Abstract
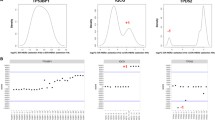
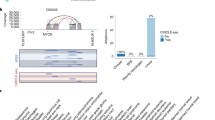
Overexpression and alternative splicing of CD44 have been implicated in tumour progression. Here we describe the identification of a high level amplification of human 11p13, encompassing the CD44 gene, in primary breast cancers and cell lines and test whether CD44 acts as the driver of this amplicon. aCGH analysis revealed 11p13 amplification in 3% (3/100) of primary breast carcinomas and in two cell lines. The minimal region of amplification was 34.38–37.62 Mb. Amplification was confirmed by dual-colour FISH in these cell lines and further validated by CISH in an independent tumour cohort. CD44 expression in primary breast cancers was significantly associated with features of basal-like breast cancer. Detection of CD44 expression in breast cancer cell lines confirmed moderate to high expression in basal-like cell lines and minimal expression in luminal cell lines. In both, primary breast cancers and cell lines, 11p13 amplification was associated with high levels of CD44 mRNA expression. CD44 alternative splicing was detected in four of nine cell lines and in tumour samples, irrespective of the amplification status. RNAi mediated knock down of CD44 failed to reveal an increased dependence on CD44 expression for proliferation or survival in amplified cell lines. Given that expression of CD44 is not an absolute requirement for the survival of cells harbouring CD44 gene amplification, CD44 is unlikely to be a driver of the 11p13 amplicon.






Similar content being viewed by others
Abbreviations
- aCGH:
-
Array chromosomal genomic hybridisation
- CISH:
-
Chromogenic in situ hybridisation
- Ck:
-
Cytokeratin
- ER:
-
Oestrogen receptor
- EGFR:
-
Epidermal growth factor receptor
- FISH:
-
Fluorescent in situ hybridisation
- PR:
-
Progesterone receptor
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
References
Naor D, Sionov RV, Ish-Shalom D (1997) CD44: structure, function, and association with the malignant process. Adv Cancer Res 71:241–319. doi:10.1016/S0065-230X(08)60101-3
Ponta H, Sherman L, Herrlich PA (2003) CD44: from adhesion molecules to signalling regulators. Nat Rev Mol Cell Biol 4:33–45. doi:10.1038/nrm1004
Joensuu H, Klemi PJ, Toikkanen S, Jalkanen S (1993) Glycoprotein CD44 expression and its association with survival in breast cancer. Am J Pathol 143:867–874
Auvinen P, Tammi R, Tammi M, Johansson R, Kosma VM (2005) Expression of CD 44s, CD 44 v 3 and CD 44 v 6 in benign and malignant breast lesions: correlation and colocalization with hyaluronan. Histopathology 47:420–428. doi:10.1111/j.1365-2559.2005.02220.x
Berner HS, Nesland JM (2001) Expression of CD44 isoforms in infiltrating lobular carcinoma of the breast. Breast Cancer Res Treat 65:23–29. doi:10.1023/A:1006417412046
Kinoshita J, Haga S, Shimizu T, Imamura H, Watanabe O, Kajiwara T (1999) The expression of variant exon v7–v8 CD44 antigen in relation to lymphatic metastasis of human breast cancer. Breast Cancer Res Treat 53:177–183. doi:10.1023/A:1006130601575
Rys J, Kruczak A, Lackowska B, Jaszcz-Gruchala A, Brandys A, Stelmach A, Reinfuss M (2003) The role of CD44v3 expression in female breast carcinomas. Pol J Pathol 54:243–247
Diaz LK, Zhou X, Wright ET, Cristofanilli M, Smith T, Yang Y, Sneige N, Sahin A, Gilcrease MZ (2005) CD44 expression is associated with increased survival in node-negative invasive breast carcinoma. Clin Cancer Res 11:3309–3314. doi:10.1158/1078-0432.CCR-04-2184
Ma W, Deng Y, Zhou L (2005) The prognostic value of adhesion molecule CD44v6 in women with primary breast carcinoma: a clinicopathologic study. Clin Oncol (R Coll Radiol) 17:258–263. doi:10.1016/j.clon.2005.02.007
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100:3983–3988. doi:10.1073/pnas.0530291100
