Abstract
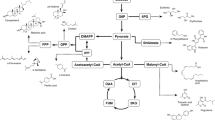
Traditional synthesis of biodiesel competes with food sources and has limitations with storage, particularly due to limited oxidative stability. Microbial synthesis of lipids provides a platform to produce renewable fuel with improved properties from various renewable carbon sources. Specifically, biodiesel properties can be improved through the introduction of a cyclopropane ring in place of a double bond. In this study, we demonstrate the production of C19 cyclopropanated fatty acids in the oleaginous yeast Yarrowia lipolytica through the heterologous expression of the Escherichia coli cyclopropane fatty acid synthase. Ultimately, we establish a strain capable of 3.03 ± 0.26 g/L C19 cyclopropanated fatty acid production in bioreactor fermentation where this functionalized lipid comprises over 32% of the total lipid pool. This study provides a demonstration of the flexibility of lipid metabolism in Y. lipolytica to produce specialized fatty acids.




Similar content being viewed by others
References
Adrio JL (2017) Oleaginous yeasts: promising platforms for the production of oleochemicals and biofuels. Biotechnol Bioeng 114:1915–1920. https://doi.org/10.1002/bit.26337
Beopoulos A, Mrozova Z, Thevenieau F, Le Dall MT, Hapala I, Papanikolaou S, Chardot T, Nicaud JM (2008) Control of lipid accumulation in the yeast Yarrowia lipolytica. Appl Environ Microbiol 74:7779–7789. https://doi.org/10.1128/aem.01412-08
Beopoulos A, Verbeke J, Bordes F, Guicherd M, Bressy M, Marty A, Nicaud JM (2014) Metabolic engineering for ricinoleic acid production in the oleaginous yeast Yarrowia lipolytica. Appl Microbiol Biotechnol 98:251–262. https://doi.org/10.1007/s00253-013-5295-x
Blazeck J, Hill A, Liu L, Knight R, Miller J, Pan A, Otoupal P, Alper HS (2014) Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat Commun 5:3131. https://doi.org/10.1038/ncomms4131
Blazeck J, Liu L, Redden H, Alper H (2011) Tuning gene expression in Yarrowia lipolytica by a hybrid promoter approach. Appl Environ Microbiol 77:7905–7914. https://doi.org/10.1128/aem.05763-11
Blazeck J, Reed B, Garg R, Gerstner R, Pan A, Agarwala V, Alper H (2013) Generalizing a hybrid synthetic promoter approach in Yarrowia lipolytica. Appl Microbiol Biotechnol 97:3037–3052. https://doi.org/10.1007/s00253-012-4421-5
Brian BL, Gardner EW (1968) A simple procedure for detecting the presence of cyclopropane fatty acids in bacterial lipids. Appl Microbiol 16:549–552
Brown JL, Ross T, McMeekin TA, Nichols PD (1997) Acid habituation of Escherichia coli and the potential role of cyclopropane fatty acids in low pH tolerance. Int J Food Microbiol 37:163–173
Chang YY, Cronan JE Jr (1999) Membrane cyclopropane fatty acid content is a major factor in acid resistance of Escherichia coli. Mol Microbiol 33:249–259
Dias MOS, Junqueira TL, Cavalett O, Cunha MP, Jesus CDF, Rossell CEV, Maciel Filho R, Bonomi A (2012) Integrated versus stand-alone second generation ethanol production from sugarcane bagasse and trash. Biores Technol 103:152–161. https://doi.org/10.1016/j.biortech.2011.09.120
Folch J, Lees M, Sloane Stanley GH (1957) A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem 226:497–509
Gaydou EM, Ralaimanarivo A, Bianchini JP (1993) Cyclopropanoic fatty acids of litchi (Litchi chinensis) seed oil. A reinvestigation. J Agric Food Chem 41:886–890. https://doi.org/10.1021/jf00030a009
Grogan DW, Cronan JE (1997) Cyclopropane ring formation in membrane lipids of bacteria. Microbiol Mol Biol Rev 61:429–441
Grogan DW, Cronan JE Jr (1984) Cloning and manipulation of the Escherichia coli cyclopropane fatty acid synthase gene: physiological aspects of enzyme overproduction. J Bacteriol 158:286–295
Hill J, Nelson E, Tilman D, Polasky S, Tiffany D (2006) Environmental, economic, and energetic costs and benefits of biodiesel and ethanol biofuels. Proc Natl Acad Sci 103:11206
Jungkind DL, Wood RC (1974) Physiological differences between cyclopropane fatty acid-deficient mutants and the parent strain of Streptococcus faecalis. Biochem Biophys Acta 337:298–310
Kai Y, Pryde EH (1982) Production of branched-chain fatty acids from sterculia oil. J Am Oil Chem Soc 59:300–305. https://doi.org/10.1007/BF02662231
Keweloh H, Diefenbach R, Rehm H-J (1991) Increase of phenol tolerance of Escherichia coli by alterations of the fatty acid composition of the membrane lipids. Arch Microbiol 157:49–53. https://doi.org/10.1007/bf00245334
Langlois A, Lebel O (2010) To cyclopropanate or not to cyclopropanate? A look at the effect of cyclopropanation on the performance of biofuels. Energy Fuels 24:5257–5263. https://doi.org/10.1021/ef100884b
Ledesma-Amaro R, Nicaud J-M (2016) Yarrowia lipolytica as a biotechnological chassis to produce usual and unusual fatty acids. Prog Lipid Res 61:40–50. https://doi.org/10.1016/j.plipres.2015.12.001
