Abstract
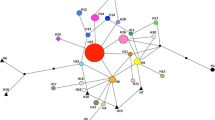
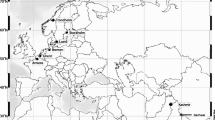
The shrub Rosa rugosa (Japanese Rose), native to East Asia, is considered one of the most troublesome invasive plant species in natural or semi-natural habitats of northern Europe and has proven very difficult to control. We aimed at disentangling the species’ invasion history in Europe, including determining the number of introductions and their geographic origin, and at investigating whether populations in the introduced and native ranges differ in genetic diversity, structure and degree of differentiation. We found that introduced (n = 16) and native (n = 16) populations had similar levels of genetic diversity at seven nuclear SSR (microsatellite) loci. European populations lack isolation by distance and are less genetically differentiated than are populations in East Asia. Multiple and at least three independent colonization events, one of which was particularly successful, gave rise to current R. rugosa populations in Europe. The geographic distribution patterns of these three genetic clusters could not be explained by natural dispersal alone, indicating that human mediated secondary dispersal is driving the expansion in Europe. One cluster representing three of the European populations was most likely derived from NW Japan, whereas the origin of the remaining thirteen populations could not clearly be resolved. The introduction and expansion in Europe occurred with no significant loss of genetic diversity. We conclude that high propagule pressure at the primary establishment phase is the most parsimonious explanation for this pattern. A potential for long distance seed dispersal, coastal habitat connectivity and an outcrossing breeding system are factors likely to have enabled populations of R. rugosa to avoid detrimental effects of genetic bottlenecks and will further increase the species’ range size and abundance in Europe. We recommend that human-mediated dispersal should be prevented in order to halt the continued expansion.





Similar content being viewed by others
References
Alexander JM, Poll M, Dietz H, Edwards PJ (2009) Contrasting patterns of genetic variation and structure in plant invasions of mountains. Divers Distrib 15(3):502–512
Allendorf FW, Lundquist LL (2003) Introduction: population biology, evolution, and control of invasive species. Conserv Biol 17(1):24–30
Besnard G, Henry P, Wille L, Cooke D, Chapuis E (2007) On the origin of the invasive Olives (Olea europaea L., Oleaceae). Heredity 99(6):608–619
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144(1):1–11. doi:10.1007/s00442-005-0070-z
Bruun HH (2005) Rosa rugosa Thunb. ex Murray. J Ecol 93(2):441–470
Bruun HH (2006) Prospects for biocontrol of invasive Rosa rugosa. Biocontrol 51(2):141–181
Chun YJ, Nason JD, Moloney KA (2009) Comparison of quantitative and molecular genetic variation of native versus invasive populations of purple loosestrife (Lythrum salicaria L., Lythraceae). Mol Ecol 18(14):3020–3035
Corander J, Waldmann P, Sillanpää MJ (2003) Bayesian analysis of genetic differentiation between populations. Genetics 163(1):367–374
Corander J, Marttinen P, Sirén J, Tang J (2008a) BAPS: Bayesian analysis of population structure—manual (version 5.2)
Corander J, Marttinen P, Sirén J, Tang J (2008b) Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinform 9:539–553
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144(4):2001–2014
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17(1):431–449
Dobson HEM, Danielson EM, van Wesep ID (1999) Pollen odor chemicals as modulators of Bumble Bee foraging on Rosa rugosa Thunb (Rosaceae). Plant Species Biol 14(2):153–166
Doorduin LJ, van den Hof K, Vrieling K, Joshi J (2010) The lack of genetic bottleneck in invasive Tansy Ragwort populations suggests multiple source populations. Basic Appl Ecol 11(3):244–250. doi:10.1016/j.baae.2009.12.007