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW, Hoey T, Gurney A, Huang EH, Simeone DM, Shelton AA, Parmiani G, Castelli C, Clarke MF (2007) Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci USA 104:10158–10163. doi:10.1073/pnas.0703478104
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ (2005) Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res 65:10946–10951. doi:10.1158/0008-5472.CAN-05-2018
Grimshaw MJ, Cooper L, Papazisis K, Coleman JA, Bohnenkamp HR, Chiapero-Stanke L, Taylor-Papadimitriou J, Burchell JM (2008) Mammosphere culture of metastatic breast cancer cells enriches for tumorigenic breast cancer cells. Breast Cancer Res 10:R52. doi:10.1186/bcr2106
Sheridan C, Kishimoto H, Fuchs RK, Mehrotra S, Bhat-Nakshatri P, Turner CH, Goulet R Jr, Badve S, Nakshatri H (2006) CD44+/CD24-breast cancer cells exhibit enhanced invasive properties: an early step necessary for metastasis. Breast Cancer Res 8:R59. doi:10.1186/bcr1610
Ponti D, Costa A, Zaffaroni N, Pratesi G, Petrangolini G, Coradini D, Pilotti S, Pierotti MA, Daidone MG (2005) Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res 65:5506–5511. doi:10.1158/0008-5472.CAN-05-0626
Isacke CM, Sauvage CA, Hyman R, Lesley J, Schulte R, Trowbridge IS (1986) Identification and characterization of the human Pgp-1 glycoprotein. Immunogenetics 23:326–332. doi:10.1007/BF00398797
Korsching E, Packeisen J, Agelopoulos K, Eisenacher M, Voss R, Isola J, van Diest PJ, Brandt B, Boecker W, Buerger H (2002) Cytogenetic alterations and cytokeratin expression patterns in breast cancer: integrating a new model of breast differentiation into cytogenetic pathways of breast carcinogenesis. Lab Invest 82:1525–1533
Natrajan R, Lambros MB, Rodriguez-Pinilla SM, Moreno-Bueno G, Tan DSP, Marchio C, Vatcheva R, Rayter S, Mahler-Araujo B, Fulford LG, Hungermann D, Mackay A, Grigoriadis A, Fenwick K, Tamber N, Hardisson D, Tutt A, Palacios J, Lord CJ, Buerger H, Ashworth A, Reis-Filho JS (2009) Tiling path genomic profiling of grade III invasive ductal breast cancers. Clin Cancer Res 15(8). doi: 10.1158/1078-0432.CCR-08-1878
Ellis IO, Galea M, Broughton N, Locker A, Blamey RW, Elston CW (1992) Pathological prognostic factors in breast cancer. II. Histological type. Relationship with survival in a large study with long-term follow-up. Histopathology 20:479–489
Singletary SE, Connolly JL (2006) Breast cancer staging: working with the sixth edition of the AJCC cancer staging manual. CA Cancer J Clin 56:37–47. doi:10.3322/canjclin.56.1.37 quiz 50-31
Tan DS, Marchio C, Jones RL, Savage K, Smith IE, Dowsett M, Reis-Filho JS (2008) Triple negative breast cancer: molecular profiling and prognostic impact in adjuvant anthracycline-treated patients. Breast Cancer Res Treat 111:27–44. doi:10.1007/s10549-007-9756-8
Bhargava R, Gerald WL, Li AR, Pan Q, Lal P, Ladanyi M, Chen B (2005) EGFR gene amplification in breast cancer: correlation with epidermal growth factor receptor mRNA and protein expression and HER-2 status and absence of EGFR-activating mutations. Mod Pathol 18:1027–1033. doi:10.1038/modpathol.3800438
Reis-Filho JS, Milanezi F, Carvalho S, Simpson PT, Steele D, Savage K, Lambros MB, Pereira EM, Nesland JM, Lakhani SR, Schmitt FC (2005) Metaplastic breast carcinomas exhibit EGFR, but not HER2, gene amplification and overexpression: immunohistochemical and chromogenic in situ hybridization analysis. Breast Cancer Res 7:R1028–R1035. doi:10.1186/bcr1341
Marchio C, Iravani M, Natrajan R, Lambros MB, Savage K, Tamber N, Fenwick K, Mackay A, Senetta R, Di Palma S, Schmitt FC, Bussolati G, Ellis LO, Ashworth A, Sapino A, Reis-Filho JS (2008) Genomic and immunophenotypical characterization of pure micropapillary carcinomas of the breast. J Pathol 215:398–410. doi:10.1002/path.2368
