Abstract
Background
The acquisition of an invasive phenotype by a tumor cell is a crucial step of malignant transformation. The underlying genetic mechanisms in gastric cancer (GC) are not well understood.
Methods
We performed whole-exome sequencing of 15 pairs of primary GC and their matched lymph node (LN) metastases (10 primary GCs with single matched LNs and 5 primary GCs with three LNs per case, respectively). Somatic alterations including single nucleotide variations, short insertions/deletions including locus-level microsatellite instability and copy number alterations were identified and compared between the primary and metastatic LN genomes.
Results
Mutation abundance was comparable between the primary GC and LN metastases, but the extent of mutation overlap or the mutation heterogeneity between primary and LN genomes varied substantially. Primary- or LN-specific mutations could be distinguished from common mutations in terms of mutation spectra and functional categories, suggesting that the mutation forces are not constant during gastric carcinogenesis. A spatial distribution revealed TP53 mutations as common mutations along with a number of region-specific mutations, such as primary-specific SMARCA4 and LN-specific CTNNB1 mutations. The subclonal architectures of common mutations were largely conserved between primary GC and LN metastatic genomes. The mutation-based phylogenetic analyses further showed that LN metastases may have arisen from homogeneous subclones of primary tumors.
Conclusions
The abundance and spatial distribution of mutations may provide clues on the evolutionary relationship between primary and matched LN genomes. Gene-level analyses further distinguished the early addicted cancer drivers such as TP53 mutations from late acquired region-specific mutations.
Similar content being viewed by others
Introduction
Gastric cancers (GCs) are one of the most common tumor types accounting for about 8% of newly diagnosed cancers and 10% of cancer mortality worldwide [1]. Endoscope-based GC screening has enabled the early detection of the disease and also has reduced the GC-related risks, such as mortality rates, especially in Eastern Asia, which has the highest regional incidence of GC. However, surgical resection remains the only curative means for GC before invasion and metastasis develop. The acquisition of the potential for invasion by GC cells is crucial in the development of gastric malignancies. A detailed molecular understanding of the invasive and metastatic potential could improve the currently used diagnostic and therapeutic modalities [2].
A stepwise model has linked the critical steps in carcinogenesis including tumor progression and metastasis with the accumulation of specific genetic aberrations. Next-generation sequencing has proven to be a powerful tool for genome-wide screening of genomic aberrations associated with cancer development, such as single nucleotide variations (SNVs), short insertions/deletions (indels), microsatellite instability (MSI), and copy number alterations (CNAs) [3]. A detailed profiling of the mutational landscape of GC genomes including the paired analyses of primary and metastatic genomes should advance our understanding of related molecular mechanisms. Along with the high-resolution platform of whole-genome sequencing, the capture-based whole-exome sequencing has been used to compare primary and metastatic GC genomes. For example, two primary and three lymph node (LN) metastatic genomes in a given patient identified a substantial level of heterogeneity, as well as the potential divergent genomic events [4]. Comparative studies have been published for primary GC genomes and peritoneal ascitic metastatic genomes [5, 6]. These studies along with our previous works [7, 8] have shown the potential of high-throughput sequencing-based multi-region analysis to identify the genomic aberrations associated with specific cellular events, such as the acquisition of malignant potential. For example, sequencing-based inference of subclonal architecture of cancer genomes using the burdens of mutant alleles may reveal the subclones arising during the progression or relapse of cancers as well as their evolutionary relationship with primary tumors [7, 8]. In addition, the sequencing of multiple regional biopsies in given individuals and the mutation-based inference of phylogenetic trees have been used in elucidating the evolutionary relationship of regional biopsies (e.g., primary and metastatic genomes) [9, 10]. Thus, the regional LN metastases of GC can be investigated in terms of genome- or exome-wide mutation abundance and their evolutionary relationship between the primary tumor genomes and their matched LN tumor genomes. LN metastasis is a major factor in predicting patient survival. The metastasis likely precedes systemic diseases that feature distant metastases [11]. Thus, mutation profiles may also provide additional mechanistic and evolutionary insights on the acquisition of tumor invasive potential with potential clinical relevance.