Downie DA (2002) Locating the sources of an invasive pest, Grape Phylloxera, using a mitochondrial DNA gene genealogy. Mol Ecol 11(10):2013–2026
Esselink GD, Smulders MJM, Vosman B (2003) Identification of cut Rose (Rosa hybrida) and rootstock varieties using robust sequence tagged microsatellite site markers. Theor Appl Genet 106(2):277–286
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes—application to human mitochondrial-DNA restriction data. Genetics 131(2):479–491
Fér T, Vašák P, Vojta J, Marhold K (2007) Out of the Alps or Carpathians? Origin of Central European populations of Rosa pendulina. Preslia 79(4):367–376
Fitzpatrick BM, Fordyce JA, Niemiller ML, Reynolds RG (2012) What can DNA tell us about biological invasions? Biol Invasions 14(2):245–253. doi:10.1007/s10530-011-0064-1
Frankham R (2005) Invasion biology—resolving the genetic paradox in invasive species. Heredity 94(4):385
Garza JC, Williamson EG (2001a) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10(2):305–318
Garza JC, Williamson EG (2001b) Webpage of John Carlos Garza—population genetic software. http://swfsc.noaa.gov/textblock.aspx?Division=FED&id=3298. Accessed 14 Sep 2009
Gaudeul M, Giraud T, Kiss L, Shykoff JA (2011) Nuclear and chloroplast microsatellites show multiple introductions in the worldwide invasion history of Common Ragweed, Ambrosia artemisiifolia. Plos One 6(3):e17658. doi:17610.11371/journal.pone.0017658
Genton BJ, Shykoff JA, Giraud T (2005) High genetic diversity in French invasive populations of Common Ragweed, Ambrosia artemisiifolia, as a result of multiple sources of introduction. Mol Ecol 14(14):4275–4285
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86(6):485–486
Goudet J (1999) PCA-GEN for windows: a program performing a principal component analysis (PCA) on genetic data (version 1.2)
Goudet J (2001) FSTAT: a program to estimate and test gene diversities and fixation indices (version 2.9.3)
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41:95–98. doi:http://www.mbio.ncsu.edu/jwb/papers/1999Hall1.pdf
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos T Roy Soc B 351(1345):1291–1298
Hanfling B, Carvalho GR, Brandl R (2002) mt-DNA sequences and possible invasion pathways of the Chinese Mitten Crab. Mar Ecol-Prog Ser 238:307–310
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59(8):1633–1638
Hellemaa P (1998) The development of coastal dunes and their vegetation in Finland. Fennia 176:1–157
Heller R, Siegismund HR (2009) Relationship between three measures of genetic differentiation G ST, D EST and G ST′: how wrong have we been? Mol Ecol 18(10):2080–2083
Henry P, Le Lay G, Goudet J, Guisan A, Jahodová S, Besnard G (2009) Reduced genetic diversity, increased isolation and multiple introductions of invasive Giant Hogweed in the western Swiss Alps. Mol Ecol 18(13):2819–2831
Isermann M (2008a) Effects of Rosa rugosa invasion in different coastal dune. In: Tokarska-Guzik B, Brock JH, Brundu G, Child L, Daehler CC, Pysek P (eds) Plant invasions: human perception, ecological impacts and management. Backhuys Publishers, Leiden, pp 289–306
Isermann M (2008b) Expansion of Rosa rugosa and Hippophae rhamnoides in coastal grey dunes: effects at different spatial scales. Flora 203(4):273–280
Jaramillo P, Garcia C (2006) Protocolo ADN Calandrinia galapagosa. Charles Darwin Research Station, Galápagos [in Spanish]
Jørgensen RH, Kollmann J (2009) Invasion of coastal dunes by the alien shrub Rosa rugosa is associated with roads, tracks and houses. Flora 204(4):289–297
Jost L (2008) G ST and its relatives do not measure differentiation. Mol Ecol 17(18):4015–4026
Kolbe JJ, Glor RE, Schettino LRG, Lara AC, Larson A, Losos JB (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431(7005):177–181
Kollmann J, Brink-Jensen K, Frandsen SI, Hansen MK (2011) Uprooting and burial of invasive alien plants: a new tool in coastal restoration? Restor Ecol 19(3):371–378. doi:10.1111/j.1526-100X.2009.00569.x
