Abstract
Objective
To develop a novel framework for evaluating the accuracy of quantitative analysis on dynamic contrast-enhanced (DCE) MRI with a specific combination of imaging technique, scanning parameters, and scanner and software performance and to test this framework with breast DCE MRI with Time-resolved angiography WIth Stochastic Trajectories (TWIST).
Materials and methods
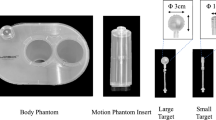
Realistic breast tumor phantoms were 3D printed as cavities and filled with solutions of MR contrast agent. Full k-space raw data of individual tumor phantoms and a uniform background phantom were acquired. DCE raw data were simulated by sorting the raw data according to TWIST view order and scaling the raw data according to the enhancement based on pharmaco-kinetic (PK) models. The measured spatial and temporal characteristics from the images reconstructed using the scanner software were compared with the original PK model (ground truth).
Results
Images could be reconstructed using the manufacturer’s platform with the modified ‘raw data.’ Compared with the ‘ground truth,’ the RMS error in all images was <10% in most cases. With increasing view-sharing acceleration, the error of the initial uptake slope decreased while the error of peak enhancement increased. Deviations of PK parameters varied with the type of enhancement.
Conclusion
A new framework has been developed and tested to more realistically evaluate the quantitative measurement errors caused by a combination of the imaging technique, parameters and scanner and software performance in DCE-MRI.








Similar content being viewed by others
References
Sullivan D (2016) The quantitative imaging biomarkers aliance (QIBA). In: Proceeding of the 24th scientific meeting, International Society for Magnetic Resonance in Medicine, Singapore, p 6875
Jackson EF (2016) Reproducibility and standardisation of MR biomarkers. In: Proceeding of the 24th scientific meeting, International Society for Magnetic Resonance in Medicine, Singapore, p 6876
Sullivan DC, Obuchowski NA, Kessler LG, Raunig DL, Gatsonis C, Huang EP, Kondratovich M, McShane LM, Reeves AP, Barboriak DP, Guimaraes AR, Wahl RL, Group R-QMW (2015) Metrology standards for quantitative imaging biomarkers. Radiology 277(3):813–825
Lim RP, Shapiro M, Wang EY, Law M, Babb JS, Rueff LE, Jacob JS, Kim S, Carson RH, Mulholland TP, Laub G, Hecht EM (2008) 3D time-resolved MR angiography (MRA) of the carotid arteries with time-resolved imaging with stochastic trajectories: comparison with 3D contrast-enhanced Bolus-Chase MRA and 3D time-of-flight MRA. AJNR Am J Neuroradiol 29(10):1847–1854
Korosec FR, Frayne R, Grist TM, Mistretta CA (1996) Time-resolved contrast-enhanced 3D MR angiography. Magn Reson Med 36(3):345–351
Mistretta CA (2009) Undersampled radial MR acquisition and highly constrained back projection (HYPR) reconstruction: potential medical imaging applications in the post-Nyquist era. J Magn Reson Imaging 29(3):501–516
Mistretta CA, Wieben O, Velikina J, Block W, Perry J, Wu Y, Johnson K (2006) Highly constrained backprojection for time-resolved MRI. Magn Reson Med 55(1):30–40
Lustig M, Donoho D, Pauly JM (2007) Sparse MRI: the application of compressed sensing for rapid MR imaging. Magn Reson Med 58(6):1182–1195
Trzasko JD, Haider CR, Borisch EA, Campeau NG, Glockner JF, Riederer SJ, Manduca A (2011) Sparse-CAPR: highly accelerated 4D CE-MRA with parallel imaging and nonconvex compressive sensing. Magn Reson Med 66(4):1019–1032
