Abstract
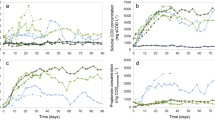
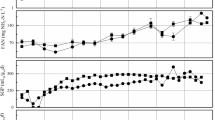
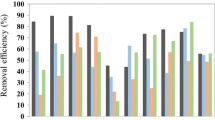
Even though full-scale digesters have been designed based on laboratory-scale tests, the substrate feeding modes of laboratory-scale tests might be different from those of full-scale digesters. The effect of substrate feeding frequencies on the performance and microbial community of laboratory-scale anaerobic digestion reactors was investigated. Feeding frequencies of twice a day, once a day, and every two days were tested in three 2-L reactors with an organic loading rate of 0.5 g-glucose/L/day under mesophilic condition. According to the results of this study, all the reactors showed similar methane production rates and SCOD removal efficiencies after sufficient time of acclimation, but frequently feeding promoted more stable digestion. Although there was no significant difference in microbial diversities from pyrosequencing analyses, the changes of archaeal community composition were observed. The decrease in feeding frequency appeared to cause shifts from acetoclastic methanogens affiliated with Methanosaeta to H2-utilizing methanogens. The increase of Methanosaeta at a frequently feeding might contribute to the stability of reactor operation. Since this study uses glucose as the substrate, there is still possibility of different results for more complex substrates, such as sludge, food waste, etc.




Similar content being viewed by others
References
Parkin GF, Owen WF (1986) Fundamentals of anaerobic digestion of wastewater sludges. J Environ Eng Landsc 112:867–920
Pohland F, Ghosh S (1971) Developments in anaerobic stabilization of organic wastes-the two-phase concept. Environ Lett 1:255–266
Chae K, Jang A, Yim S, Kim IS (2008) The effects of digestion temperature and temperature shock on the biogas yields from the mesophilic anaerobic digestion of swine manure. Bioresour Technol 99:1–6
Kim JK, Oh BR, Chun YN, Kim SW (2006) Effects of temperature and hydraulic retention time on anaerobic digestion of food waste. J Biosci Bioeng 102:328–332
Chen Y, Cheng JJ, Creamer KS (2008) Inhibition of anaerobic digestion process: a review. Bioresour Technol 99:4044–4064
Li Y, Park SY, Zhu J (2011) Solid-state anaerobic digestion for methane production from organic waste. Renew Sustain Energy Rev 15:821–826
Bombardiere J, Espinosa-Solares T, Domaschko M, Chatfield M (2007) Thermophilic anaerobic digester performance under different feed-loading frequency. In: Applied biochemistry and biotecnology, Springer, Berlin, p 765–775
Golkowska K, Sibisi-Beierlein N, Greger M (2012) Kinetic considerations on thermophilic digestion of maize silage at different feeding modes. Chem Ing Tech 84:1551–1558
Inglis SF, Gooch CA (2007) Biogas production fluctuations associated with feeding frequency for a mixed digester. In: Sixth International Dairy Housing Conference, American Society of Agricultural and Biological Engineers
Piao ZH, Kim CH, Kim JY (2013) Effect of feeding frequency on performance of laboratory-scale anaerobic digestion reactor. In: The 1st IWWG-ARB Symposium, Hokkaido University, Japan
Li Y, Zhang R, He Y, Zhang C, Liu X, Chen C, Liu G (2014) Anaerobic co-digestion of chicken manure and corn stover in batch and continuously stirred tank reactor (cstr). Bioresour Technol 156:342–347
Escudero A, Lacalle A, Blanco F, Pinto M, Díaz I, Domínguez A (2014) Semi-continuous anaerobic digestion of solid slaughterhouse waste. J Environ Chem Eng 2:819–825
Wang M, Sun X, Li P, Yin L, Liu D, Zhang Y, Li W, Zheng G (2014) A novel alternate feeding mode for semi-continuous anaerobic co-digestion of food waste with chicken manure. Bioresour Technol 164:309–314
Conklin A, Stensel HD, Ferguson J (2006) Growth kinetics and competition between Methanosarcina and Methanosaeta in mesophilic anaerobic digestion. Water Environ Res 78:486–496
De Vrieze J, Verstraete W, Boon N (2013) Repeated pulse feeding induces functional stability in anaerobic digestion. Microb Biotechnol 6:414–424
Lee S-H, Kang H-J, Lee YH, Lee TJ, Han K, Choi Y, Park H-D (2012) Monitoring bacterial community structure and variability in time scale in full-scale anaerobic digesters. J Environ Monitor 14:1893–1905
Shelton DR, Tiedje JM (1984) General method for determining anaerobic biodegradation potential. Appl Environ Microbiol 47:850–857
Chun J, Kim KY, Lee J-H, Choi Y (2010) The analysis of oral microbial communities of wild-type and toll-like receptor 2-deficient mice using a 454 gs flx titanium pyrosequencer. BMC Microbiol 10:1
