Abstract
The objective of this study was to determine the probable pathogen causing the recently discovered hepatopancreas browned disease (HPBD) in Chinese mitten crab (Eriocheir sinensis), an important freshwater aquaculture species. Hepatopancreas material was collected from 30 E. sinensis individuals confirmed as infected using plasma coagulase tests. Genes nuc and femA, both specific for Staphylococcus aureus, were amplified by reverse transcription-PCR from three randomly selected HPBD-infected crabs, specific bands were observed at 191 and 397 bp, and amplified partial sequences aligned perfectly (100%) with published nuc and femA gene sequences using BLAST and GenBank. Four major developmental stages of hepatopancreas histopathological changes were observed in 7-week reverse infection challenge tests in 90 pathogen-induced animals; hepatopancreas material changed colour from orange to yellow, red to brown, and light brown to almost fully necrotised. Optical and transmission electron microscopy revealed similar hepatopancreas pathological characteristics to those of HPBD-infected E. sinensis. The hepatopancreas browning rate of infected crabs (88.89 ± 5.09%) was significantly higher than controls (7.78 ± 1.92%). These results suggest S. aureus is the probable pathogen causing HPBD in E. sinensis, and infection leads to inflammation, atrophy, and complication of hepatopancreatic necrosis.
Similar content being viewed by others
Introduction
Eriocheir sinensis is the most important freshwater crab in China due to its wide culture area (> 10,000 km2), high annual output (> 700,000 tons), high annual production value (> 8 billion US dollars), and nice delicacy (Fisheries Bureau, Ministry of Agriculture 2016). In Jiangsu Province, a major E. sinensis aquaculture area in China, we showed that the hepatopancreas of healthy pond-raised crabs is orange-yellow, but it can be red-brown or brown in crabs suffering from hepatopancreas browned disease (HPBD), which is the first named disease by our research team all over the world. Symptoms can occur at any stage of the E. sinensis life cycle and include massive effusion, hepatopancreas colour changes, obvious inflammation and ulceration, atrophy and necrosis of the hepatopancreas, and a tarnished carapace. In recent ten years, a lot of cases about hepatopancreas necrosis has spread rapidly with extensive E. sinensis farming, leading to heavy economic losses (Song et al. 2007a; b; c; Chang et al. 2008; Zhang and Pan 2013). Once the hepatopancreas of E. sinensis becomes necrotic, the edible value is lost since it is the most prized organ in terms of flavour (Chang et al. 2008).
Staphylococcus aureus, a common Gram-positive pathogenic bacterium, is widely distributed in air, water, and dust. S. aureus produces numerous toxins and invasive enzymes, including haemotoxin, leukocidin, enterotoxin, exfoliative toxin, toxic shock syndrome toxin, and plasma-coagulase, all of which work to destroy the immune system of the host (Morandi et al. 2009; Vandenesch et al. 2012). S. aureus causes skin suppurative infection, septicaemia, pneumonia, endocarditis, enteritis, meningitis, and toxic shock syndrome in humans and other animals (Emmerson 1994; Liu et al. 2014). Many of the symptoms are a direct result of the actions of its toxins and invasive enzymes. According to the US Center for Disease Control and Prevention, S. aureus infections are the second most common type of bacterial infection worldwide, second only behind Escherichia coli (Purcell and Fergie 2006). Much intensive histopathological research on S. aureus infections has focused on human and animal models (Conaughty et al. 2006; Salgado et al. 2005; Wang et al. 2010). Meanwhile, various aquatic animals have been suffered from S. aureus infections (Suo et al. 2008; Wang et al. 2014; Wu and Su 2014).
In the present study, we performed a microbiological examination of HPBD-infected E. sinensis individuals to investigate histopathological changes of the hepatopancreas and evaluate inflammatory responses with a reverse infection challenge test. The primary objective was to elucidate the probable pathogen responsible for HPBD.
Materials and methods
Experimental crabs
During June 2016, 30 HPBD-infected E. sinensis individuals (mean weight 24.65 ± 3.86 g) and 180 healthy crabs of a similar size (mean weight 25.38 ± 0.72 g) were selected from a Chinese mitten crab farm at the Freshwater Fisheries Research Institute of Jiangsu Province, CN. All healthy E. sinensis individuals were vigorous, had a bright carapace, intact appendages, and an orange-yellow hepatopancreas, as observed by gently opening the abdomen umbilicus connected to the abdomen carapace.