Herrmann KH, Baltzer PA, Dietzel M, Krumbein I, Geppert C, Kaiser WA, Reichenbach JR (2011) Resolving arterial phase and temporal enhancement characteristics in DCE MRM at high spatial resolution with TWIST acquisition. J Magn Reson Imaging 34(4):973–982
Tudorica LA, Oh KY, Roy N, Kettler MD, Chen Y, Hemmingson SL, Afzal A, Grinstead JW, Laub G, Li X, Huang W (2012) A feasible high spatiotemporal resolution breast DCE-MRI protocol for clinical settings. Magn Reson Imaging 30(9):1257–1267
Platel B, Mus R, Welte T, Karssemeijer N, Mann R (2014) Automated characterization of breast lesions imaged with an ultrafast DCE-MR protocol. IEEE Trans Med Imaging 33(2):225–232
Janka R, Wenkel E, Geppert C, Kiefer B, Uder M (2010) Breast MRI using TWIST: Doubling the spatial resolution “for free”. In: Proceeding of the 18th scientific meeting, International Society for Magnetic Resonance in Medicine, Stockholm, Sweden, p 4553
Song T, Laine AF, Chen Q, Rusinek H, Bokacheva L, Lim RP, Laub G, Kroeker R, Lee VS (2009) Optimal k-space sampling for dynamic contrast-enhanced MRI with an application to MR renography. Magn Reson Med 61(5):1242–1248
Le Y, Kroeker R, Geppert C, Dale B, Kipfer HD, Lin C (2013) Impact of TWIST View Sharing on Lesion Enhancement Profile in Dynamic Breast MRI. In: Proceeding of the 21th scientific meeting, International Society for Magnetic Resonance in Medicine, Salt Lake City, p 3375
Rashidnasab A, Elangovan P, Yip M, Diaz O, Dance DR, Young KC, Wells K (2013) Simulation and assessment of realistic breast lesions using fractal growth models. Phys Med Biol 58(16):5613–5627
Witten TA, Sander LM (1981) Diffusion-limited aggregation, a kinetic critical phenomenon. Phys Rev Lett 47(19):1400–1403
RepRap.org (2015) Prusa i3 Build Manual. http://reprap.org/wiki/Prusa_i3_Build_Manual. Accessed 6 March 2015
Tofts PS, Brix G, Buckley DL, Evelhoch JL, Henderson E, Knopp MV, Larsson HB, Lee TY, Mayr NA, Parker GJ, Port RE, Taylor J, Weisskoff RM (1999) Estimating kinetic parameters from dynamic contrast-enhanced T(1)-weighted MRI of a diffusable tracer: standardized quantities and symbols. J Magn Reson Imaging 10(3):223–232
Lopata RG, Backes WH, van den Bosch PP, van Riel NA (2007) On the identifiability of pharmacokinetic parameters in dynamic contrast-enhanced imaging. Magn Reson Med 58(2):425–429
Parker GJ, Roberts C, Macdonald A, Buonaccorsi GA, Cheung S, Buckley DL, Jackson A, Watson Y, Davies K, Jayson GC (2006) Experimentally-derived functional form for a population-averaged high-temporal-resolution arterial input function for dynamic contrast-enhanced MRI. Magn Reson Med 56(5):993–1000
Huang W, Li X, Chen Y, Chang MC, Oborski MJ, Malyarenko DI, Muzi M, Jajamovich GH, Fedorov A, Tudorica A, Gupta SN, Laymon CM, Marro KI, Dyvorne HA, Miller JV, Barbodiak DP, Chenevert TL, Yankeelov TE, Mountz JM, Kinahan PE, Kikinis R, Taouli B, Fennessy F, Kalpathy-Cramer J (2014) Variations of dynamic contrast-enhanced magnetic resonance imaging in evaluation of breast cancer therapy response: a multicenter data analysis challenge. Transl Oncol 7(1):153–166
Rakow-Penner R, Daniel B, Yu H, Sawyer-Glover A, Glover GH (2006) Relaxation times of breast tissue at 1.5T and 3T measured using IDEAL. J Magn Reson Imaging 23(1):87–91
