Abstract
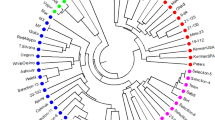
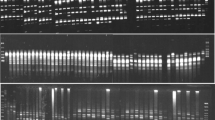
Pistachio (Pistacia vera L.) is considered to be the most widely cultivated species in the genus Pistacia and regarded as the most probable ancestral species within the genus. As one of the oldest nut crops in human history, pistachio nuts have a high nutritional value and are commercially important. In the present study, the genetic variation of pistachio genotypes was investigated by nuclear SSR markers. The transferability of the SSR markers was high across genotypes, thus, flanking regions of the SSRs are conserved in the studied genotypes. High level of polymorphism was detected among the studied genotypes. The seven SSR primer pairs generated a total of 18 alleles that 13 of them were polymorphic among the genotypes. The range of the polymorphic alleles was 1 (for Ptms9, Ptms40, Ptms41, and Ptms42) to 5 (for Ptms7 locus) with an average of 2.57. The amplified allele sizes ranged from 120 to 250 bp. Pair-wise genetic similarity coefficients varied from 0.20 to 0.75. The UPGMA dendrogram differentiated the genotypes into two major clusters. The present results may be used for the conservation, core collection and future breeding of the pistachio.



Similar content being viewed by others
References
Abdelkrim J, Robertson BC, Stanton JAL, Gemmell NJ (2009) Fast, cost effective development of species-specific microsatellite markers by genomic sequencing. BioTechniques 46:185–192
Ahmad R, Struss D, Southwick SM (2003) Development and characterization of microsatellite markers in Citrus. J Am Soc Hortic Sci 128:584–590
Ahmad R, Ferguson L, Southwick SM (2005) Molecular marker analyses of pistachio rootstocks by simple sequence repeats and sequence-related amplified polymorphisms. J Hortic Sci Biol 80(3):382–386
Allentoft ME, Schuster S, Holdaway RN, Hale ML, McLay E (2009) Identification of microsatellites from an extinct moa species using high throughput (454) sequence data. BioTechniques 46:195–200
Barazani O, Atayev A, Yakubov B, Kostiukovsky V, Popov K, Golan-Goldhirsh A (2003) Genetic variability in Turkmen populations of Pistacia vera L. Genet Resour Crop Evol 50:383–389
Buckler ES, Thornsberry JM (2002) Plant molecular diversity and applications to genomics. Curr Opin Plant Biol 5(2):107–111
Emanuelli F, Lorenzi S, Grzeskowiak L, Catalano V, Stefanini M, Troggio M (2013) Genetic diversity and population structure assessed by SSR and SNP markers in a large germplasm collection of grape. BMC Plant Biol 13(1):39
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
FAO (2014) FAO STAT data base. http://apps.fao.org. Accessed February 12
Fares K, Guasmi F, Touil L, Triki T, Ferchichi A (2009) Genetic diversity of pistachio tree using inter-simple sequence repeat markers ISSR supported by morphological and chemical markers. Biotechnology 8:24–34
Ferdinandez YSN, Somers DJ, Coulman BE (2001) Estimation the genetic relationship of hybrid bromegrass to smooth bromegrass and meadow bromegrass using RAPD markers. Plant Breed 120:149–153
Flint-Garcia SA, Thornsberry JM, Buckler ES (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Hamrick JL, Godt MJW (1996) Conservation genetics of endemic plant species. In: Acise JC, Hamrick JL (eds) Plant population genetics, breeding and genetic resource. Sinauer, Sunderland, pp 43–46
Hormaza JI, Pinney K, Polito VS (1998) Genetic diversity of pistachio (Pistacia vera, Anacardiaceae) germplasm based on randomly amplified polymorphic DNA (RAPD) markers. Econ Bot 52:78–87
de Jesus ON, de Oliveira e Silva S, Amorim EP, Ferreira CF, de Campos JMS, de Gaspari e Silva G, Figueira A (2013) Genetic diversity and population structure of Musa accessions in ex situ conservation. BMC Plant Biol 13:41
Levinson G, Gutman GA (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol 4:203–221
Liu Q, Song Y, Liu L, Zhang M, Sun J, Zhang S (2015) Genetic diversity and population structure of pear (Pyrus spp.) collections revealed by a set of core genome-wide SSR markers. Tree Genet Genomes 11(6):1–22
Maggs DH (1973) Genetic resources in pistachio. Plant Genet Resour Newsl 29:7–15
Malysheva-Otto LV, Ganal MW, Roder MS (2006) Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.). BMC Genet 7:6
Mantel N (1967) The detection of disease clustering and generalized regression approach. Cancer Res 27:209–220
Murray MG, Thompson WF (1980) Rapid Isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci Usa 76:5269–5273
Nimmakayala P, Kaur P, Bashet AZ, Bates GT, Langham R, Reddy OUK (2005) Molecular characterization of Sesamum using SSRs and AFLPs. Plant and animal genomes XIII conference, San Diego, pp 15–19
Parfitt DA, Badenes ML (1997) Phylogeny of the genus Pistacia as determined from analysis of the chloroplast genome. Proc Natl Acad Sci USA 94:7987–7992
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ (2000) NTSYS-pc numerical taxonomy and multivariate analysis system. Version 2.1. Exeter Software, Setauket
Romesburg HC (1990) Cluster analysis for researchers. Krieger Publishing Company, Malabar
Zohary D (1996) The genus Pistacia L. In: Padulosi S, Caruso T, Barone E (eds) Taxonomy, distribution, conservation and uses of Pistacia genetic resources. IPGRI, Palermo: 1‑11318 Conservation Genetics, (2010)11:311–318
Zohary D, Hopf M (1994) Domestication of plants in the old world. Clarendon Press, Oxford
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A. Khadivi declares that he has no competing interests.
Rights and permissions
About this article
Cite this article
Khadivi, A. Assessment of Genetic Variability in Pistachio (Pistacia vera L.) with Nuclear SSR Molecular Markers. Erwerbs-Obstbau 60, 289–294 (2018). https://doi.org/10.1007/s10341-018-0372-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10341-018-0372-z