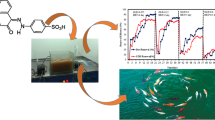
Abstract
The effect of the addition of compatible solutes (ectoine and trehalose) on the denitrification process of saline wastewater was studied. In saline wastewater, it was observed that the initial concentration of nitrates was 500 mg N l−1. A fatty substance isolated from oiled bleaching earth (waste of vegetable oil refining process) was used as a source of carbon. The consortium, which was responsible for the denitrification process originated from the wastewater of the vegetable oil industry. The consortium of microorganisms was identified by the use of restriction fragment length polymorphism of 16S rRNA gene amplicons and sequencing techniques. It was noted that ectoine affects significantly the activity of lipase and nitrate reductase, and resulted in faster denitrification compared to saline wastewater with the addition of trehalose or control saline wastewater (without compatible solutes). It was observed that relative enzyme activities of lipase and nitrate reductase increased by 32 and 35%, respectively, in the presence of 1 mM ectoine. This resulted in an increase in specific nitrate reduction rate in the presence of 1 mM ectoine to 5.7 mg N g−1 VSS h−1, which was higher than in the absence of ectoine (3.2 mg N g−1 VSS h−1). The addition of trehalose did not have an effect on nitrate removals. Moreover, it was found that trehalose was used up completely by bacteria as a source of carbon in the denitrification process. The fatty acids were biodegraded by 74% in the presence of 1 mM ectoine.
Similar content being viewed by others
Introduction
A common problem with wastewaters is that one of the components is usually predominant, which makes their utilization challenging. This can be illustrated by the example of saline wastewaters with a high content of nitrates originating from the production of explosive materials (Glass and Silverstein 1999), chemical fertilizers (Leakowić et al. 2000), regeneration of ion-exchange resins (Cyplik et al. 2010) and others. They can be characterized by a marginal content of phosphorus, microelements, and most importantly, a source of organic carbon, which is concurrently a source of energy for the denitrification (Cyplik et al. 2012). Moreover the high osmotic potential of brines usually limits the possibility of introducing microorganisms for biological treatment. The presence of sodium chloride and oxyanions at concentration above 2,000 mg/l create additional problems that hamper this process.
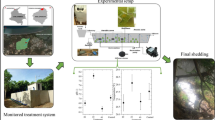
Therefore the denitrification process of brines requires the external supplementation with carbon source, which can be simply realized by combining wastes from vegetable oil production.
The fat industry represents one of the branches of the food industry that produces waste materials that are very difficult to biodegrade (Arvanitoyannis and Kassaveti 2007; Chandra and Sathiavelu 2009). Fatty waste products are difficult to utilize because of the large amounts of contaminants they contain that can subsequently contaminate water and soil. Among the various types of waste products from the vegetable oil industry, oiled bleaching earth creates the biggest problem for their utilization. It contains from 10 to 60% fatty substances, depending on the applied technology. In Poland, fat industry plants produce annually 40 kt of oiled bleaching earth, and it is assumed that this amount may increase. Previous studies on the biodegradation of biodiesel under anaerobic conditions (Cyplik et al. 2011) confirmed that fatty acid methyl and ethyl esters may be used as a carbon source, which does not alter the community structure and biodegradation potential. The application of fatty substances to enhance the denitrification efficiency of high salinity wastewater with a high content of nitrates seems like a plausible strategy to resolve the above-mentioned issues.
An important physiological mechanism of microorganisms that determines whether they can function in a particular habitat, is their ability to react to osmotic changes in the environment. The growth of microorganisms in an environment with low water activity is possible in part because of the cellular synthesis or intake of saccharides, alcohols and amino acids from the environment, which stabilize and protect cells from high salinity (Pastor et al. 2010). Compatible solutes contain a suitable composition of cytoplasm, and they increase the cell’s osmotic potential, and result in the maintenance of an appropriate turgor pressure. Compatible solutes stabilize the structure of proteins and nucleic acids (Argüelles 2000; Pastor et al. 2010). In addition, they also level off the differences between the osmotic pressure of the cell and the osmotic pressure of the environment (Roessler and Müller 2001).