Strain culture
Hepatopancreas material was removed from each of 30 HPBD-infected and 30 healthy E. sinensis individuals, immediately inoculated into corresponding Luria-Bertani (LB) liquid culture medium (5 mL), and placed in an incubator shaker (SPX-250-ZX, Xinbiao Tengda Instrument Equipment, Beijing, CN) for 24 h at 37 °C. Rejuvenated bacterial solutions were obtained by centrifuging for 5 min at 6000g (Eppendorf 5804R, Hamburg, Germany).
Plasma coagulase test (PCT, test tube method)
For the experimental group, 0.5 mL of freeze-dried rabbit plasma (Hope Bio-Technology, Qingdao, CN) was added to each of the 60 aforementioned bacterial solutions (0.3 mL) in a test tube (5 mL). For the positive control group, freeze-dried rabbit plasma (0.5 mL) was added to the S. aureus standard strain (ATCC29213, Maojie Microorganism, Nanjing, CN) in a test tube (5 mL). For the negative control group, freeze-dried rabbit plasma (0.5 mL) was added to LB liquid culture medium (0.3 mL) in a test tube (5 mL). All samples were placed in an incubator shaker (Peiying Experimental Equipment, Suzhou, CN) at 37 °C and observed for at least 6 h (once every 0.5 h). Solidification in the test tube represented a positive PCT result as follows:
Positive PCT rate = (number of positive reactions/30) × 100%
RNA extraction, purity, and concentration determination
Three solutions were randomly sampled from 30 rejuvenated bacterial solutions. To 0.3 mL of each solution in an Eppendorf centrifuge tube (1.5 mL), 1.0 mL of ice-cold TRIzol (Invitrogen, Carlsbad, USA) was added, and mixtures were incubated at room temperature for 5 min. Chloroform (0.2 mL; Nanjing Chemical Reagent, Jiangsu, CN) was then added, vortexed for 15 s, and samples incubated for 3 min at room temperature. After centrifugation (Eppendorf centrifuge 5804R) at 12,000g for 15 min at 4 °C, the resulting supernatant (0.6 mL) was placed in another Eppendorf centrifuge tube (1.5 mL), isopropyl alcohol (0.6 mL; Nanjing Chemical Reagent) was added, mixed gently for 15 s, and samples incubated for 10 min at room temperature. The supernatant was carefully removed after centrifugation at 12,000g for 10 min at 4 °C, and ethyl alcohol (1.0 mL, 75%, prepared by double distilled water processed by diethyl pyrocarbonate; Jiancheng Bioengineering Institute, Nanjing, CN) was slowly added along the tube wall. The supernatant was then carefully removed after centrifugation at 12,000g for 10 min at 4 °C, the precipitate was air-dried for 5 min at room temperature, RNAse-free double-distilled water (30 μL) was added to dissolve RNA, and samples were stored at – 70 °C.
RNA samples were diluted 100-fold with 1× TRIS-EDTA buffer. The optical density (OD) was determined at 260 and 280 nm using a spectrophotometer (Shimadzu UV-2450, Japan), and an OD260/280 value of 1.96 indicated high purity (1.8-2.1 indicates high purity). RNA concentration (OD260 × 100 × 0.04 μg/μL = 0.4 μg/μL) was calculated.
First strand cDNA synthesis (25 μL reaction system)
Before first strand cDNA synthesis, three centrifuge tubes filled with RNA were gently mixed and centrifuged at 800g for 20 s at 4 °C. First strand cDNA synthesis then comprised the following steps: to three RNAse-free centrifuge tubes (0.2 mL, AXYGEN, USA) which had been sterilised beforehand was successively added 5 μL RNA (2 μg), 2 μL Oligo dT(18) (10 μM), and 8 μL double-distilled water. Samples were incubated for 5 min at 75 °C then at 0 °C for 5 min in an ice bath. Next, 0.5 μL RNase inhibitor (40 U/μL), 2.5 μL 10× AMV reaction buffer, 1.5 μL dNTPs (10 mM), 1.0 μL DTT (200 mM), 1.0 μL reverse transcriptase (AMV), and 3.5 μL double-distilled water were successively added, mixed, centrifuged at 800g for 20 s at 4 °C, and incubated for 60 min at 42 °C, then for 10 min at 85 °C, then for 5 min at 0 °C in an ice bath to prepare products for the next PCR step. All the above experimental reagents for cDNA synthesis and the associated kits were purchased from Jiancheng Bioengineering Institute.
