Abstract
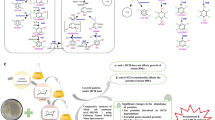
Various hydrocarbons have been released into the environment as a result of industrialization. An effective way of removing these materials without further environmental contamination is microbial bioremediation. Mycobacterium gilvum PYR-GCK, a bacteria isolated from a PAH polluted estuary, was studied using comparative shotgun proteomics to gain insight on its molecular activity while using pyrene and glucose as sole carbon and energy sources. Based on annotated genomic information, a confirmation analysis was first performed to confirm its pyrene degradation activity, using gas chromatography–mass spectrometry technology. One dimensional gel electrophoresis and liquid chromatography–mass spectrometry technologies employed in the proteomics analysis revealed the expression of pyrene degrading gene products along with upregulated expression of proteins functioning in the glyoxylate and shikimate pathways, in the pyrene-induced cells. The study also revealed the pathway of pyrene degraded intermediates, via partial gluconeogenesis, into the pentose phosphate pathway to produce precursors for nucleotides and amino acids biosynthesis.


Similar content being viewed by others
References
Adhikari D (2008) Microbial response to different carbon source amendments in agricultural soils as monitored by culture-independent techniques. ProQuest, UMI Dissertation Publishing, p 82
Baek S, Kweon O, Kim S-J et al (2009) ClassRHO: a platform for classification of bacterial rieske non-heme iron ring hydroxylating oxygenases. J Microbiol Methods 76:307–309
Bai NJ, Pai MR, Murthy PS, Venkitasubramanian TA (1975) Pathways of carbohydrate metabolism in Mycobacterium tuberculosis H37Rv. Can J Microbiol 21:1688–1691
Betts JC, Lukey PT, Robb LC, McAdam RA, Duncan K (2002) Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol Microbiol 43(3):717–731
Boonchan S, Brit ML, Stanley GA (1998) Surfactant-enhanced biodegradation of high molecular weight polycyclic aromatic hydrocarbons by Stenotrophomonas maltophilia. Biotechnol Bioeng 59:482–494
Bradford MM (1976) Rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brezna B, Khan A, Cerniglia CE (2003) Molecular characterization of dioxygenases from polycyclic aromatic hydrocarbon-degrading Mycobacterium spp. FEMS Microbiol Lett 223:177–183
Cerniglia CE (1992) Biodegradation of polycyclic aromatic hydrocarbons. Biodegradation 3:351–368
Cheung PY, Kinkle BK (2001) Mycobacterium diversity and pyrene mineralization in petroleum contaminated soils. Appl Environ Microbiol 67:2222–2229
Cohen-Bazire G, Sistrom WR, Stainer RY (1957) Kinetic studies of pigment synthesis by non-sulfur purple bacteria. J Cell Comp Physiol 49:25–68
Dean-Ross D, Cerniglia CE (1996) Degradation of pyrene by Mycobacterium flavescens. Appl Microbiol Biotechnol 46:307–312
Doroshenko VG, Tsyrenzhapova IS, Krylov AA et al (2010) Pho regulon promoter-mediated transcription of the key pathway gene aroGFbr improves the performance of an l-phenylalanine-producing Escherichia coli strain. Appl Microbiol Biotechnol 88(6):1287–1295
Fuente JL, Rumbero A, Martin JF, Liras P (1997) δ-1-Piperideine-6-carboxylate dehydrogenase, a new enzyme that forms a-aminoadipate in Streptomyces clavuligerus and other cephamycin C-producing Actinomycetes. Biochem J 327:59–64
Gibson DT, Subramanian V (1984) Microbial degradation of aromatic hydrocarbons. In: Gibson DT (ed) Microbial degradation of organic compounds. Marcel Dekker Inc., New York, pp 181–252
Grosser RJ, Warshawsky D, Vestal JR (1991) Indigenous and enhanced mineralization of pyrene, benzo(a)pyrene, and carbazole in soils. Appl Environ Microbiol 57:3462–3469