Compatible solutes are amphiphilic molecules capable of increasing the total water content, as well as the cytoplasmic volume of cells, what might be responsible for reverse osmotic inhibition (Cayley et al. 1992). As proposed by Wiggins 1990, the structure-forming and breaking properties of compatible solutes may indirectly influence the hydration shells and thus the activities of the proteins involved.
Ectoine is a substance which is considered as a major protein stabilizer. Trehalose regulates the osmotic pressure both in the environment and inside the cell. It is accumulated in the periplasm and cytoplasm. The presence of trehalose in the cellular membranes decreases the temperature of phase transitions for lipids. This enables the liquid–crystal membrane structure to be maintained despite water deficiency or a notable drop of temperature (Leslie et al. 1995). Unlike ectoine, the synthesis of trehalose is dependant on the presence of nitrogen and compatible solutes in the environment.
Because the denitrification process was carried out in an environment with increased osmotic potential, the ability of bacteria to synthesize and accumulate compatible solutes was studied. The effect of compatible solutes on the denitrification process and the biodegradation of fatty acids was determined as well.
Experimental procedures
Composition of wastewater
The composition of synthetic wastewaters is presented in Table 1. In wastewaters, the initial concentration of nitrates was 500 mg N l−1 (pH 7.2). Fatty substances (1.5 g l−1) were used as a carbon source (C:N = 3).
Microorganisms and their genetic identification
A consortium of microorganisms isolated from a biological waste treatment plant was used in these studies. Identification of species included in this mixed population of microorganisms was achieved by the PCR–RFLP technique and sequencing. Total bacterial DNA was isolated using the Genomic Mini kit from A&A Biotechnology (Gdynia, Poland). A mixture of 16S rDNA was amplified on a matrix of genomic DNA. The PCR reaction was carried out in volume of 25 μl in a Biometra T Gradient apparatus. The reaction mixture contained 100 ng genomic DNA, 1 × reaction buffer (10 mM Tris–HCl, 50 mM KCl, 0.1% Triton X-100, 2.5 mM MgCl2), 200 μM dNTP, 0.44 μM of each primer oligonucleotide, 2 U DNA polymerase (DyNAzyme II, FinnZymes). The thermal cycling profile was: initial denaturation at 95°C for 5 min, hybridization at 48°C for 1 min, elongation at 72°C for 1 min, 30 cycles. The following oligonucleotide primers were used in the reaction: SD-B 5′AGAGTTTGATCCTGGCTCAGT3′ and SD-U 5′ACGGCTACCTTGTTACGACTT3′.
PCR amplicons were purified with the Clean-up kit (A&A Biotechnology), and cloned into the pGEM-T Easy vector (Promega, Madison, USA). Chemically competent cells of E. coli JM109 (Promega) were transformed with this vector. Transformants were cultured for 16 h at 37°C in LB (Luria–Bertani) medium with agar (Difco, USA) and ampicillin at a concentration of 100 μg ml−1. One hundred microliters of 100 mM isopropyl-β-d-thiogalactoside and 20 μl of 50 mg ml−1 bromo-chloro-indolyl-galactopyranoside (X-gal) were plated on Petri dishes. Plasmids were isolated with the use of the Plasmid Mini Kit (A&A Biotechnology). Reamplification of 16S rDNA was carried out on a matrix of plasmid DNA. PCR amplicons were digested with the restriction enzymes, HinfI and TaqI. Products of the digestion were separated by electrophoresis in a 2% agarose gel in 1× TBE buffer. DNA in the gel was visualized by the addition of ethidium bromide at 5 μg ml−1. Plasmid DNA that was not subjected to restriction digestion was sequenced (Genomed Warszawa, Poland). The results from sequencing were analyzed with the use of VectorNTI software, and BLASTN and Ribosomal Database Project databases.