Primer design and PCR
The S. aureus-specific genes Nuc and femA encode a thermostable nuclease (Mehrotra et al. 2000) and a methicillin-resistant protein (Pinto et al. 2005), respectively. The nuc gene is highly conserved in different strains (Gao et al. 2003). Nowadays, S. aureus can be accurately and rapidly identified by PCR amplification of specific genes (Ardic et al. 2005). Specific primers for amplification of nuc and femA genes (Table 1) were designed using Primer Premier 5.0 according to gene sequences published in GenBank (ACC. No. FJ809757 and ACC. No. JN258593), and synthesised by a commercial corporation (Jinsirui Biology, Nanjing, CN). To each 0.2 mL centrifuge tube on ice, 5 μL 10× Taq buffer, 4 μL MgCl2 (25 mM), 1 μL dNTPs (10 mM), 1 μL primer-F (10 μM), 1 μL primer-R (10 μM), 1 μL cDNA, 0.5 μL Taq DNA polymerase (Fermentas EP0405, LT), and 36.5 μL sterilised double-distilled water was added. All amplifications were performed with a 5 min denaturation step at 95 °C, followed by 32 cycles of denaturation at 95 °C for 30 s, annealing at 56 °C for 30 s, extension at 72 °C for 45 s, and a final extension at 72 °C for 10 min, using a multigene gradient PCR amplifier (Labnet TC9600-G, USA). Amplified products were separated using a 1.5% agarose gel (Biowest 111860, ES) followed by UV visualisation (BIO-RAD Gel Doc XR, USA) after ethidium bromide staining.
Gene sequencing and BLAST alignment
All amplified products were sequenced by Sangon Biotech, Shanghai, CN. Sequence alignments were performed using BLAST and GenBank (National Center for Biotechnology Information, US National Library of Medicine).
Reverse infection challenge test
LD50 test
Pilot trial
Rejuvenated bacterial solutions from HPBD-infected E. sinensis individuals were transferred to normal saline (0.7%) at a cell density of 2.0 × 1010 CFU/mL. The exposure doses of pathogen were diluted in geometric progression at an equal ratio of 2 including the following: (1) 2.0 × 1010, (2) 1.0 × 1010, (3) 5.0 × 109, (4) 2.5 × 109, (5) 1.25 × 109, (6) 6.25 × 108, (7) 3.13 × 108 CFU/mL. The seven treatments were randomly assigned to aquaria. Five healthy E. sinensis individuals of similar size (mean weight 25.37 ± 0.63 g/crab) were stocked into each of seven aquaria. All the crabs were immersed in their assigned bacterial solutions as the natural infection route. The proximate maximal lethal dose (Dm) was determinated in a treatment, where the crabs were all died within 96 h.
Formal trial
According to the modified Karber method, the Dm, the highest dose, was diluted in geometric progression at an equal ratio of 3 including the following: (1) Dm, (2) Dm/3, (3) Dm/9, (4) Dm/27, (5) Dm/81, (6) Dm/243, (7) Dm/729 CFU/mL. The seven treatments were randomly assigned to an aquarium. Ten crabs of similar size (mean weight 25.18 ± 0.55 g/crab) were stocked into each of seven aquaria. The same infection pattern with the pilot trial was performed to all the crabs. The mortality in each treatment was recorded in 96 h. LD50 of pathogen-induced E. sinensis was calculated according to the following formulas.
LD50a = lg−1 [Xm−i (∑P-0.5)]
Sx50b = i[(∑P−∑P2)/(n-1)]1/2
95% confidence interval of LD50c = lg−1(x50 ± 1.96 Sx50)
95% mean confidence interval of LD50 = LD50 ± (MaxLD50−MinLD50)/2
aXm, lg (Dm); i, lg a–lg b, a > b, adjacent doses; P, mortality rate of each treatment; ∑P, sum of mortality rate
bSx50, standard error of lg LD50; n, initial number of crab; ∑P2, quadratic sum of mortality rate
cx50, lg LD50
Chronic infection
A total of 90 healthy E. sinensis individuals (mean weight 24.87 ± 0.77 g/crab) were immersed in the bacterial solutions (LD50/729 CFU/mL) to generate the reverse infection challenge test group. An additional 90 healthy crabs (mean weight 25.06 ± 0.65 g/crab) were immersed in the same way but with sterile saline to generate the control group. The two treatments were randomly assigned to triplicate aquaria (1.0 × 0.6 × 0.5 m), with 30 crabs in each of the six separate aquaria, and fed once daily a homemade diet of sterilised surimi at a rate of 10% of body weight. The trial was conducted in a closed laboratory that was sterilised beforehand. In all six aquaria, the assigned solutions were changed every 5 days, and the conditions were as follows: depth 0.2 m, temperature 26.5–28.2 °C, dissolved oxygen 6.2–6.7 mg/L, pH 6.9–7.6.