Li X, Huang W, Yankeelov TE, Tudorica A, Rooney WD, Springer CS Jr (2005) Shutter-speed analysis of contrast reagent bolus-tracking data: preliminary observations in benign and malignant breast disease. Magn Reson Med 53(3):724–729
Henderson E, Rutt BK, Lee TY (1998) Temporal sampling requirements for the tracer kinetics modeling of breast disease. Magn Reson Imaging 16(9):1057–1073
Le Y, Geppert C, Kroeker R, Dale B, Lin C (2012) Simulations of the impact of TWIST view sharing on the measured enhancement in breast DCE MRI. In: Proceeding of the 20th scientific meeting, International Society for Magnetic Resonance in Medicine, Melbourne, p 2404
Le Y, Nickel D, Kroeker R, Geppert C, Dale B, Kipfer H, Lin C (2014) A simulation study of the flexible TWIST view sharing impact on the breast DCE MRI. In: Proceeding of the 22th scientific meeting, International Society for Magnetic Resonance in Medicine, Milan, Italy, p 6436
Mann RM, Mus RD, van Zelst J, Geppert C, Karssemeijer N, Platel B (2014) A novel approach to contrast-enhanced breast magnetic resonance imaging for screening: high-resolution ultrafast dynamic imaging. Invest Radiol 49(9):579–585
Agliozzo S, De Luca M, Bracco C, Vignati A, Giannini V, Martincich L, Carbonaro LA, Bert A, Sardanelli F, Regge D (2012) Computer-aided diagnosis for dynamic contrast-enhanced breast MRI of mass-like lesions using a multiparametric model combining a selection of morphological, kinetic, and spatiotemporal features. Med Phys 39(4):1704–1715
Carton AK, Bakic P, Ullberg C, Derand H, Maidment ADA (2011) Development of a physical 3D anthropomorphic breast phantom. Med Phys 38(2):891–896
Liney GP, Tozer DJ, Turnbull LW (1999) A simple and realistic tissue-equivalent breast phantom for MRI. J Magn Reson Imaging 10(6):968–971
Tiemann D, Wilbig J, Ezzat R, Terzenback D, Kompan IN, Laue H, Rascher-Friesenhausen R (2011) Perfusion mamma phantom adapted to physical MRI characteristics of breast tissue including tumorlike structure. In: Paper presented at the European Society for Magnetic Resonance in Medicine and Biology, Leipzig, Germany, October 6–8
Ahmed A, Gibbs P, Pickles M, Turnbull L (2013) Texture analysis in assessment and prediction of chemotherapy response in breast cancer. J Magn Reson Imaging 38(1):89–101
Kale MC, Fleig JD, Imal N (2013) Assessment of feasibility to use computer aided texture analysis based tool for parametric images of suspicious lesions in DCE-MR mammography. Comput Math Methods Med 2013:872676
Wang TC, Huang YH, Huang CS, Chen JH, Huang GY, Chang YC, Chang RF (2014) Computer-aided diagnosis of breast DCE-MRI using pharmacokinetic model and 3-D morphology analysis. Magn Reson Imaging 32(3):197–205
Teruel JR, Heldahl MG, Goa PE, Pickles M, Lundgren S, Bathen TF, Gibbs P (2014) Dynamic contrast-enhanced MRI texture analysis for pretreatment prediction of clinical and pathological response to neoadjuvant chemotherapy in patients with locally advanced breast cancer. NMR Biomed 27(8):887–896
Baish JW, Jain RK (2000) Fractals and cancer. Can Res 60(14):3683–3688
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
This study was sponsored by Siemens Healthcare.
Rights and permissions
About this article
Cite this article
Le, Y., Nickel, M.D., Kannengiesser, S. et al. A novel framework for evaluating the image accuracy of dynamic MRI and the application on accelerated breast DCE MRI. Magn Reson Mater Phy 31, 309–320 (2018). https://doi.org/10.1007/s10334-017-0648-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10334-017-0648-6