Culture conditions
Loosely capped 250-ml Duran-Schott bottles contained 50 ml of wastewater. The initial inoculum was adjusted to an optical density of 0.10 ± 0.03 at 600 nm, and the bacteria were cultured at 25°C and 100 rpm. Limited aeration was achieved by flushing with nitrogen to remove dissolved oxygen from the medium and the headspace (Cyplik et al. 2011). All the experiments were carried out in three replicates.
Determination of nitrate reductase activity
The activity of nitrate reductase was determined based on measurements of the amount of nitrites produced from nitrates during the enzymatic reaction. Reaction was carried out at pH 7.2. Samples were incubated at 37°C for 15 min. A sample of 100 ml was taken from the culture and centrifuged at 4,000×g. Obtained biomass was suspended at 0.1 M phosphate buffer (pH 7.2) containing 1% NaCl, then centrifuged. This step was repeated twice. The biomass was condensed to the final volume 20 ml. Then, the cells were broken with the use of glass beads of 0.3 mm diameter for 30 min in a bead mill. After the cells were broken and separated from the glass beads, the entire sample was centrifuged at 5,000×g. A reaction mixture with following composition was prepared in a vial to a final volume of 3 ml: 1 ml KNO3 (100 mM), 0.5 ml Tris-HCI (100 mM), 0.5 ml methyl viologen (dichloride 1,1′–dimethyl-4,4′–bipyridine) (120 μM), 1 ml NaCl (1 M) and 50 μl crude extract. The reaction was initiated by the addition 0.1 ml sodium hydrogen sulfite (0.5 M). Samples were incubated for 15 min in a water bath at 37°C. Then, 0.1 ml formaldehyde was added to stop the reaction.
Determination of lipolytic activity
Lipolytic activity was determined based on the hydrolysis reaction of p-nitrophenyl phosphate (p-NPP) (Lotrakul and Dharmsthiti 1997). Enzymatic activity was calculated as the amount of μM p-NP released within 1 h, at 37°C from 1 ml of culture. The reaction mixture was composed of 540 μl of solution A (2 mM p-NPP dissolved in 2-propanol) and 4,860 μl of solution B (4 g Triton × 100 and 0.1 g Arabic gum in 100 ml phosphate buffer, pH 7.2). The mixture was emulsified for 10 min in an ultrasonic bath, and then heated for 5 min at 37°C in a water bath. After an initial warming of the mixture, 1 ml of culture was added. The entire mixture was incubated for 15 min at 37°C. The sample was centrifuged at 3,600×g for 15 min at 25°C.
Determination of nitrates and nitrites
The nitrate content of the brine was determined by spectrophotometry via reaction with sodium salicylate. The amount of produced nitrites in a reaction mixture, equal to the amount of reduced nitrates, was determined spectrophotometrically (λ = 530 nm) with sulfanilic acid and 1-naphthylamine. Volatile suspended solid (VSS) was determined using a standard method (Hermanowicz 1976).
Determination of trehalose, glycerol and ectoine
Compatible solutes from cells were isolated according to the methods described by Onraedt et al. (2004); Roustan and Sablayrolles (2002). Trehalose and glycerol were determined by HPLC (1200 series, Agilent Technologies, Santa Clara, CA). Analysis was performed isocratically at a flow rate 0.6 ml min−1, at 60°C on an Aminex HPX-87H 300 × 7.8 mm column (Bio-Rad, Hercules, CA). The mobile phase consisted of 0.001 N H2SO4. Thirty microliter samples were injected into the column.
Determinations of ectoine were carried out on the same HPLC apparatus. Analysis was performed isocratically at a flow rate of 2 ml min−1, at 70°C, on a Nucleosil 100-5 NH2 column (Mecherery-Nagel, Germany). The mobile phase was acetonitrile:water (70:30). Samples were filtered through 0.22 μm Millex-GS filters (Millipore, USA), and were injected in a volume of 30 μl.