Hepatosomatic index and histopathological changes
Every seven days after the chronic infection, three crabs were randomly sampled from each of the six aquaria (nine per treatment), and the hepatosomatic index (HSI) (hepatopancreas weight/body weight × 100%) was calculated. At the same time, a little hepatopancreas tissues were removed and cleaned with phosphate-buffered solution (0.1 M, pH 7.4; Senbeijia Biology, Nanjing, CN). Samples were immediately placed in glutaraldehyde stationary liquid (2.5%, precooled to 4 °C; Senbeijia Biology), cut into small pieces (1 mm3), transferred to Eppendorf centrifuge tubes (1.5 mL) filled with glutaraldehyde stationary liquid, and stored in a refrigerator at 4 °C. After a series of treatments including sampling, cleaning (phosphate-buffered solution; Senbeijia Biology), fixation (osmic acid; Pelco, USA), stepwise dehydration in acetone (Nanjing Chemical Reagent), embedding in Epon resin (Spi-Chem, USA), polymerisation in a baking oven at 60°C (Biobase, Jinan, CN), ultrathin sectioning (Leica EMUC6, Wetzlar, DE), drying at 70 °C on a Leica electric heating plate, and staining using toluidine blue (Senbeijia Biology), samples were visualised using an optical microscope (OM, Olympus-BX51, JP) and images were captured using an imaging system (Nikon digital sight DS-L1, JP) to reveal histopathological changes of the hepatopancreas. Similarly, after a series of treatments including sampling, cleaning (phosphate-buffered solution), stepwise dehydration using ethyl alcohol and acetone (Nanjing Chemical Reagent), embedding using acetone and Spurr resin, solidifying using a baking oven at 37, 45, and 60 °C), ultrathin sectioning (Leica EMUC6), and staining (3% uranyl acetate and lead citrate, Spi-Chem), a transmission electron microscope (TEM, JEM1230, Japan Electron Optics Laboratory) was used to capture images of histopathological changes of the hepatopancreas ultrastructure.
Hepatopancreas browned rate and statistical analysis
At the end of the trial, all remaining crabs in each of the six aquaria were dissected to assess the hepatopancreas condition with naked eye and OM, and the hepatopancreas browned rate (HPBR) was calculated based on both dead and dissected crabs. SPSS 19.0 software was used to compare means and for variance analysis, and a p value < 0.01 was considered statistically significant.
Results
PCT analysis
The results of PCT analysis showed that all 30 HPBD-infected E. sinensis individuals gave a positive reaction, all 30 healthy E. sinensis individuals gave a negative reaction (Table 2).
PCR amplification and BLAST alignments of nuc and femA genes
PCR amplification electrophoretograms of nuc and femA genes from three randomly sampled HPBD-infected E. sinensis individuals showed that all yielded specific bands at 191 bp (Fig. 1) and 397 bp (Fig. 2).
PCR-amplified nuc gene partial sequences aligned perfectly (100%) with S. aureus strain ATCC 6538 and others using BLAST and GenBank. Similarly, PCR-amplified femA gene partial sequences aligned perfectly (100%) with S. aureus strain USA300-SUR1-24 and others using BLAST and GenBank.
LD50
The results of LD50 test of pathogen for E. sinensis using modified Karber method are showed in Table 3.
HSI
The HSI of pathogen-induced E. sinensis displayed a decreasing trend over time, while no significant changes were found in the control group (Table 4).
Histopathological changes of hepatopancreas
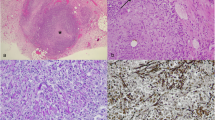
The microstructure of the hepatopancreas of healthy and HPBD-infected E. sinensis individuals was observed with the naked eye, OM, and TEM (Fig. 3). Healthy animals displayed an orange-yellow hepatopancreas, regular monolayer column-like epithelia, and intact ultrastructure (Fig. 3a, b, c). By contrast, HPBD-infected crabs exhibited a red-brown hepatopancreas with evidence of atrophy, epithelium apoptosis, hepatic tubule cavity enlargement, and evident ultrastructure damage (Fig. 3d, e, f).
