Abstract
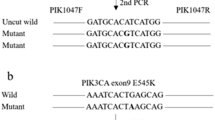
Somatic mutations in PIK3CA (encoding a class I phosphoinositide 3 kinase (PI3K) subunit) modulate PI3K signalling to influence tumour behaviour and occur in up to 40% of breast cancers. Inhibitors of PI3K signalling are entering clinical trials, but the impact of PIKC3A mutation on tumour response has yet to be clarified. This study investigated the potential utility of circulating free DNA (cfDNA) as a source for PIK3CA mutation detection in patients with breast cancer. cfDNA extracted (QIAamp Virus spin kit) from blood and matched archival tumour from 46 patients with metastatic breast cancer and 30 patients with localised, operable breast cancer was assessed for hotspot PIK3CA mutations using Amplification Refractory Mutation System (ARMS™) allele-specific PCR and Scorpion probes. PIK3CA mutations were detected in 13/46 (28%) plasma-derived and 10/46 (21%) serum-derived cfDNA samples from metastatic breast cancer patients. In 41 cases with matched tumour and plasma-derived cfDNA data, concordance (same mutation status in plasma and tumour) was 95%. Where a PIK3CA mutation was present in tumour, the ‘pick up’ in plasma-derived cfDNA was 80%. PIK3CA mutations were present in tumours from 14/30 (47%) localised breast cancers, but no PIK3CA mutations were detected in matched cfDNA. These data demonstrate feasibility and potential utility of cfDNA for PIK3CA mutation detection in patients with metastatic breast cancer. Studies are underway to qualify PIK3CA mutation in cfDNA as a predictive biomarker allowing patient stratification in clinical trials of mechanism-based therapeutics that target PI3K signalling pathways.

Similar content being viewed by others
References
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, Yan H, Gazdar A, Powell SM, Riggins GJ, Willson JK, Markowitz S, Kinzler KW, Vogelstein B, Velculescu VE (2004) High frequency of mutations of the PIK3CA gene in human cancers. Science 304(5670):554
Kang S, Bader AG, Vogt PK (2005) Phosphatidylinositol 3-kinase mutations identified in human cancer are oncogenic. Proc Natl Acad Sci USA 102(3):802–807
Gymnopoulos M, Elsliger MA, Vogt PK (2007) Rare cancer-specific mutations in PIK3CA show gain of function. Proc Natl Acad Sci USA 104(13):5569–5574
Samuels Y, Diaz LA Jr, Schmidt-Kittler O, Cummins JM, Delong L, Cheong I, Rago C, Huso DL, Lengauer C, Kinzler KW, Vogelstein B, Velculescu VE (2005) Mutant PIK3CA promotes cell growth and invasion of human cancer cells. Cancer Cell 7(6):561–573
Ihle NT, Williams R, Chow S, Chew W, Berggren MI, Paine-Murrieta G, Minion DJ, Halter RJ, Wipf P, Abraham R, Kirkpatrick L, Powis G (2004) Molecular pharmacology and antitumor activity of PX-866, a novel inhibitor of phosphoinositide-3-kinase signaling. Mol Cancer Ther 3(7):763–772
Serra V, Markman B, Scaltriti M, Eichhorn PJ, Valero V, Guzman M, Botero ML, Llonch E, Atzori F, Di Cosimo S, Maira M, Garcia-Echeverria C, Parra JL, Arribas J, Baselga J (2008) NVP-BEZ235, a dual PI3K/mTOR inhibitor, prevents PI3K signaling and inhibits the growth of cancer cells with activating PI3K mutations. Cancer Res 68(19):8022–8030
Eichhorn PJ, Gili M, Scaltriti M, Serra V, Guzman M, Nijkamp W, Beijersbergen RL, Valero V, Seoane J, Bernards R, Baselga J (2008) Phosphatidylinositol 3-kinase hyperactivation results in lapatinib resistance that is reversed by the mTOR/phosphatidylinositol 3-kinase inhibitor NVP-BEZ235. Cancer Res 68(22):9221–9230
Saal LH, Holm K, Maurer M, Memeo L, Su T, Wang X, Yu JS, Malmstrom PO, Mansukhani M, Enoksson J, Hibshoosh H, Borg A, Parsons R (2005) PIK3CA mutations correlate with hormone receptors, node metastasis, and ERBB2, and are mutually exclusive with PTEN loss in human breast carcinoma. Cancer Res 65(7):2554–2559
Campbell IG, Russell SE, Choong DY, Montgomery KG, Ciavarella ML, Hooi CS, Cristiano BE, Pearson RB, Phillips WA (2004) Mutation of the PIK3CA gene in ovarian and breast cancer. Cancer Res 64(21):7678–7681