Determinations of fatty acids
Fatty acids were extracted according to the Folch method (Folch et al. 1957). Chromatographic separation of methyl esters was carried out on a gas chromatograph (Hewlett Packard 5890, USA) with a flame ionization detector at 260°C, using “split/splitless” injection (1:50). A capillary column (Innowax 30 m × 0.25 mm × 0.25 μm) was used as a stationary phase and helium gas as a carrier gas (1.5 ml min−1). Chromatographic separation was achieved in a temperature ramp from 60 to 200°C at a rate of 12°C min−1, and the final temperature was maintained for 20 min. The injector temperature was 240°C. Identification of acids was based on retention times of internal standards (Supelco 37 Component Fame Mix, USA). However, quantitative interpretation was based on peak areas as compared to an internal standard–heptadecanoic acid methyl ester.
Kinetics of denitrification
During the initial experiments several calculations were carried out in order to determine the most suitable model. Based on the obtained data, it was concluded, that the zero order kinetic equation was most fit for further modeling (Foglar et al. 2005; Dhamole et al. 2007).
Results and discussion
The effect of the addition of compatible solutes on the activity of enzymes in denitrification and fat degradation
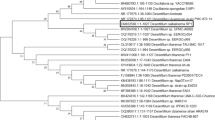
The consortium used to carry out the denitrification process was identified by restriction fragment length polymorphism of 16S rRNA gene amplicons and sequencing. It was found to consist of 12 dominant bacterial strains with the closest match to strains such as: Alacligenes faecalis, Pseudomonas alcaligenes, Pseudomonas putida, Klebsiella oxytoca, Stenotrophomonas maltophilia, Defluvibacter lusatiae, Hafnia alvei, Microbacterium foliorum, Microbacterium phyllosphaerae, Microbacterium profundi, Ochrobactrum intermedium, Sphingobacterium sp. All the isolated bacteria were capable of nitrate reduction. Alcaligenes faecalis, Pseudomonas alcaligenes, Pseudomonas putida, Klebsiella oxytoca, Stenotrophomonas maltophilia, Ochrobactrum intermedium, Sphingobacterium sp. are well known for the ability to completely reduce nitrate to elemental nitrogen. None of the described microorganisms could synthesize ectoine, however some Pseudomonas putida species were previously reported to be capable of betaine production ((Kets et al. 1996). Trehalose could be synthesized by Stenotrophomonas maltophilia (Roder et al. 2005).
The wastewater denitrification process was conducted after the addition of 1 mM compatible solutes (trehalose or ectoine) and a fatty substance (1.5 g l−1) isolated from oiled bleaching earth, and their effect on the rate of denitrification was determined. On the first day denitrification, the bacteria adapted to a new environment (lag phase) and no decrease in the nitrate level was observed. The nitrite concentration, which is an intermediate denitrification product, increased slightly in all cultures at that time, but did not exceed 8 mg N l−1. A considerable increase in the nitrate reduction rate was observed on the subsequent days. In particular, the addition of ectoine accelerated the process, and caused the complete removal of nitrates after 5 d of the process, and that of nitrites after 7 d. The addition of trehalose did not affect significantly the acceleration of the denitrification process; it proceeded as in wastewater without the addition of any compatible solutes. Although the nitrates were removed completely on the 7th day (on the 8th day in wastewater without compatible solutes), the complete removal of nitrite occurred in both cases on the 9th day of the process. However it is also plausible that the lag phase could be explained otherwise. The shape of the nitrate degradation curve could be the result of the microbial consortium acclimation to either salinity or nitrous acid. Additionally the base producing nature of denitrification processes increased the pH, making the solution more basic over the course of reaction. This increase in pH may have created an environment more conducive to denitrification, increasing the rate of the reaction after an initial lag. It has been shown by other researchers that denitrification occurs more rapidly at elevated pH values (Rust et al. 2000). In this conditions nitrite peaking may have presented some nitrous acid toxicity, although this seems unlikely. The pH was monitored during all experiments when cultures were terminated, so the phosphate buffer was strong enough to keep the pH in range 7.2–8.1 and from spectacular rising pH during denitrification.