Marchio C, Natrajan R, Shiu KK, Lambros MB, Rodriguez-Pinilla SM, Tan DS, Lord CJ, Hungermann D, Fenwick K, Tamber N, Mackay A, Palacios J, Sapino A, Buerger H, Ashworth A, Reis-Filho JS (2008) The genomic profile of HER2-amplified breast cancers: the influence of ER status. J Pathol 216:399–407. doi:10.1002/path.2423
Arriola E, Marchio C, Tan DS, Drury SC, Lambros MB, Natrajan R, Rodriguez-Pinilla SM, Mackay A, Tamber N, Fenwick K, Jones C, Dowsett M, Ashworth A, Reis-Filho JS (2008) Genomic analysis of the HER2/TOP2A amplicon in breast cancer and breast cancer cell lines. Lab Invest 88:491–503. doi:10.1038/labinvest.2008.19
Coe BP, Ylstra B, Carvalho B, Meijer GA, Macaulay C, Lam WL (2007) Resolving the resolution of array CGH. Genomics 89:647–653. doi:10.1016/j.ygeno.2006.12.012
Tan DS, Lambros MB, Natrajan R, Reis-Filho JS (2007) Getting it right: designing microarray (and not ‘microawry’) comparative genomic hybridization studies for cancer research. Lab Invest 87:737–754. doi:10.1038/labinvest.3700593
Gunnarsson R, Staaf J, Jansson M, Ottesen AM, Goransson H, Liljedahl U, Ralfkiaer U, Mansouri M, Buhl AM, Smedby KE, Hjalgrim H, Syvanen AC, Borg A, Isaksson A, Jurlander J, Juliusson G, Rosenquist R (2008) Screening for copy-number alterations and loss of heterozygosity in chronic lymphocytic leukemia—a comparative study of four differently designed, high resolution microarray platforms. Genes Chromosom Cancer 47:697–711. doi:10.1002/gcc.20575
Hicks J, Krasnitz A, Lakshmi B, Navin NE, Riggs M, Leibu E, Esposito D, Alexander J, Troge J, Grubor V, Yoon S, Wigler M, Ye K, Borresen-Dale AL, Naume B, Schlicting E, Norton L, Hagerstrom T, Skoog L, Auer G, Maner S, Lundin P, Zetterberg A (2006) Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 16:1465–1479. doi:10.1101/gr.5460106
Lambros MB, Simpson PT, Jones C, Natrajan R, Westbury C, Steele D, Savage K, Mackay A, Schmitt FC, Ashworth A, Reis-Filho JS (2006) Unlocking pathology archives for molecular genetic studies: a reliable method to generate probes for chromogenic and fluorescent in situ hybridization. Lab Invest 86:398–408. doi:10.1038/labinvest.3700390
Reis-Filho JS, Pinheiro C, Lambros MB, Milanezi F, Carvalho S, Savage K, Simpson PT, Jones C, Swift S, Mackay A, Reis RM, Hornick JL, Pereira EM, Baltazar F, Fletcher CD, Ashworth A, Lakhani SR, Schmitt FC (2006) EGFR amplification and lack of activating mutations in metaplastic breast carcinomas. J Pathol 209:445–453. doi:10.1002/path.2004
Hudson DL, Sleeman J, Watt FM (1995) CD44 is the major peanut lectin-binding glycoprotein of human epidermal keratinocytes and plays a role in intercellular adhesion. J Cell Sci 108(Pt 5):1959–1970
Roscic-Mrkic B, Fischer M, Leemann C, Manrique A, Gordon CJ, Moore JP, Proudfoot AE, Trkola A (2003) RANTES (CCL5) uses the proteoglycan CD44 as an auxiliary receptor to mediate cellular activation signals and HIV-1 enhancement. Blood 102:1169–1177. doi:10.1182/blood-2003-02-0488
Nielsen TO, Hsu FD, Jensen K, Cheang M, Karaca G, Hu Z, Hernandez-Boussard T, Livasy C, Cowan D, Dressler L, Akslen LA, Ragaz J, Gown AM, Gilks CB, van de Rijn M, Perou CM (2004) Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 10:5367–5374. doi:10.1158/1078-0432.CCR-04-0220
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T, Clark L, Bayani N, Coppe JP, Tong F, Speed T, Spellman PT, DeVries S, Lapuk A, Wang NJ, Kuo WL, Stilwell JL, Pinkel D, Albertson DG, Waldman FM, McCormick F, Dickson RB, Johnson MD, Lippman M, Ethier S, Gazdar A, Gray JW (2006) A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10:515–527. doi:10.1016/j.ccr.2006.10.008
Chin L, Gray JW (2008) Translating insights from the cancer genome into clinical practice. Nature 452:553–563. doi:10.1038/nature06914
Woodward WA, Sulman EP (2008) Cancer stem cells: markers or biomarkers? Cancer Metastasis Rev 27:459–470. doi:10.1007/s10555-008-9130-2