Histopathological changes in the hepatopancreas microstructure of healthy and HPBD-infected E. sinensis individuals observed with the naked eye, optical microscopy (OM), and transmission electron microscopy (TEM). An orange-yellow hepatopancreas (white arrow) is evident in healthy E. sinensis (a). A large number of intact and regular monolayer column-like epithelia constitute healthy hepatic tubule sections. (★) represents OM (× 400 magnification, toluidine blue staining) (b). The hepatopancreas in healthy E. sinensis has numerous reticular hepatic tubules, intact intensely-dyed big cell nuclei (▽) and endoplasmic reticulum with an ordered structure (※) via TEM (× 10,000 magnification, uranyl acetate, and lead citrate staining) (c). The hepatopancreas is red-brown and atrophic (black arrow) in HPBD-infected E. sinensis (d). Hepatic tubule sections in HPBD-infected E. sinensis are filled with a large number of microorganisms (*), as shown by OM (× 400 magnification), and apoptotic column-like epithelia leading to the hepatic tubule cavity are enlarged (☆) (e). Cell nuclei in HPBD-infected E. sinensis are pyknotic, show evidence of karyorrhexis and karyolysis, and the endoplasmic reticulum has a disordered structure (※) based on TEM (× 10,000 magnification) (f). Epithelia are occupied by a large number of progenitive microorganisms (*) and damage to the cell structure is extensive (f)
Histopathological changes in the hepatopancreas of pathogen-induced E. sinensis were observed with the naked eye, and with OM and TEM (Fig. 4). As shown in Fig. 4, four major developmental stages were apparent (stages 1-4) in which the colour of the hepatopancreas is progressively browner (darker), turning from orange-yellow, red-brown, light-brown, to almost complete necrosis. Stage 1 (Fig. 4a, b, c) corresponds to the infection stage, and although pathogens begin to reproduce in the hepatic tubule, there were no significant pathological characteristics in terms of appearance or behaviour at this stage. Stage 2 (Fig. 4d, e, f) corresponds to the outbreak stage, and large numbers of pathogens occupied almost the whole hepatic tubule section. Additionally, the hepatopancreas changed colour from orange-yellow to red-brown at this stage, and hepatopancreas atrophy and ultrastructure changes were also evident. Stage 3 (Fig. 4g, h, i) corresponds to the apoptosis stage, during which hepatopancreas atrophy was increased, and the colour had changed to light-brown. A large number of pathogen cells had disappeared due to epithelium apoptosis, and the hepatic tubule was hollow and degenerated at this stage. Stage 4 (Fig. 4j, k, l) corresponds to the necrosis stage, during which extreme atrophy of the hepatopancreas was apparent, and it was light-brown in colour with massive effusion. Hepatic tubule dissociation and a shapeless ultrastructure suggested that necrosis of the hepatopancreas was advanced at this stage.
Histopathological changes in the hepatopancreas of pathogen-induced E. sinensis observed with the naked eye, OM, and TEM, revealing the four major developmental stages. a–c Stage 1; the hepatopancreas (white arrow) is orange-yellow under normal conditions (a). Only part of the hepatic tubule section (☆) is infected by pathogens (black arrow), and the boundaries of the column-like epithelia are clear (b). Intact, intensely-dyed big cell nuclei (▽) and abundant endoplasmic reticulum (※) are observed (c). Some hollow areas surrounding pathogens (black arrow) begin to form (c). d–f Stage 2; the hepatopancreas (white arrow) changes colour from orange-yellow to red-brown (d). Extensive replication of pathogens (black arrow) is evident in almost the entire hepatic tubule section (☆) (e). Reduction in endoplasmic reticulum (※) and cell nuclei (▽) pyknosis (f). Hollow areas surrounding pathogens (black arrow) are more abundant and larger (f). g–i Stage 3; the hepatopancreas (white arrow) changes colour from red-brown to light-brown, and significant atrophy is evident (g). The cavity (◎) in the hepatic tubule section is enlarged (☆) (h). A large number of pathogen cells (black arrow) have disappeared following the appearance of apoptotic column-like epithelia (h), and some pyknotic and dead pathogen cells (black arrow) are observed (i). The tubular structure of the endoplasmic reticulum (※) is seriously disordered, and the cell nucleus (▽) displays further pyknotic symptoms (i). j–l Stage 4; the hepatopancreas (white arrow) is lighter in colour, an atrophy and necrosis are pronounced and accompanied by a massive effusion in the body (j). The hepatic tubule section (☆) has disintegrated into several pieces, and some dead pathogen cells remain (black arrow) (k). Almost no organelles are visible except gathered cell nuclei (▽), which have undergone extensive karyorrhexis and karyolysis (l). The entire cytoplasm (&) is disordered (l). a, d, g, and j = observed with the naked eye. b, e, h, and k = observed by OM and toluidine blue staining at × 200 magnification. c, f, i, and l = observed by TEM at × 5000 magnification following uranyl acetate and lead citrate staining
HPBR
The HPBR of pathogen-induced E. sinensis was 88.89 ± 5.09%, which was significantly higher than that if the control group (7.78 ± 1.92%; Fig. 5).