In all cultures, the maximal nitrite concentration did not exceed 185 mg N l−1 (Fig. 1). These results confirm the calculated specific nitrate reduction rate, which was: 5.7 mg N g−1 VSS h−1 for denitrification in the presence of ectoine, 3.4 mg N g−1 VSS h−1 with trehalose, and 3.2 mg N g−1 VSS h−1 without any compatible solutes addition (Table 2). During the denitrification with a hydrophilic carbon source (methanol, acetic acid) in a sodium chlorate-free environment, considerably higher values of specific nitrate reduction rate, above 100 mg N g−1 VSS h−1 were obtained (Foglar et al. 2005; Rodríguez et al. 2007). The salinity of wastewater decreased the value of the specific nitrate reduction rate to 12–23 mg N g−1 VSS h−1 (Glass and Silverstein 1999; Peyton et al. 2001).
When fatty substances are being used as a carbon source, the key factors influencing denitrification are the activities of enzymes participating in nitrate reduction (nitrate reductase) and fat hydrolyzing enzymes (lipases). Nitrate reductase is an enzyme located in the bacterial cell membrane and determines the first stage of denitrification, that is reduction of nitrates to nitrites (Kim et al. 2006). Lipases are enzymes secreted outside the cell and are responsible for fat hydrolysis to glycerol and fatty acids. These enzymes have great catalytic potential, and require an insoluble substrate at the fat–water phase boundary (Gupta et al. 2004). They determine the supply of the organic carbon source (electron donor) essential for denitrification. It is of particular importance if high molecular weight compounds of hydrophobic character are used as a carbon source.
The measured activities of nitrate reductase and lipase confirm the important effect of ectoine on the denitrification process. At the beginning (first day), the activity of lipases did not exceed 3 μM p-nitrophenol ml−1 min−1. Subsequently, the activity increased and already on the second day reached 9 μM p-nitrophenol ml−1 min−1. The highest lipase activity was obtained on the fifth day (15.8 μM p-nitrophenol ml−1 min−1). This activity remained until the end of the denitrification process. Lipase activity was similar during denitrification with the addition of trehalose, and denitrification without compatible solutes. The enzyme activity also increased from the initial 3 μM p-nitrophenol ml−1 min−1 (first day) until the fourth day (lipase activity: 11–12 μM p-nitrophenol ml−1 min−1). However, these activities were lower by 32%, than during denitrification with ectoine (Fig. 2a). Changes in nitrate reductase were similar. Initially no activity of this enzyme was observed. It resulted from the fact that the bacteria used for denitrification were grown under aerobic conditions, and needed time for adaptation to new environmental conditions, including synthesis of cytochrome aa3, which transfers electrons onto nitrate in the respiratory chain. The activity of nitrate reductase during denitrification with ectoine addition increased until the fourth day of the process, reaching a maximum of 49 mg N ml−1 min−1. The increase in nitrate reductase activity in cultures with trehalose and without the addition of compatible solutes proceeded similarly to denitrification with ectoine addition, but the activity was lower by 35% (Fig. 2b). A similar phenomenon was observed by Werber and Mevarech (1978), but they measured the activity of reductases in the genus isolated in the Dead Sea. The authors stated that after transferring the Halobacteria from aerobic to anaerobic conditions, nitrate reductase gene expression occurred at 85 h of culturing, and its activity was 30 mg N min−1 g−1 protein. The enzyme activity increased rapidly, and at 200 h of culturing reached 400 mg N min−1 g−1 protein.