Levine DA, Bogomolniy F, Yee CJ, Lash A, Barakat RR, Borgen PI, Boyd J (2005) Frequent mutation of the PIK3CA gene in ovarian and breast cancers. Clin Cancer Res 11(8):2875–2878
Bachman KE, Argani P, Samuels Y, Silliman N, Ptak J, Szabo S, Konishi H, Karakas B, Blair BG, Lin C, Peters BA, Velculescu VE, Park BH (2004) The PIK3CA gene is mutated with high frequency in human breast cancers. Cancer Biol Ther 3(8):772–775
Lee JW, Soung YH, Kim SY, Lee HW, Park WS, Nam SW, Kim SH, Lee JY, Yoo NJ, Lee SH (2005) PIK3CA gene is frequently mutated in breast carcinomas and hepatocellular carcinomas. Oncogene 24(8):1477–1480
Berns K, Horlings HM, Hennessy BT, Madiredjo M, Hijmans EM, Beelen K, Linn SC, Gonzalez-Angulo AM, Stemke-Hale K, Hauptmann M, Beijersbergen RL, Mills GB, van de Vijver MJ, Bernards R (2007) A functional genetic approach identifies the PI3 K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell 12(4):395–402
Leon SA, Shapiro B, Sklaroff DM, Yaros MJ (1977) Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res 37(3):646–650
Kimura H, Kasahara K, Kawaishi M, Kunitoh H, Tamura T, Holloway B, Nishio K (2006) Detection of epidermal growth factor receptor mutations in serum as a predictor of the response to gefitinib in patients with non-small-cell lung cancer. Clin Cancer Res 12(13):3915–3921
Kimura H, Suminoe M, Kasahara K, Sone T, Araya T, Tamori S, Koizumi F, Nishio K, Miyamoto K, Fujimura M, Nakao S (2007) Evaluation of epidermal growth factor receptor mutation status in serum DNA as a predictor of response to gefitinib (IRESSA). Br J Cancer 97(6):778–784
Sorenson GD (2000) Detection of mutated KRAS2 sequences as tumor markers in plasma/serum of patients with gastrointestinal cancer. Clin Cancer Res 6(6):2129–2137
Yancovitz M, Yoon J, Mikhail M, Gai W, Shapiro RL, Berman RS, Pavlick AC, Chapman PB, Osman I, Polsky D (2007) Detection of mutant BRAF alleles in the plasma of patients with metastatic melanoma. J Mol Diagn 9(2):178–183
Daniotti M, Vallacchi V, Rivoltini L, Patuzzo R, Santinami M, Arienti F, Cutolo G, Pierotti MA, Parmiani G, Rodolfo M (2007) Detection of mutated BRAFV600E variant in circulating DNA of stage III-IV melanoma patients. Int J Cancer 120(11):2439–2444
Board RE, Thelwell NJ, Ravetto PF, Little S, Ranson M, Dive C, Hughes A, Whitcombe D (2008) Multiplexed assays for detection of mutations in PIK3CA. Clin Chem 54(4):757–760
Coombs NJ, Gough AC, Primrose JN (1999) Optimisation of DNA and RNA extraction from archival formalin-fixed tissue. Nucleic Acids Res 27(16):e12
Diehl F, Li M, Dressman D, He Y, Shen D, Szabo S, Diaz LA Jr, Goodman SN, David KA, Juhl H, Kinzler KW, Vogelstein B (2005) Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc Natl Acad Sci USA 102(45):16368–16373
Wang JY, Hsieh JS, Chang MY, Huang TJ, Chen FM, Cheng TL, Alexandersen K, Huang YS, Tzou WS, Lin SR (2004) Molecular detection of APC, K-ras, and p53 mutations in the serum of colorectal cancer patients as circulating biomarkers. World J Surg 28(7):721–726
Hayes DF, Cristofanilli M, Budd GT, Ellis MJ, Stopeck A, Miller MC, Matera J, Allard WJ, Doyle GV, Terstappen LW (2006) Circulating tumor cells at each follow-up time point during therapy of metastatic breast cancer patients predict progression-free and overall survival. Clin Cancer Res 12(14 Pt 1):4218–4224
Ramirez JL, Sarries C, de Castro PL, Roig B, Queralt C, Escuin D, de Aguirre I, Sanchez JM, Manzano JL, Margeli M, Sanchez JJ, Astudillo J, Taron M, Rosell R (2003) Methylation patterns and K-ras mutations in tumor and paired serum of resected non-small-cell lung cancer patients. Cancer Lett 193(2):207–216
Acknowledgements
Ruth Board was funded via a Cancer Research UK and AstraZeneca Clinical Pharmacology Fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Board, R.E., Wardley, A.M., Dixon, J.M. et al. Detection of PIK3CA mutations in circulating free DNA in patients with breast cancer. Breast Cancer Res Treat 120, 461–467 (2010). https://doi.org/10.1007/s10549-010-0747-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-010-0747-9