Discussion
This is the first work to characterise and name HPBD, a new disease leading to hepatopancreatic necrosis in pond-raised Chinese mitten crab. Microbiological examination of HPBD-infected E. sinensis and reverse infection challenge tests were performed to identify the probable pathogenic factor responsible for the disease. The hepatopancreas of all 30 HPBD-infected E. sinensis individuals displayed a positive reaction in PCT analysis, and in all three randomly selected samples, nuc and femA genes specific for S. aureus were detected by RT-PCR and BLAST alignment of the amplified products. The same pathological characteristics were observed for hepatopancreas tissue from pathogen-induced and HPBD-infected E. sinensis. Together, these results suggest that S. aureus extremely likely to be the pathogenic factor responsible for HPBD in E. sinensis.
Due to the invasiveness of standard S.aureus produces, the test tube method is widely used to identify pathogens rapidly (GB 4789.10 2010). The precision rate for identifying S. aureus using PCT was 97.4%, which compares favourably with the rate obtained using an automatic microorganism identification instrument (BioMérieux-VITEK-2, Lyons, FR) (Peng et al. 2013). Two kinds of freeze-dried rabbit plasma have been used for PCT analysis, and the precision rate was 100% for identification of S. aureus with a MID-Staph Staphylococcus biochemistry identification system (Microgen, UK) (Shi et al. 2012). In agreement, S. aureus was identified by PCT analysis using freeze-dried rabbit plasma from all 30 HPBD-infected E. sinensis in the present study. This lays the groundwork for subsequent molecular biology detection. S. aureus can be accurately identified via PCR amplification of the nuc or femA genes using a broad range of samples. In the current work, both genes were successfully amplified, indicating the presence of S. aureus in all HPBD-infected E. sinensis individuals.
Four major developmental stages of hepatopancreas histopathological changes with decreasing HSI in pathogen-induced E. sinensis were observed, suggesting an increasing hepatopancreatic atrophy and pathogen invades and damages the column-like epithelia of the hepatopancreas in this crab species. Bacterial diseases are common in aquatic animals, examples of which include bacterial septicaemia in freshwater fish caused by Aeromonas hydrophila (Zhou et al. 2011), red-skin disease in freshwater fish caused by Pseudomonas fluorescens (Deng et al. 2010), Edwardsiella tarda infection of turbot and other fish (Feng et al. 2009), early mortality syndrome, also known as acute hepatopancreas necrosis syndrome (AHPND) in Penaeus vannamei caused by Vibrio parahaemolyticus (Tran et al. 2013), and gill-rot diseases of silver carp and other freshwater fish caused by Klebsiella pneumoniae (Tang et al. 2007). Most pathogenic bacteria previously shown to infect aquatic organisms are Gram-negative. In the present work, HPBD in E. sinensis was also shown to be caused by a Gram-positive bacterial pathogen. Based on similar hepatopancreatic necrosis in crustaceans, HPBD in E. sinensis appears to be similar to AHPND in Penaeus vannamei. The colour of the hepatopancreas turned from deep-brown, white and brown, to almost white in AHPND-infected Penaeus vannamei, and histological sections of the hepatopancreas revealed lesions in the acute phase of AHPND infection characterised by extensive sloughing and necrosis of the tubular epithelial cells in a reverse gavage challenge test (Tran et al. 2013). Additionally, the advanced stage of AHPND infection was characterised by hemocytic inflammation, bacterial infection, and massive sloughing and necrosis in immersion challenge tests (Tran et al. 2013).