Changes in ectoine and trehalose concentration in wastewater were observed during the denitrification process (Fig. 3). Until the second day of the process, the concentration of both compounds in wastewater did not differ significantly and was 1 mM. After the second day, the concentration of trehalose began to decrease considerably, and on the fourth day the compound was no longer observed in wastewater. The ectoine concentration decreased on the 8th day to 0.1 mM. The level of ectoine in cell biomass was 0.16 mg per 1 g of dry mass, which is 80% of ectoine contained in wastewater at the end of the process. As the final ectoine concentration in wastewater was 0.1 mM, the amount of metabolized ectoine, used as carbon source was 10%. No trehalose was found in the bacterial biomass; thus, it must be acknowledged that this compound was fully used as a carbon source during the denitrification process. The fact that glycerol was produced during fat hydrolysis, and that glycerol is a substance included among compatible solutes, and at the same time a perfect carbon source for denitrification, is worth mentioning. Its small concentration in wastewater, below 0.003 mM, and in the cell biomass (below 0.06 mM) is evidence that this compound was used immediately by the bacteria for energetic processes and metabolized fully.
The results on the trehalose and ectoine content in the cell biomass indicate that denitrifying bacteria preferred amino acids (ectoine) as a compatible solute, while they used sugars (trehalose) only as a carbon source and metabolized them. It should be emphasized that the ability to accumulate and metabolize compatible solutes applies to numerous heterotrophic bacteria, cyanobacteria, phototrophic bacteria, methane bacteria, fungi and yeast (Welsh 2000), including bacteria that do not possess the ability to produce them (Jebbar et al. 1992; Nagata and Wang 2005). Studies of Zhang et al. (2008) can serve as an example. They used ectoine to increase the efficiency of ethanol fermentation conducted by the bacterium Zymomonas mobilis. It was observed that relative enzyme activities of glucokinase (GK), glucose-6-phosphate dehydrogenase (G-6-PDH), and alcohol dehydrogenase (ADH) increased by 29.9, 11.6, and 7.7%, respectively, in the presence of 0.5 mM ectoine. This resulted in an increase in volumetric ethanol productivity in the presence of 1 mM ectoine to 1.1 g l−1 h−1, which was higher than in the absence of ectoine (0.7 g l−1 h−1). Examples of the use of ectoine as a protective substance in enzymatic processes are known. According to Wang and Zhang (2010) to improve the production of biodiesel by enzymatic conversion of triglycerides in cottonseed oil, compatible solutes were added to the solvent-free methanolysis system to prevent competitive methanol inhibition of the immobilized lipase (Novozym® 435). The results indicated that the addition of ectoine increased biodiesel synthesis using a three-step methanol addition process. The concentration of methyl ester (ME) reached a maximum of 95% in the presence of 1.1 mM ectoine, which constituted an increase by 20.9% compared to that in the absence of ectoine. Bacteria used for denitrification did not have the ability to produce ectoine and trehalose, and the amounts that they accumulated from the environment were much smaller than in halotolerant or halophilic bacteria, which are capable of growing in a high salinity environment (from 3 to 18% NaCl) (Onraedt et al. 2004). Such high osmotic potentials force microorganisms to accumulate compatible solutes in the amounts directly proportional to the salinity of the environment.
Fatty acid biodegradation
Fatty acids with chains containing more than four carbon atoms have a limited solubility in water, and those with more than twelve carbon atoms are practically insoluble. Fatty acids form a monomolecular layer on the surface of water, with the carboxyl groups directed towards water (Ron and Rosenberg 2002). Fat degradation products, such as glycerol or fatty acids are, for many types of microorganisms, a perfect source of carbon and energy. Heterotrophic denitrifying bacteria need an organic carbon source to conduct life processes (growth, respiration). Carbon sources used in waste treatment technology are very diverse; for instance, methanol, ethanol, glucose, acetate, formic acid, and industrial wastes, including molasses, whey, wastewater, and even cellulose (Mateju et al. 1992). The use of a hydrophobic carbon source enabled the total removal of nitrate removal from wastewater. Fatty acids with the shortest chains were metabolized first during denitrification (Fig. 4). C12 and C14 fatty acids, at initial amounts of 36 and 29 mg l−1, respectively, were removed completely in all cultures already on the sixth day of the process. Bacteria also used acids of higher carbon atom numbers (C18 and C18:2). The initial concentration of these acids in wastewater was 70 and 75 mg l−1, respectively. On the sixth day of the process, a complete removal of these fatty acids from wastewater was observed. The fatty acids that prevailed in wastewater (C16 and C18:1) were biodegraded faster when ectoine was added. On the sixth day of the process, in the culture medium containing ectoine, the C16 acid concentration decreased from the initial 660 to 175 mg l−1, while the C18:1 acid concentration dropped from 625 to 245 mg l−1.