Li Q, Du W, Liu D (2008) Perspectives of microbial oils for biodiesel production. Appl Microbiol Biotechnol 80:749–756. https://doi.org/10.1007/s00253-008-1625-9
Da Silva Lima, Nd Maciel MRW, Batistella CB, Filho RM (2006) Optimization of biodiesel production from castor oil. Appl Biochem Biotechnol 130:405–414. https://doi.org/10.1385/abab:130:1:405
Liu L, Markham K, Blazeck J, Zhou N, Leon D, Otoupal P, Alper HS (2015) Surveying the lipogenesis landscape in Yarrowia lipolytica through understanding the function of a Mga2p regulatory protein mutant. Metab Eng 31:102–111. https://doi.org/10.1016/j.ymben.2015.07.004
Machida S, Shiraiwa Y, Suzuki I (2016) Construction of a cyanobacterium synthesizing cyclopropane fatty acids. Biochem Biophys Acta 1861:980–987. https://doi.org/10.1016/j.bbalip.2016.05.012
Markham KA, Alper HS (2018) Synthetic biology expands the industrial potential of Yarrowia lipolytica. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2018.05.004
Markham KA, Cordova L, Hill A, Alper HS (2017) Yarrowia lipolytica as a cell factory for oleochemical biotechnology. In: Lee SY (ed) Consequences of microbial interactions with hydrocarbons, oils, and lipids production of fuels and chemicals. Springer International Publishing, Cham, pp 1–19. https://doi.org/10.1007/978-3-319-31421-1_223-2
Markham KA, Palmer CM, Chwatko M, Wagner JM, Murray C, Vazquez S, Swaminathan A, Chakravarty I, Lynd NA, Alper HS (2018) Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation. Proc Natl Acad Sci 115:2096
Milla S, Elina L, Kurt F, Jorma K, Pirjo K, Jaanika B (2015) Macroalgae in biofuel production. Phycol Res 63:1–18. https://doi.org/10.1111/pre.12078
Mizoguchi T, Tsukatani Y, Harada J, Takasaki S, Yoshitomi T, Tamiaki H (2013) Cyclopropane-ring formation in the acyl groups of chlorosome glycolipids is crucial for acid resistance of green bacterial antenna systems. Bioorg Med Chem 21:3689–3694. https://doi.org/10.1016/j.bmc.2013.04.030
Munoz-Rojas J, Bernal P, Duque E, Godoy P, Segura A, Ramos JL (2006) Involvement of cyclopropane fatty acids in the response of Pseudomonas putida KT2440 to freeze-drying. Appl Environ Microbiol 72:472–477. https://doi.org/10.1128/aem.72.1.472-477.2006
Murphy DJ (1993) Structure, function and biogenesis of storage lipid bodies and oleosins in plants. Prog Lipid Res 32:247–280. https://doi.org/10.1016/0163-7827(93)90009-L
Papanikolaou S, Aggelis G (2002) Lipid production by Yarrowia lipolytica growing on industrial glycerol in a single-stage continuous culture. Bioresour Technol 82:43–49
Peralta-Yahya PP, Zhang F, del Cardayre SB, Keasling JD (2012) Microbial engineering for the production of advanced biofuels. Nature 488:320–328. https://doi.org/10.1038/nature11478
Poger D, Mark AE (2015) A ring to rule them all: the effect of cyclopropane fatty acids on the fluidity of lipid bilayers. J Phys Chem B 119:5487–5495. https://doi.org/10.1021/acs.jpcb.5b00958
Rakicka M, Lazar Z, Dulermo T, Fickers P, Nicaud JM (2015) Lipid production by the oleaginous yeast Yarrowia lipolytica using industrial by-products under different culture conditions. Biotechnol Biofuels 8:104. https://doi.org/10.1186/s13068-015-0286-z
Schmid KM (1999) Cyclopropane fatty acid expression in plants. US Patent US 5936139 A, 1999/08/10
Shabala L, Ross T (2008) Cyclopropane fatty acids improve Escherichia coli survival in acidified minimal media by reducing membrane permeability to H+ and enhanced ability to extrude H+. Res Microbiol 159:458–461. https://doi.org/10.1016/j.resmic.2008.04.011
Sherkhanov S, Korman TP, Clarke SG, Bowie JU (2016) Production of FAME biodiesel in E. coli by direct methylation with an insect enzyme. Sci Rep 6:24239. https://doi.org/10.1038/srep24239
Wagner JM, Williams EV, Alper HS (2018) Developing a piggyBac transposon system and compatible selection markers for insertional mutagenesis and genome engineering in Yarrowia lipolytica. Biotechnol J 13:1800022. https://doi.org/10.1002/biot.201800022
Wang AY, Grogan DW, Cronan JE Jr (1992) Cyclopropane fatty acid synthase of Escherichia coli: deduced amino acid sequence, purification, and studies of the enzyme active site. Biochemistry 31:11020–11028
Yu XH, Rawat R, Shanklin J (2011) Characterization and analysis of the cotton cyclopropane fatty acid synthase family and their contribution to cyclopropane fatty acid synthesis. BMC Plant Biol 11:97. https://doi.org/10.1186/1471-2229-11-97
Acknowledgements
This work was funded through the Office of Naval Research (ONR) under grant N00014-15-1-2785 and the Welch Foundation under grant F-1753. The authors acknowledge Andrew Hill for his initial work on CFA synthase enzymes in Yarrowia.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Markham, K.A., Alper, H.S. Engineering Yarrowia lipolytica for the production of cyclopropanated fatty acids. J Ind Microbiol Biotechnol 45, 881–888 (2018). https://doi.org/10.1007/s10295-018-2067-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2067-8