Heitkamp MA, Freeman JP, Cerniglia CE (1987) Naphthalene biodegradation in environmental microcosms: estimates of degradation rates and characterization of metabolites. Appl Environ Microbiol 53:1–13
HSDB: Hazardous Substances Data Bank. Polycyclic aromatic hydrocarbon (CASRN: 130498-29-2), Toxicology Data Network, United States National Library of Health Medicine. (http://toxnet.nlm.nih.gov/cgi-bin/sis/search/r?dbs?hsdb:@term?@DOCNO?7092)
Ishihama Y, Oda Y, Tabata T et al (2005) Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol Cell Proteomics 4:1265–1272
Kanaly RA, Harayama S (2000) Biodegradation of high-molecular weight polycyclic aromatic hydrocarbons by bacteria. J Bacteriol 182:2059–2067
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kazunga C, Aitken MD (2000) Products from the incomplete metabolism of pyrene by polycyclic aromatic hydrocarbon-degrading bacteria. Appl Environ Microbiol 66:1917–1922
Kim TJ, Lee EY, Kim YJ, Cho KS, Ryu HW (2003) Degradation of polyaromatic hydrocarbons by Burkholderia cepacia 2A-12. World J Microbiol Biotechnol 19:411–417
Kim Y-K, Engesser K-H, Cerniglia CE (2004a) Numerical and genetic analysis of polycyclic aromatic hydrocarbon-degrading Mycobacteria. Microb Ecol 50:110–119
Kim YH, Moody JD, Freeman JP et al (2004b) Evidence for the existence of PAH-quinone reductase and catechol-O-methyltransferase in Mycobacterium vanbaalenii PYR-1. J Ind Microbiol Biotechnol 31:507–516
Kim Y-H, Engesser K-H, Cerniglia CE (2005) Numerical and ribosomal RNA gene sequence analysis of polycyclic aromatic hydrocarbon-degrading mycobacteria. Microb Ecol 50:110–119
Kim YH, Cho K, Yun SH et al (2006) Analysis of aromatic catabolic pathways in Pseudomonas putida KT 2440 using a combined proteomic approach: 2-DE/MS and cleavable isotope-coded affinity tag analysis. Proteomics 6:1301–1318
Kim S-J, Kweon O, Jones RC et al (2007) Complete and integrated pyrene degradation pathway in Mycobacterium Vanbaalenii PYR-1 based on systems biology. J Bacteriol 189(2):464–472
Kondrashov FA, Koonin EV, Morgunov IG et al (2006) Evolution of glyoxylate cycle enzymes in Metazoa: evidence of multiple horizontal transfer events and pseudogene formation. Biol Direct 1:31
Kweon O, Kim SJ, Holland RD et al (2011) Polycyclic aromatic hydrocarbon metabolic network in Mycobacterium vanbaalenii PYR-1. J Bacteriol 193(17):4326–4337
Liu H, Sadygov R, Yates J (2004) A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem 76:4193–4201
Lorenz M, Fink G (2002) Life and death in a macrophage: role of the glyoxylate cycle in virulence. Eukaryot Cell 1(5):657–662
Maeda H, Dudareva N (2012) The shikimate pathway and aromatic aminoacid biosynthesis in plants. Annu Rev Plant Biol 63:73–105
Nakajima M, Nishino Y, Tamura M et al (2009) Microbial conversion of glucose to a novel chemical building block, 2-pyrone-4,6-dicarboxylic acid. Metab Eng 11(4–5):213–220
Noor E, Eden E, Milo R, Alon U (2010) Central carbon metabolism as a minimal biochemical walk between precursors for biomass and energy. Mol Cell 39:809–820
Powell DW, Weaver CM, Jennings JL et al (2004) Cluster analysis of mass spectrometry data reveals a novel component of SAGA. Mol Cell Biol 24:7249–7259
Qi SW, Chaudhry MT, Zhang Y et al (2007) Comparative proteomes of Corynebacterium glutamicum grown on aromatic compounds revealed novel proteins involved in aromatic degradation and a clear link between aromatic catabolism and gluconeogenesis via fructose-1,6-bisphosphatase. Proteomics 7:3775–3787
Seo JS, Keum YS, Li QX (2009) Bacterial degradation of aromatic compounds. Int J Environ Res Public Health 6:278–309
Shuttleworth KL, Cerniglia CE (1995) Environmental aspects of PAH biodegradation. Appl Biochem Biotechnol 54:291–302