Conclusions
The presented results confirm a significant effect of ectoine on the denitrification rate. Ectoine increased lipase activity, thus the availability of organic compounds for the process increased, and influenced denitrification by increasing the nitrate reductase activity. No significant effect of trehalose on the denitrification process was observed in the study. It was confirmed that fatty acids constitute a perfect carbon source for the denitrification process. Therefore, it is possible to conduct an effective denitrification along with the simultaneous biodegradation of fatty acids isolated from oiled bleaching earth in an environment of high osmotic potential.
References
Argüelles JC (2000) Physiological roles of trehalose in bacteria and yeasts: a comparative analysis. Arch Microbiol 174:217–224
Arvanitoyannis IS, Kassaveti A (2007) Current and potential uses of composted olive oil waste. Int J Food Sci Technol 42:281–295
Cayley S, Lewis BA, Record MT Jr (1992) Origins of the osmoprotective properties of betaine and proline in Escherichia coli K-12. J Bacteriol 174:1586–1595
Chandra M, Sathiavelu S (2009) Waste management in the oil industry in the Mediterranean region by composting. Clean Technol Environ Policy 11:293–298
Cyplik P, Czaczyk K, Piotrowska-Cyplik A, Marecik R, Grajek W (2010) Removal of nitrates from brine using Haloferax mediterranei archeon. Environ Prot Eng 36:5–16
Cyplik P, Schmidt M, Szulc A, Marecik R, Lisiecki P, Heipieper HJ, Owsianiak M, Vainshtein M, Chrzanowski Ł (2011) Relative-quantitative PCR to assess bacterial community dynamics during degradation of diesel and biodiesel fuels under various aeration conditions. Bioresour Technol 102:4347–4352
Cyplik P, Marecik R, Piotrowska-Cyplik A, Olejnik A, Drożdżyńska A, Chrzanowski Ł (2012) Biological denitrification of high nitrate processing wastewaters from explosives production plant. Water Air Soil Pollut. doi:10.1007/s11270-011-0984-5
Dhamole PB, Nair RR, D’souza SF, Lele SS (2007) Denitrification of high strength nitrate waste. Bioresour Technol 98:247–252
Foglar L, Briŝki F, Sipos L, Vukovic M (2005) High nitrate removal from synthetic wastewater with the mixed bacterial culture. Bioresour Technol 96:879–888
Folch J, Lees M, Sloane-Stanley GH (1957) A simple method for isolation and purification of total lipids from animal tissues. J Biol Chem 226:497–509
Glass C, Silverstein J (1999) Denitrification of high-nitrate, high salinity wastewater. Water Res 33:223–229
Gupta R, Gupta N, Rathi P (2004) Bacterial lipases: an overview of production, purification and biochemical properties. Appl Microbiol Biotechnol 64:763–781
Hermanowicz W (1976) Physicochemical testing of water and sewage. Arkady, Poland
Jebbar M, Talibart R, Gloux K, Bernard T, Blanco C (1992) Osmoprotection of Escherichia coli by ectoine—uptake and accumulation characteristics. J Bacteriol 174:5027–5035
Kets EPW, de Bond JAM, Heipieper HJ (1996) Physiological response of Pseudomonas putida S12 subject to reduced water activity. FEMS Microbiol Lett 139:133–137
Kim SH, Song SH, Yoo YJ (2006) Characterization of membrane-bound of nitrate reductase from denitrifying bacteria Ochrobactrum antrophi SY 509. Biotechnol Bioprocess Eng 11:32–37
Leakowić S, Mijatović I, Serjan-Stefanović S, Hodzić E (2000) Nitrogen removal from fertilizer wastewater by ion exchange. Water Res 34:185–190