Le Roux J, Wieczorek AM (2009) Molecular systematics and population genetics of biological invasions: towards a better understanding of invasive species management. Ann Appl Biol 154(1):1–17. doi:10.1111/j.1744-7348.2008.00280.x
Lockwood JL, Hoopes MF, Marchetti MP (2007) Invasion ecology. Blackwell, Oxford
Lombaert E, Guillemaud T, Cornuet JM, Malausa T, Facon B, Estoup A (2010) Bridgehead effect in the worldwide invasion of the biocontrol Harlequin Ladybird. PLoS ONE 5(3):e9743. doi:10.1371/Journal.Pone.0009743
Marrs RA, Sforza R, Hufbauer RA (2008a) Evidence for multiple introductions of Centaurea stoebe micranthos (Spotted Knapweed, Asteraceae) to North America. Mol Ecol 17(19):4197–4208
Marrs RA, Sforza R, Hufbauer RA (2008b) When invasion increases population genetic structure: a study with Centaurea diffusa. Biol Invasions 10(4):561–572
McFadyen REC (1998) Biological control of weeds. Annu Rev Entomol 43:369–393
Miller N, Estoup A, Toepfer S, Bourguet D, Lapchin L, Derridj S, Kim KS, Reynaud P, Furlan L, Guillemaud T (2005) Multiple transatlantic introductions of the western Corn Rootworm. Science 310(5750):992. doi:10.1126/science.1115871
Miura O (2007) Molecular genetic approaches to elucidate the ecological and evolutionary issues associated with biological invasions. Ecol Res 22(6):876–883
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29(1):1–10
NOBANIS (2008) Rosa rugosa (Rosaceae, Angiosperms). Available from: http://nobanis.org/speciesInfo.asp?taxaID=763. Accessed 15 Aug 2012
Novak SJ, Mack RN (2005) Genetic bottleneck in alien plant species: Influence of mating systems and introduction dynamics. In: Sax DF, Stachowicz JJ, Gaines SD (eds) Species Invasions: Insights into ecology, evolution and biogeography. Sinauer Associates, Inc. Publishers, Sunderland, pp 201–228
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13(5):1143–1155
Okada M, Ahmad R, Jasieniuk M (2007) Microsatellite variation points to local landscape plantings as sources of invasive pampas grass (Cortaderia selloana) in California. Mol Ecol 16(23):4956–4971
Østergaard J (1953) Floristiske meddelelser: Rosa rugosa. Botanisk Tidsskrift 50:103–107 [in Danish]
Oyant LHS, Crespel L, Rajapakse S, Zhang L, Foucher F (2008) Genetic linkage maps of Rose constructed with new microsatellite markers and locating QTL controlling flowering traits. Tree Genet Genomes 4(1):11–23
Paetkau D, Slade R, Burden M, Estoup A (2004) Genetic assignment methods for the direct, real-time estimation of migration rate: a simulation-based exploration of accuracy and power. Mol Ecol 13(1):55–65
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90(4):502–503
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95(6):536–539
Prentis PJ, Sigg DP, Raghu S, Dhileepan K, Pavasovic A, Lowe AJ (2009) Understanding invasion history: genetic structure and diversity of two globally invasive plants and implications for their management. Divers Distrib 15(5):822–830
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94(17):9197–9201
Rosenthal DM, Ramakrishnan AP, Cruzan MB (2008) Evidence for multiple sources of invasion and intraspecific hybridization in Brachypodium sylvaticum (Hudson) Beauv. in North America. Mol Ecol 17(21):4657–4669
Ross CA, Auge H, Durka W (2008) Genetic relationships among three native North-American Mahonia species, invasive Mahonia populations from Europe, and commercial cultivars. Plant Syst Evol 275(3–4):219–229
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of 3 noncoding regions of chloroplast DNA. Plant Mol Biol 17(5):1105–1109
Thiele J, Kollmann J, Andersen UR (2009) Ecological and socioeconomic correlates of plant invasions in Denmark: the utility of environmental assessment data. Ambio 38(2):89–94
Thiele J, Isermann M, Otte A, Kollmann J (2010) Competitive displacement or biotic resistance? Disentangling relationships between community diversity and invasion success of tall herbs and shrubs. J Veg Sci 21(2):213–220. doi:10.1111/j.1654-1103.2009.01139.x