Story SP, Kline EL, Hughes TA et al (2004) Degradation of aromatic hydrocarbons by Sphingomonas paucimobilis strain EPA505. Arch Environ Contam Toxicol 47:168–176
Talley JW, Ghosh U, Tucker SG et al (2002) Particle-scale understanding of the bioavailability of PAHs in sediment. Environ Sci Technol 36:477–483
Tang YJ, Shui W, Myers S, Feng X et al (2009) Central metabolism in Mycobacterium smegmatis during transition from O2-rich to O2-poor conditions as studied by isotopomer-assisted metabolite analysis. Biotechnol Lett 31:1233–1240
Tzin V, Galili G (2010) New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol Plant 3(6):956–972
Walter U, Beyer M, Klein J, Rehm HJ (1991) Degradation of pyrene by Rhodococcus sp. UW1. Appl Microbiol Biotechnol 34:671–676
Wang X, Sun S, Ma H, Liu Y (2006) Sources and distribution of aliphatic and polyaromatic hydrocarbons in sediments of Jiaozhou Bay, Qingdao, China. Mar Pollut Bull 52:129–138
Wayne LG, Lin KY (1982) Glyoxylate metabolism and adaptation of mycobacterium tuberculosis to survival under anaerobic conditions. Infect Immun 37(3):1042
Acknowledgments
This work was supported by the Development of Biohydrogen Production Technology using Hyperthermophilic Archaea program and the Marine and Extreme Genome Research Center program of the Ministry of Land, Transport, and Maritime Affairs, a KBSI grant (K32403) to Y.H.C., and the National Research Foundation of Korea Grant funded by the Korean Government (No. 2012-0009212 to Y.G.C).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10532_2013_9622_MOESM1_ESM.pdf
Online Resource 1: The gas chromatogram of the extracted pyrene residue at a 0 h (day 0) and b 48 h (day 2) from the pyrene-induced Mycobacterium gilvum PYR-GCK culture. Quantification was based on the integration of FID-peak areas of absolute pyrene identification and 2-nanodecanone was used as an injected internal reference standard. Supplementary material 1 (PDF 231 kb)
10532_2013_9622_MOESM2_ESM.pdf
Online Resource 2: Mass-fragment spectra of the degraded pyrene substrate showing metabolites identification of a phthalic acid and b 1,10-phenanthrene dicarboxylic acid in Mycobacterium gilvum PYR-GCK. Relative identification of pyrene metabolites were accomplished based on the library similarity search on the GC/MS software program as well as metabolites structures and fragment patterns from published papers. Supplementary material 2 (PDF 244 kb)
10532_2013_9622_MOESM3_ESM.pdf
Online Resource 3: List of all identified proteins in Mycobacterium gilvum PYR-GCK by Mascot database search. Gene locus tags are numbers assigned to each gene in the annotated genome sequence. Proteins were identified and normalized based on NSAF, PAF and emPAI (explained in “Materials and methods”). Significantly regulated proteins are based on quantification in at least 2 of 3 biological replicates, a mean regulation factor ratio of <−0.5 or ≥2 and a p value of ≤0.05 (in bold fonts). Parameters (ratio and p value) with glucose or pyrene written signify samples with binary expression in the corresponding substrate induced samples. All values are acquired from triplicate analyses. Supplementary material 3 (PDF 1625 kb)
Rights and permissions
About this article
Cite this article
Badejo, A.C., Choi, CW., Badejo, A.O. et al. A global proteome study of Mycobacterium gilvum PYR-GCK grown on pyrene and glucose reveals the activation of glyoxylate, shikimate and gluconeogenetic pathways through the central carbon metabolism highway. Biodegradation 24, 741–752 (2013). https://doi.org/10.1007/s10532-013-9622-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-013-9622-9