Hur M, Kim Y, Song H-R, Kim JM, Im Choi Y, Yi H (2011) Effect of genetically modified poplars on soil microbial communities during the phytoremediation of waste mine tailings. Appl Environ Microbiol 77:7611–7619
Kim B-S, Kim JN, Yoon S-H, Chun J, Cerniglia CE (2012) Impact of enrofloxacin on the human intestinal microbiota revealed by comparative molecular analysis. Anaerobe 18:310–320
Huber T, Faulkner G, Hugenholtz P (2004) Bellerophon: a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20:2317–2319
Kim O-S, Cho Y-J, Lee K, Yoon S-H, Kim M, Na H, Park S-C, Jeon YS, Lee J-H, Yi H (2012) Introducing eztaxon-e: a prokaryotic 16 s rrna gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Micr 62:716–721
APHA (2005) Standard methods for the examination of water and wastewater, 21st edn. APHA, Washington
Hill D (1982) A comprehensive dynamic model for animal waste methanogenesis. T ASAE 25:1374–1380
Ward AJ, Hobbs PJ, Holliman PJ, Jones DL (2008) Optimisation of the anaerobic digestion of agricultural resources. Bioresour Technol 99:7928–7940
Chen W-H, Han S-K, Sung S (2003) Sodium inhibition of thermophilic methanogens. J Environ Eng Landsc 129:506–512
O’connor OA, Young L (1989) Toxicity and anaerobic biodegradability of substituted phenols under methanogenic conditions. Environ Toxicol Chem 8:853–862
Boon N, De Windt W, Verstraete W, Top EM (2002) Evaluation of nested pcr–dgge (denaturing gradient gel electrophoresis) with group-specific 16 s rrna primers for the analysis of bacterial communities from different wastewater treatment plants. FEMS Microbiol Ecol 39:101–112
Hugenholtz P, Tyson GW, Webb RI, Wagner AM, Blackall LL (2001) Investigation of candidate division tm7, a recently recognized major lineage of the domain bacteria with no known pure-culture representatives. Appl Environ Microbiol 67:411–419
Ariesyady HD, Ito T, Okabe S (2007) Functional bacterial and archaeal community structures of major trophic groups in a full-scale anaerobic sludge digester. Water Res 41:1554–1568
Narihiro T, Terada T, Kikuchi K, Iguchi A, Ikeda M, Yamauchi T, Shiraishi K, Kamagata Y, Nakamura K, Sekiguchi Y (2009) Comparative analysis of bacterial and archaeal communities in methanogenic sludge granules from upflow anaerobic sludge blanket reactors treating various food-processing, high-strength organic wastewaters. Microbes Environ 24:88–96
Romano RT, Zhang R, Teter S, Mcgarvey JA (2009) The effect of enzyme addition on anaerobic digestion of josetall wheat grass. Bioresour Technol 100:4564–4571
Ouverney CC, Armitage GC, Relman DA (2003) Single-cell enumeration of an uncultivated tm7 subgroup in the human subgingival crevice. Appl Environ Microbiol 69:6294–6298
Chouari R, Le Paslier D, Daegelen P, Ginestet P, Weissenbach J, Sghir A (2005) Novel predominant archaeal and bacterial groups revealed by molecular analysis of an anaerobic sludge digester. Environ Microbiol 7:1104–1115
Qiao J-T, Qiu Y-L, Yuan X-Z, Shi X-S, Xu X-H, Guo R-B (2013) Molecular characterization of bacterial and archaeal communities in a full-scale anaerobic reactor treating corn straw. Bioresour Technol 143:512–518
Zhang H, Banaszak JE, Parameswaran P, Alder J, Krajmalnik-Brown R, Rittmann BE (2009) Focused-pulsed sludge pre-treatment increases the bacterial diversity and relative abundance of acetoclastic methanogens in a full-scale anaerobic digester. Water Res 43:4517–4526
Gorlas A, Robert C, Gimenez G, Drancourt M, Raoult D (2012) Complete genome sequence of methanomassiliicoccus luminyensis, the largest genome of a human-associated archaea species. J Bacteriol 194:4745
Liu Y, Whitman WB (2008) Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann NY Acad Sci 1125:171–189
Acknowledgments
This study was supported by the Korea Ministry of Environment as “Knowledge-based environmental service (Waste to energy) Human Resource Development Project”, the Korea Ministry of Education through the Brain Korea 21 Plus research program, and the National Research Foundation of Korea (NRF) Grant (No. 2015R1A5A7037372) funded by the Korean Government (MSIP). Authors also appreciate the technical support of the Institute of Construction and Environmental Engineering and Engineering Research Institute, Seoul National University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Piao, Z.H., Lee, J. & Kim, J.Y. Effect of substrate feeding frequencies on the methane production and microbial communities of laboratory-scale anaerobic digestion reactors. J Mater Cycles Waste Manag 20, 147–154 (2018). https://doi.org/10.1007/s10163-016-0556-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10163-016-0556-2