Ueda Y, Takeshita D, Ando T (1996) Pollination in Rosa rugosa Thun. ex Murray. Acta Horticulturae 424:309–310
USDA (2009) Rosa rugosa. http://www.plants.usda.gov/java/nameSearch?keywordquery=rosa+rugosa&mode=sciname. Accessed 3 Oct 2009
Vogel V, Pedersen JS, Giraud T, Krieger MJB, Keller L (2010) The worldwide expansion of the Argentine Ant. Divers Distrib 16(1):170–186. doi:10.1111/j.1472-4642.2009.00630.x
Wang LY, Ikeda H, Liu TL, Wang YJ, Liu JQ (2009) Repeated range expansion and glacial endurance of Potentilla glabra (Rosaceae) in the Qinghai-Tibetan Plateau. J Integr Plant Biol 51(7):698–706
Ward S (2006) Genetic analysis of invasive plant populations at different spatial scales. Biol Invasions 8(3):541–552
Ward SM, Gaskin JF, Wilson LM (2008) Ecological genetics of plant invasion: what do we know? Invasive Plant Sci Manag 1(1):98–109. doi:10.1614/ipsm-07-022.1
Weber E (2003) Invasive plant species of the world: a reference guide to environmental weeds. CABI Publishing, Wallingford
Weidema I (2006) NOBANIS - invasive alien species fact sheet - Rosa rugosa. From: Online Database of the North European and Baltic Network on Invasive Alien Species - NOBANIS www.nobanis.org. Accessed 16 May 2012
Weidema I, Ravn HP, Vestergaard P, Johnsen I, Bruun HH (2007) Rynket rose (Rosa rugosa) i Danmark. Rapport fra workshop på Biologisk Institut, Københavns Universitet 5–6 September 2006. Biologisk Institut, Københavns Universitet, Skov- og Landskab, Københavns Universitet, samt Skov- og Naturstyrelsen [in Danish]
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6):1358–1370
Williamson-Natesan EG (2005) Comparison of methods for detecting bottlenecks from microsatellite loci. Conserv Genet 6(4):551–562
Wilson JRU, Dormontt EE, Prentis PJ, Lowe AJ, Richardson DM (2009) Something in the way you move: dispersal pathways affect invasion success. Trends Ecol Evol 24(3):136–144
Yan Z, Denneboom C, Hattendorf A, Dolstra O, Debener T, Stam P, Visser PB (2005) Construction of an integrated map of Rose with AFLP, SSR, PK, RGA, RFLP, SCAR and morphological markers. Theor Appl Genet 110(4):766–777
Zhang YP, Uyemoto JK, Kirkpatrick BC (1998) A small-scale procedure for extracting nucleic acids from woody plants infected with various phytopathogens for PCR assay. J Virol Methods 71(1):45–50
Zhang LH, Byrne DH, Ballard RE, Rajapakse S (2006) Microsatellite marker development in Rose and its application in tetraploid mapping. J Am Soc Hortic Sci 131(3):380–387
Acknowledgments
We cordially thank Jean-Philippe Bizoux, Dirk-Jan ten Brink, Ulla Carlsson-Granér, Maarten Christenhusz, Charles Coyle, Tinne Gaardmann, Barbara Giles, Marcin Górniak, Jin-Seok Kim, Hajime Matsushima, Teruyoshi Nagamitsu, Liv S. Nilsen, Triin and Mari Reitalu, Olav Skarpaas, Sonia Vanderhoeven, Valentina Vetrova, Volker Wissemann and Valentin V. Yakubov for providing leaf material or DNA samples. Sylvia Mathiasen and Ruth Jacobsen are thanked for their assistance in the laboratory and Bjørn Hermansen for GIS support. Katharine Marske, Jaco Le Roux and two anonymous reviewers gave valuable comments on a previous version of this manuscript. This study was supported by FORMAS (the Swedish Research Council for Environment, Agricultural Sciences and Spatial Planning) and the Danish National Research Foundation. All authors thank the Danish National Research Foundation for support to the Center for Macroecology, Evolution and Climate and to the Centre for Social Evolution.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kelager, A., Pedersen, J.S. & Bruun, H.H. Multiple introductions and no loss of genetic diversity: invasion history of Japanese Rose, Rosa rugosa, in Europe. Biol Invasions 15, 1125–1141 (2013). https://doi.org/10.1007/s10530-012-0356-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-012-0356-0