Abstract
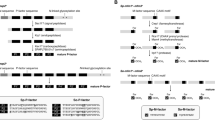
Genes involved in mating type determination and recognition were examined in Metschnikowia and related species, to gather insights on factors affecting mating compatibility patterns among haplontic, heterothallic yeast species of the genus. We confirmed the universality of the special mating locus organisation found in Clavispora lusitaniae across and exclusive to the family Metschnikowiaceae (i.e., Metschnikowia and Clavispora). Timing of the divergence between idiomorphs was confirmed to coincide with the origin of the larger (CUG-ser) clade comprising the Debaryomycetaceae and the Metschnikowiaceae, exclusive of Cephaloascus fragrans. The sequence of the a mating pheromone is highly conserved within the large-spored Metschnikowia species, including Metschnikowia orientalis and Metschnikowia hawaiiana, but not Metschnikowia drosophilae or Metschnikowia torresii, which have a pattern of their own, as do other clades in the genus. In contrast, variation in α pheromones shows a more continuous, although imperfect correlation with phylogenetic distance as well as with in vivo mating compatibility.






Similar content being viewed by others
References
Adhikari H, Cullen PJ (2015) Role of phosphatidylinositol phosphate signaling in the regulation of the filamentous-growth mitogen-activated protein kinase pathway. Eukaryot Cell 14:427–440
Audhya A, Foti M, Emr SD (2000) Distinct roles for the yeast phosphatidylinositol 4-kinases, Stt4p and Pik1p, in secretion, cell growth and organelle membrane dynamics. Mol Biol Cell 11:2673–2689
Beh CT, McMaster CR, Kozminski KG, Menon AK (2012) A detour for yeast oxysterol binding proteins. J Biol Chem 287:11481–11488
Bennett RJ, Turgeon BG (2016) Fungal sex: the ascomycota. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.FUNK-0005-2016
Butler G, Rasmussen MD, Lin MF et al (2009) Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature 459:657–662
Cevheroglu O, Kumas G, Melinda G, Becker JM, Son CD (2017) The yeast Ste2p G protein-coupled receptor dimerizes on the cell plasma membrane. Biochim Biophys Acta 1859:698–711
Dignard D, El-Naggar AL, Logue ME, Butler G, Whiteway M (2007) Identification and characterization of MFA1, the gene encoding Candida albicans a-factor pheromone. Eukaryot Cell 6:487–494
Gastaldi S, Zamboni M, Bolasco G, Di Segni G, Tocchini-Valentini GP (2016) Analysis of random PCR-originated mutants of the yeast Ste2 and Ste3 receptors. Microbiologyopen 5:670–686
Gonçalves-Sá J, Murray A (2011) Asymmetry in sexual pheromones is not required for ascomycete mating. Curr Biol 21:1337–1346
Guzmán B, Lachance MA, Herrera CM (2013) Phylogenetic analysis of the angiosperm-floricolous insect-yeast association: have yeast and angiosperm lineages co-diversified? Mol Phlogenet Evol 68:61–175
Hull CM, Johnson AD (1999) Identification of a mating type-like locus in the asexual pathogenic yeast Candida albicans. Science 285:1271–1275
Jones SK Jr, Bennett RJ (2011) Fungal mating pheromones: choreographing the dating game. Fungal Genet Biol 48:668–676
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kozakov D, Hall DR, Xia B, Porter KA, Padhorny D, Yueh C, Beglov D, Vajda S (2017) The ClusPro web server for protein-protein docking. Nat Protoc 12:255–278
Kurtzman CP (2003) Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. FEMS Yeast Res 4:233–245
Kurtzman CP (2011) Discussion of teleomorphic and anamorphic ascomycetous yeasts and yeast-like taxa. In: Kurtzman CP, Fell JW, Boekhout T (eds) The yeasts, a taxonomic study, vol 2. Elsevier, Amsterdam, pp 293–307
Kurtzman CP, Robnett CJ (2013) Relationships among genera of the Saccharomycotina (Ascomycota) from multigene phylogenetic analysis of type species. FEMS Yeast Res 13:23–33
Lachance MA (2011) Metschnikowia Kamienski (1899). In: Kurtzman CP, Fell JW, Boekhout T (eds) The yeasts, a taxonomic study, vol 2. Elsevier, Amsterdam, pp 575–620
Lachance MA (2012) In defense of yeast sexual life cycles: the forma asexualis, an informal proposal. Yeast Newsl 61:24–25
Lachance MA (2016) Paraphyly and (yeast) classification. Int J Syst Evol Microbiol 66:4924–4929
Lachance MA, Bowles JM (2004) Metschnikowia similis sp. nov. and Metschnikowia colocasiae sp. nov., two ascomycetous yeasts isolated from Conotelus spp. (Coleoptera: Nitidulidae) in Costa Rica. Stud Mycol 50:69–76
Lachance MA, Ewing CP, Bowles JM, Starmer WT (2005) Metschnikowia hamakuensis sp. nov., Metschnikowia kamakouana sp. nov. and Metschnikowia mauinuiana sp. nov., three endemic yeasts from Hawaiian nitidulid beetles. Int J Syst Evol Microbiol 55:1369–1377
Lachance MA, Collens JD, Peng XF, Wardlaw AM, Bishop L, Hou LY, Starmer WT (2016a) Spatial scale, genetic structure, and speciation of Hawaiian endemic yeasts. Pac Sci 70:389–408
Lachance MA, Hurtado E, Hsiang T (2016b) A stable phylogeny of the large-spored Metschnikowia clade. Yeast 33:261–275