In this study, we aimed to identify the mutations of primary GC and their matched local LN metastases. For this, we performed whole exome-based mutation analyses for 15 pairs of primary and matched LN metastatic GC genomes. Among the cases, 10 cases were sequenced for primary tumor and their single matched lymph nodes (P and L, respectively) as well as matched normal genomes. For additional five cases, we obtained three lymph nodes (P for primary and L1–L3 for 3 matched LNs) per case and sequenced with matched normal genomes. By comparing the somatic mutations between the primary and metastatic genomes, three mutational categories were distinguished: common and primary-/LN-specific mutations. This categorization is associated with the relative timing of mutation acquisition, with common mutations as early changes and region-specific mutations representing those occurring late during the evolution of the cancer genome. Mutation features, such as sequence properties associated with individual mutation categories, as well as the mutation frequency were investigated to identify the mutation abundance and their subclonal architecture along with phylogenetic trees associated with the regional LN metastases.
Materials and methods
Patients
Fifteen gastric cancer patients who underwent gastrectomy with combined LN dissection between March 2014 and August 2017 were enrolled in the study. This study was approved by an institutional review board (UC15SISI0180 and KC18SESI0028). The primary tumor specimens were snap-frozen. The frozen section were stained with hematoxylin and eosin (H&E) for histologic examination by board-certified pathologists for the tumor purity (> 70%). After the positive LNs were confirmed by pathologists, the areas with metastatic cancer tissues were identified in the slide that had been stained with H&E. Ten micrometer thick sections were utilized for microdissection. Tissue sections were micro-dissected using a PixCell II LCM system (Arcturus, Mountain View, CA, USA) by the operator. For matched normal genome DNA, patient’s blood was also collected in EDTA-treated tumors. The clinicopathologic features of 15 gastric cancers are summarized in Table 1. One case (GC15) is selected to show the H&E section images for primary and one lymph nodes (Supplementary Fig. 1).
Whole-exome sequencing
We used the DNeasy Blood and Tissue Kit (Qiagen, Germany) to extract the genomic DNA according to the manufacturer’s recommendations. For each of 15 GC patients, the genomic DNA was obtained from primary tumor sites and available LNs as well as from the patient blood. The exomic DNA was captured using the Agilent SureSelect Human All Exome 50 Mb kit (Agilent, USA). Genomic DNA libraries were prepared and 100 bp paired-end sequencing reads were generated using the Illumina HiSeq2000 platform according to the manufacturer’s recommendations (Illumina, USA). Sequencing information, including coverage, is available in Supplementary Table 1.
Somatic mutations
The Burrows–Wheeler aligner [12] was used to align the sequencing reads onto the human reference genome (hg19). Genome Analysis ToolKit [13] was used with appropriated reference datasets to further realign the sequencing reads and recalibrate the score. The SamTools sequencing management tool [14] was used for additional data processing. Somatic alterations were identified by comparing the tumor and matched normal genome-derived sequencing data. For SNV and indels, we used MuTect [15] and Indelocator [13]. The ANNOVAR tool [16] was used to curate the somatic SNVs and indels on coding sequences and their functional consequences with respect to codon changes. Each SNV and indel was assigned to mutation spatial categories (common and primary-/LN-specific categories). Common mutations were those observed both in the primary and LN genomes while primary- or LN-specific mutations were those observed only in the primary or LN genomes, respectively. In addition, we performed joint calling of somatic mutations so as not to overestimate the extent of heterogeneity. For this, the mutations called in either primary or LN genomes were reexamined for the presence of sequencing reads supporting the mutations in the other biopsy of the given case [7, 10]. In the five cases with three matched LNs were examined, primary- and LN-specific mutations were also identified as those exclusive to primary or LN genomes.
MSI analyses
To identify the locus-level MSI calls, we used our inhouse MSI calling algorithm [17]. For 146,000 reference microsatellite or tandem repeats identified on coding sequences, we obtained the repeat length distribution from the tumor and matched normal genomes. Kolmogorov–Smirnov test was applied to estimate the significance for the differences in two repeat length distribution. False discovery rate (FDR) < 0.05 was considered an MSI call for each comparison. We have previously shown that the abundance of locus-level MSI calls can distinguish the MSI-H-vs.-MSI-L or MSS cases. Across the 15 GC cases, 10, 7 and 72 MSI calls were identified in GC8, GC9 and GC10 genomes, respectively, but none or one MSI call was identified in the remaining MSS cases (e.g., no MSI calls found in GC7 in spite of the high mutation burden). We validated our finding using the capillary sequencing for five Bethesda markers: BAT25, BAT25, D2S123, D5S346, and D17S250. A substantial fraction of MSI calls overlap with those of indels. Given the high sensitivity of our inhouse indel calling algorithm [17], we only considered MSI calls that do not overlap with indels.