The colour of the hepatopancreas of seven normal saline-induced E. sinensis individuals in the control group (90 samples) also changed from orange-yellow to red-brown, suggesting HPBD in E. sinensis may not always be caused by the pathogen. Alternatively, and perhaps more likely, these individuals were presumably already carrying pathogens. Indeed, the hepatopancreas colour of all apparently healthy E. sinensis individuals used for control was assessed with the naked eye, and where pathogen was detected, individuals died. Similarly, not all pathogen-induced E. sinensis individuals (88.89%) displayed pathological symptoms of HPBD, possibly because some pathogen-induced crabs died before pathogen attack, hence the hepatopancreas colour was not significantly altered. In addition, in the samples displaying a colour change observed by the naked eye and OM was the presence of pathogen. Thus, analysis by the naked eye does not appear to be a precise method for evaluating the inflammatory responses following pathogen infection. The method should be improved using flow cytometry or real-time PCR in future studies.
Overall, the findings strongly suggest that S. aureus is extremely likely to be the pathogenic factor responsible for HPBD in E. sinensis. This is the first report to name the disease and identify the probable pathogen. This study also provides insight into pathogenic factors associated with HPND in E. sinensis based on a complication of hepatopancreatic necrosis. S. aureus damages the column-like epithelia of the hepatopancreas and alters the colour of this organ, but the mechanisms remain unknown. There may be other pathogenic factors involved, and future studies should investigate this possibility as well as the prevention and treatment of HPBD and HPND in E. sinensis using medicines and immunopotentiator screening. Decreasing the incidence of HPBD and HPND in pond-raised E. sinensis could ultimately prevent the heavy economic losses currently plaguing the aquaculture industry.
References
Ardic N, Ozyurt M, Sareyyupoglu B, Haznedaroglu T (2005) Investigation of erythromycin and tetracycline resistance genes in methicillin-resistant Staphylococci. Int J Antimicrob Agents 26(3):213–218
Chang GL, Cheng YX, Wu XG, Yang XZ, Wang ZK, Shen H (2008) Comparative studies of micro-ultrastructure and fatty acid composition of normal and albino hepatopancreas of Chinese mitten crab Eriocheir sinensis. Acta Hydrobiologica Sinica 32(5):687–693
Conaughty JM, Chen J, Martinez OV, Chiappetta G, Brookfield KF, Eismont FJ (2006) Efficacy of linezolid versus vancomycin in the treatment of methicillin-resistant Staphylococcus aureus discitis a controlled animal model. Spine 31(22):830–832
Deng XW, Xie ZX, Liu JB, Xie ZQ, Xie LJ, Pang YS (2010) Isolation and identification of Pseudomonas fluorescens in tilapia. Guangxi Agricultural Sciences 41:612–615
Emmerson M (1994) Nosocomial Staphylococcus outbreaks. Scand J Infect Dis Suppl 93(2):47–54
Feng SM, Zhan WB, Sheng XZ, Han JG, Wang J, Yang K, Li J, Zhang L (2009) Immune response to Edwardsiella tarda in turbot (Scophthal musmaximus). Periodical of Ocean University of China 39:203–208
Fisheries Bureau, Ministry of Agriculture (2016) China Fishery Statistical Yearbook of 2016. China Agriculture Press, Beijing
Gao ZQ, Xing H, Sun HC, Li HD (2003) Cloning and sequencing of the nuc gene of Staphylococcus aureus. Shanghai Laboratory Animal Science 23, 3-6.