Jarvinen AK, Autio R, Kilpinen S, Saarela M, Leivo I, Grenman R, Makitie AA, Monni O (2008) High-resolution copy number and gene expression microarray analyses of head and neck squamous cell carcinoma cell lines of tongue and larynx. Genes Chromosom Cancer 47:500–509. doi:10.1002/gcc.20551
Fukuda Y, Kurihara N, Imoto I, Yasui K, Yoshida M, Yanagihara K, Park JG, Nakamura Y, Inazawa J (2000) CD44 is a potential target of amplification within the 11p13 amplicon detected in gastric cancer cell lines. Genes Chromosom Cancer 29:315–324. doi:10.1002/1098-2264(2000)9999:9999<::AID-GCC1047>3.0.CO;2-E
Savage K, Lambros MB, Robertson D, Jones RL, Jones C, Mackay A, James M, Hornick JL, Pereira EM, Milanezi F, Fletcher CD, Schmitt FC, Ashworth A, Reis-Filho JS (2007) Caveolin 1 is overexpressed and amplified in a subset of basal-like and metaplastic breast carcinomas: a morphologic, ultrastructural, immunohistochemical, and in situ hybridization analysis. Clin Cancer Res 13:90–101. doi:10.1158/1078-0432.CCR-06-1371
Chin K, DeVries S, Fridlyand J, Spellman PT, Roydasgupta R, Kuo WL, Lapuk A, Neve RM, Qian Z, Ryder T, Chen F, Feiler H, Tokuyasu T, Kingsley C, Dairkee S, Meng Z, Chew K, Pinkel D, Jain A, Ljung BM, Esserman L, Albertson DG, Waldman FM, Gray JW (2006) Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10:529–541. doi:10.1016/j.ccr.2006.10.009
Andre F, Job B, Dessen P, Tordai A, Michiels S, Liedtke C, Richon C, Yan K, Wang B, Vassal G, Delaloge S, Hortobagyi GN, Symmans WF, Lazar V, Pusztai L (2009) Molecular characterization of breast cancer with high-resolution oligonucleotide comparative genomic hybridization array. Clin Cancer Res 15:441–451. doi:10.1158/1078-0432.CCR-08-1791
Charafe-Jauffret E, Ginestier C, Monville F, Finetti P, Adelaide J, Cervera N, Fekairi S, Xerri L, Jacquemier J, Birnbaum D, Bertucci F (2006) Gene expression profiling of breast cell lines identifies potential new basal markers. Oncogene 25:2273–2284. doi:10.1038/sj.onc.1209254
Pegram M, Slamon D (2000) Biological rationale for HER2/neu (c-erbB2) as a target for monoclonal antibody therapy. Semin Oncol 27:13–19
Guan Y, Kuo WL, Stilwell JL, Takano H, Lapuk AV, Fridlyand J, Mao JH, Yu M, Miller MA, Santos JL, Kalloger SE, Carlson JW, Ginzinger DG, Celniker SE, Mills GB, Huntsman DG, Gray JW (2007) Amplification of PVT1 contributes to the pathophysiology of ovarian and breast cancer. Clin Cancer Res 13:5745–5755. doi:10.1158/1078-0432.CCR-06-2882
Kao J, Pollack JR (2006) RNA interference-based functional dissection of the 17q12 amplicon in breast cancer reveals contribution of coamplified genes. Genes Chromosom Cancer 45:761–769. doi:10.1002/gcc.20339
Acknowledgments
This work was funded by Breakthrough Breast Cancer grants to JSR-F and CMI. We acknowledge NHS funding to the NIHR Biomedical Research Centre. We thank Kay Savage and Suzanne Parry (Breakthrough Histopathology Laboratory, The Institute of Cancer Research) who performed the TMA immunohistochemical staining.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Pamela Klingbeil and Rachael Natrajan have contributed equally to this work. RN and RV performed aCGH and data analysis and RV performed FISH and CISH. CM and JSR-F interpreted the immunohistochemical and in situ hybridisation experiments. JP and HB provided the breast cancer samples for aCGH. PK and GE performed all the cell based assays. CMI and JSR-F designed the experiments. PK, RN, JSR-F and CMI wrote the manuscript.
Rights and permissions
About this article
Cite this article
Klingbeil, P., Natrajan, R., Everitt, G. et al. CD44 is overexpressed in basal-like breast cancers but is not a driver of 11p13 amplification. Breast Cancer Res Treat 120, 95–109 (2010). https://doi.org/10.1007/s10549-009-0380-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-009-0380-7