Leslie SB, Israeli E, Lighthart B, Crowe JH, Crowe LM (1995) Trehalose and sucrose protect both membranes and proteins in intact bacteria during drying. Appl Environ Microbiol 10:3592–3597
Lotrakul P, Dharmsthiti S (1997) Lipase production by Aeromonas sobria LP004 in a medium containing whey and soybean meal. World J Microbiol Biotechnol 13:163–166
Mateju V, Cizinska S, Krajci J, Janoch T (1992) Biological water denitrification-a review. Enzyme Microb Technol 14:170–183
Nagata S, Wang C (2005) Efficient utilization of ectoine by halophilic Brevibacterium species and Escherichia coli subjected to osmotic downshock. J Biosci Bioeng 99:61–67
Onraedt A, De Muynck C, Walcarius B, Soetaert W, Vandamme E (2004) Ectoine accumulation in Brevibacterium epidermis. Biotechnol Lett 26:1481–1485
Pastor JM, Salvador M, Argandoña M, Bernal V, Reina-Bueno M, Csonka LN, Iborra JL, Vargas C, Nieto JJ, Cánovas M (2010) Ectoines in cell stress protection: uses and biotechnological production. Biotechnol Adv 28:782–801
Peyton BM, Mormile MR, Petersen JN (2001) Nitrate reduction with Halomonas compisalis: kinetics of denitrification at pH 9 and 12,5% NaCl. Water Res 35:4237–4242
Roder A, Hoffmann E, Hagemann M, Berg G (2005) Synthesis of the compatible solutes glucosylglycerol and trehalose by salt-stressed cells of Stenotrophomonas strains. FEMS Microbiol Lett 243:219–226
Rodríguez L, Villaseñor J, Fernández FJ (2007) Use of agro-food wastewaters for the optimisation of the denitrification process. Water Sci Technol 55:63–70
Roessler M, Müller V (2001) Osmoadaptation in bacteria and archaea: common principles and differences. Environ Microbiol 3:743–754
Ron EZ, Rosenberg E (2002) Biosurfactants and oil bioremediation. Curr Opin Biotech 13:249–252
Roustan J-L, Sablayrolles J-M (2002) Trehalose and glycogen in wine-making yeasts: methodological aspects and variability. Biotechnol Lett 24:1059–1064
Rust CM, Aelion CM, Flora JRV (2000) Control of ph during denitrification in subsurface sediment microcosms using encapsulated phosphate buffer. Water Res 34:1447–1454
Wang Y, Zhang L (2010) Ectoine improves yield of biodiesel catalyzed by immobilized lipase. J Mol Catal B 62:90–95
Welsh DT (2000) Ecological significance of compatible solute accumulation by microorganisms: from single cells to global climate. FEMS Microbiol Rev 24:263–290
Werber MM, Mevarech M (1978) Induction of a dissimilatory reduction pathway of nitrate in Halobacterium of the Dead Sea. Arch Biochem Biophys 186:60–65
Wiggins PM (1990) Role of water in some biological processes. Microbiol Rev 54:432–449
Zhang L, Lang Y, Wang C, Nagata S (2008) Promoting effect of compatible solute ectoine on the ethanol fermentation by Zymomonas mobilis CICC10232. Process Biochem 43:642–646
Acknowledgment
This work was financially supported by the Ministry of Science and Higher Education, Poland (Grants No. N N523 419837 and N N523 176 333) years 2008–2012.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Cyplik, P., Piotrowska-Cyplik, A., Marecik, R. et al. Biological denitrification of brine: the effect of compatible solutes on enzyme activities and fatty acid degradation. Biodegradation 23, 663–672 (2012). https://doi.org/10.1007/s10532-012-9542-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-012-9542-0