GB 4789.10 (2010) National food safety standard, food microbiological examination: Staphylococcus aureus. National Standard of People’s Republic of China
Liu ZS, Lu SY, Cui SS (2014) Zoonosis. China Science Press, Beijing
Mehrotra M, Wang GH, Johnson WM (2000) Multiplex PCR for detection of genes for Staphylococcus aureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. J Clin Microbiol 38(3):1032–1035
Morandi S, Brasca M, Andrighetto C, Lombardi A, Lodi R (2009) Phenotypic and genotypic characterization of Staphylococcus aureus strains from Italian dairy products. International Journa1 of Medical Microbiology 2009(1):1–7
Peng Y, Luan Y, Wu SZ, Huang HQ (2013) Comparison of four identification methods of Staphylococcus aureus. Chinese Journal of Health Laboratory Technology 23:2919–2923
Pinto B, Chenoll E, Aznar R (2005) Identification and typing of food-borne Staph ylococcus aureus by PCR-based techniques. Syst Appl Microbiol 28:340–352
Purcell K, Fergie J (2006) Epidemic of community-acquired methicillin-resistant Staphylococcus aureus infections: a 14-year study at Driscoll Children’s Hospital. Digest of the World Core Medical Journals 159(10):980–985
Salgado CJ, Jamali AA, Mardini S, Buchanan K, Veit B (2005) A model for chronic osteomyelitis using Staphylococcus aureus in goats. Clin Orthop Relat Res 436(436):246–250.
Shi W, Lu MF, Cai ZH, Wu QP, Teng KL, Chen ZW (2012) The application study of freeze dried rabbit plasma in Staphylococcus aureus testing. Food and Fermentation Technology 48:44–47
Song XH, Cheng JX, Zhu MX, Cai CF, Yang CG, Shen ZH (2007a) Pathogenic factors of albinism in hepatopancreas of Chinese mitten crab Eriocheir sinensis (Decapoda: Grapsidae). Journal of Fishery Sciences of China 14(5):762–769
Song XH, Zhu MX, Wang YL, Li Y, Cao P, Shen ZH (2007b) The histopathological changes in tissues of Eriocheir sinensis with hepatopancreas albinism. J Fish China 31:257–263
Song XH, Yang CG, Cheng JX, Shen ZH, Wu LK, Cai CF, Gong CL (2007c) Effects of different feeding models on growth of Chinese mitten crab and their relation to its incidence of hepatopancreas albinism. J Fish China 31(2):496–503
Suo YJ, Yu HW, Ling W, Jia YM (2008) Analysis on the contamination of Staphylococcus aureus in food. Journal of Chinese Institute of Food Science and Technology 3:88–93
Tang Y, Zhang F, Sun HC, Geng XX, Tu Z, Wan YJ (2007) Isolation and identification of Klebslella pneumoniae from silver carp. Journal of southwest university (Natural science edition) 29(6):73–76
Tran L, Nunan L, Redman RM, Lightner DV, Fitzsimmons K (2013) EMS/AHPND: infectious disease caused by bacteria. Global Aquacuhure Advocat 7(8):18–20
Vandenesch F, Lina G, Henry T (2012) Staphylococcus aureus hemolysins, bi-component leukocidins, and cytolytic peptides: a redundant arsenal of membrane-damaging virulence factors? Front Cell Infect Microbiol 2(2):12
Wang HC, Hang H, Liu Y, Jiang ZH, Li JR (2014) Construction of predictive model for the growth of Staphylococcus aureus in bread shrimp. Guangdong Agricultural Sciences 22, 104-108.
Wang DC, Wang X, Zhou WJ, Peng XH, Gao Q (2010) Establishment and evaluation of a mouse model of skin abscess by infection with an isolated methicillin-resistant Staphylococcus aureus strain. Acta Laboratorium Animalis Scientia Sinica 3:221–224
Wu X, Su YC (2014) Growth of Staphylococcus aureus and enterotoxin production in pre-cooked tuna meat. Food Control 42:63–70
Zhang Y, Pan LD (2013) Histopatholgical study on hepatopancreas symptoms of Eriocheir sinensis. Guangdong Agricultural Sciences 40(9):118–133
Zhou G, Zhou QL, Xie J, Xi BW, Xu P (2011) Research advances of Aeromonas hydrophila epidemiology. Journal of Anhui Agricultural Sciences 39:14149–14151
Funding
This work was supported by the Modern Fisheries Industry Technology System Project of Jiangsu Province (grant number JFRS-01), the Important Research and Development Plan Project (Modern-agriculture) of Jiangsu Province (grant number BE2017383), and the Sanxin Aquaculture Project of Jiangsu Province (grant numbers D2015-5, D2016-1), China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed by the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fu, L., Gao, T., Jiang, H. et al. Staphylococcus aureus causes hepatopancreas browned disease and hepatopancreatic necrosis complications in Chinese mitten crab, Eriocheir sinensis. Aquacult Int 27, 1301–1314 (2019). https://doi.org/10.1007/s10499-019-00387-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10499-019-00387-1