybicki LA, Weng DE, Crowe JP (2007) The 2003 revised TNM staging system for breast cancer: results of stage re-classification on survival and future comparisons among stage groups. Ann Surg Oncol 14:143–147
Fordyce CA, Heaphy CM, Bisoffi M, Wyaco JL et al (2006) Telomere content correlates with stage and prognosis in breast cancer. Breast Cancer Res Treat 99:193–202
Rogalla P, Rohen C, Bonk U, Bullerdiek J (1996) Telomeric repeat fragment lengths are not correlated to histological grading in 85 breast cancers. Cancer Lett 106:155–161
Hiyama E, Gollahon L, Kataoka T, Kuroi K, Yokoyama T, Gazdar AF, Hiyama K, Piatyszek MA, Shay JW (1996) Telomerase activity in human breast tumors. 88:116–122
Ning H, Li TT, Zhao L, Li T, Li J, Liu J, Liu Z, Fan D (2006) TRF2 promotes multidrug resistance in gastric cancer cells. Cancer Biol Ther 5:950–956
Acknowledgments
We thank the Anatomic Pathology Research Services at Virginia Commonwealth University for providing anonymized breast tissues for antibody optimization and for use of their Nikon microscopic workstation. This study was supported by a National Institutes of Health KO1 CA105050-01A131 (to LWE). It was also funded, in part, by a grant (CJC) from the National Institute of Environmental Health (R01 ES12074). Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIEHS, NIH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Diehl, M.C., Idowu, M.O., Kimmelshue, K.N. et al. Elevated TRF2 in advanced breast cancers with short telomeres. Breast Cancer Res Treat 127, 623–630 (2011). https://doi.org/10.1007/s10549-010-0988-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-010-0988-7