Marinoni G (2003) Sexual cycle and speciation in the giant-spored Metschnikowia species. Dissertation, University of Western Ontario
Marinoni G, Lachance MA (2004) Speciation in the large-spored Metschnikowia clade and establishment of a new species, Metschnikowia borealis comb. nov. FEMS Yeast Res 4:587–596
McNeill J, Barrie FR, Buck WR, Demoulin V, Greuter W, Hawksworth DL, Herendeen PS, Knapp S, Marhold K et al (2012) International code of nomenclature for algae, fungi, and plants (Melbourne Code). Regnum Veg 154. ARG Gantner Verlag, Koenigstein
Merlini L, Dudin O, Martin SG (2013) Mate and fuse: how yeast cells do it. Open Biol 3:130008
Michaelis S, Barrowman J (2012) Biogenesis of the Saccharomyces cerevisiae pheromone a-factor, from yeast mating to human disease. Microbiol Mol Biol Rev 76:626–651
Raicu V, Stoneman MR, Fung R, Melnichuk M, Jasnma DB, Pisterzi LF, Rath S, Fox M, Wells JW, Saldin DK (2009) Determination of supramolecular structure and spatial distribution of protein complexes in living cells. Nat Photonics 3:107–113
Reedy JL, Floyd AM, Heitman J (2009) Mechanistic plasticity of sexual reproduction and meiosis in the Candida pathogenic species complex. Curr Biol 19:891–899
Riley R, Haridas S, Wolfe KH, Lopes MR, Hittinger CT, Göker M, Salamov AA, Wisecaver JH, Long TM, Calvey CH, Aerts AL, Barry KW, Choi C, Clum A, Coughlan AY, Deshpande S, Douglass AP, Hanson SJ, Klenk HP, LaButti KM, Lapidus A, Lindquist EA, Lipzen AM, Meier-Kolthoff JP, Ohm RA, Otillar RP, Pangilinan JL, Peng Y, Rokas A, Rosa CA, Scheuner C, Sibirny AA, Slot JC, Stielow JB, Sun H, Kurtzman CP, Blackwell M, Grigoriev IV, Jeffries TW (2016) Comparative genomics of biotechnologically important yeasts. Proc Natl Acad Sci USA 113:9882–9887
Robles LM, Millan-Pacheco C, Pastor N, Del Rio G (2017) Structure-function studies of the alpha pheromone receptor from yeast. Rev Esp Cienc Quim Biol 20:16–26
Rogers DW, McConnel E, Greig D (2012) Molecular quantification of Saccharomyces cerevisiae α-pheromone secretion. FEMS Yeast Res 12:668–674
Rogers DW, Denton JA, McConnell E, Greig D (2015) Experimental evolution of species recognition. Curr Biol 25:1753–1758
Sengupta P, Cochran BH (1990) The PRE and PQ box are functionally distinct yeast pheromone response elements. Mol Cell Biol 10:6809–6812
Shen Y, Maupetit J, Derreumaux P, Tuffery P (2014) Improved PEP-FOLD approach for peptide and miniprotein structure prediction. J Chem Theor Comput 10:4745–4758
Sherwood RK, Scaduto CM, Torres SE, Bennett RJ (2014) Convergent evolution of a fused sexual cycle promotes the haploid lifestyle. Nature 506:387
Soll DR, Daniels KJ (2016) Plasticity of Candida albicans biofilms. Microbiol Mol Biol Rev 80:565–595
Srikantha T, Daniels KJ, Pujol C, Sahni N, Yi S, Soll DR (2012) Nonsex genes in the mating type locus of Candida albicans play roles in a/α biofilm formation, including impermeability and fluconazole resistance. PLoS Pathog 8:e1002476
Stoneman MR, Paprocki JD, Biener G, Yokoi K, Shevade A, Kuchin S, Raicu V (2017) Quaternary structure of the yeast pheromone receptor Ste2 in living cells. Biochim Biophys Acta 1859:1456–1464
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Uddin MS, Hauser M, Naider F, Becker JM (2016) The N-terminus of the yeast G protein-coupled receptor Ste2p plays critical roles in surface expression, signaling and negative regulation. Biochim Biophys Acta 1858:715–724
Umanah GKE, Huang L, Ding F, Arshava B, Farley AR, Link AJ, Naider F, Becker JM (2010) Identification of residue to residue contact between a peptide ligand and its G protein-coupled receptor using periodate-mediated dihydroxyphenylalanine cross-linking and mass spectrometry. J Biol Chem 285:39425–39436
Wolfe KH, Butler G (2017) Evolution of mating in the Saccharomycotina. Annu Rev Microbiol 71:197–214
Wolfe KH, Armisén D, Proux-Wera E, ÓhÉigeartaigh SS, Azam H, Gordon JL, Byme KP (2015) Clade- and species-specific features of genome evolution in the Saccharomycetaceae. FEMS Yeast Res 15:fov035
Wu CI (2001) The genic view of the process of speciation. J Evol Biol 14:851–865
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Methods 12:7–8
Yi S, Sahni N, Daniels KJ, Lu KL, Srikantha T, Huang G, Garnaas AM, Soll DR (2011) Alternative mating type configurations (a/α versus a/a or α/α) of Candida albicans result in alternative biofilms regulated by different pathways. PLoS Biol 9:e1001117
Acknowledgements
Funding from the Natural Science and Engineering Research Council of Canada is gratefully acknowledged. Thanks are extended to K. Bensch for her assistance in obtaining the MycoBank numbers and to A. Spera for early contributions to exploration of the mating type locus.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, D.K., Hsiang, T. & Lachance, MA. Metschnikowia mating genomics. Antonie van Leeuwenhoek 111, 1935–1953 (2018). https://doi.org/10.1007/s10482-018-1084-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-018-1084-y