Copy number profiling
VarScan2 [18] was used for CNA profiling to obtain read depth differences between the tumor and matched normal sequencing data. The read depth ratio of genomic bins was corrected for GC contents and further log2-transformed. The segmentation was done using circular binary segmentation algorithm [19] and visualized using IGV browser [20].
Phylogenetic analysis
To construct phylogenetic trees for five cases with one primary tumor and three matched LN genomes sequenced, we generated a binary matrix of somatic mutations per case. Exonic and non-exonic SNVs with coverage > 10 sequencing reads were collected and used to generate mutation matrix. Maximum parsimony trees were inferred using branch-and-bound algorithm with PHYLIP software [21] as we have previously described [22].
Results
Somatic mutations of primary and LN GC genomes
To obtain mutational profiles of 15 pairs of primary GCs and their matched LN metastases, we performed whole-exome sequencing. The sequencing information of 15 pairs of tumor and LN genomes, as well as their matched normal genomes are presented in Supplementary Table 1. The clinicopathological information of the 15 GC patients is summarized in Table 1. Somatic mutations were identified by comparing the tumor genomes (primary and LN genomes, respectively) with matched normal genomes. Somatic SNVs, indels and MSI events were identified using MuTect [15], Indelocator [13] and our in-house MSI caller [17], respectively. A total of 11,601 somatic variants were identified (10,877 SNVs and 724 indels/MSI). The full list of somatic variants is presented in Supplementary Table 2.
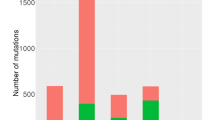
The mutation abundance of 10 GC pairs [one primary tumor and one matched LN; 7 microsatellite stable (MSS) and 3 microsatellite instability high (MSI-H) cases] as well as 5 GC pairs (one primary tumor and three matched LNs; 5 MSS cases) are shown in Fig. 1a. For MSS GC genomes, the number of somatic variants was 66–780 for primary tumors and 54–784 for LN metastases. MSI-H GC genomes showed an elevated mutation abundance compared to MSS genomes (e.g., for GC10, both primary and LN genomes harbored > 3000 somatic mutations). But, there was no clear-cut mutation abundance between the MSS and MSI-H genomes. For example, two MSS genomes (GC7 and GC15) also showed highly elevated mutation abundance (> 300 mutations both for primary and LN genomes) and the mutation abundance of one MSI-H case (GC8; 229/234 mutations in primary and LN genomes, respectively) was comparable to those of MSS GC genomes. Of note, the mutation abundance of primary tumor and matched LN genomes was not significantly different (P = 0.103; paired t test) suggesting that the mutation abundance of primary GC genomes are largely comparable to those of matched LN genomes. However, the mutational heterogeneity (i.e., the extent of overlap between the mutations from primary and LN genomes) was variable across the cases. For example, only 18 mutations were overlapping between 1390 and 893 primary and LN mutations (1.2 and 2.0%, respectively) for GC9/MSI-H case while the corresponding percentages were 96.1 and 86.3% for GC6/MSS case.
Mutation abundance of 15 GC genomes. a The mutation abundance or the number of somatic variants are shown in y-axis for primary GC genomes and their matched metastatic LN genomes across 15 GC pairs. Seven MSS (left) and 3 MSI-H GC genomes with single matched LN genomes (middle) are separately shown from five MSS GC cases with three matched LN genomes (right) with different scales. Grey represents the common mutations (e.g., those observed both in primary and LN genomes) while green and red represent the number of variants specific to primary and metastatic LN genomes, respectively. b The functional consequences of exonic variants and splice site mutations are shown for their relative fraction. For each case, the fractions are separately shown for the common, primary-specific and LN-specific mutations. c Similarly shown for the six mutation spectra. d Relative fractions are shown for mutation signatures. Eight mutation signatures with known mutagens or genomic events are shown with other signatures combined
For three types of mutations (‘common’ mutations as those both observed in primary and LN genomes and the lesion-specific mutations as ‘primary-’ or ‘LN-specific’ mutations, respectively), the functional consequence of coding proteins (Fig. 1b) were investigated along with the mutation spectra and signatures (Fig. 1c, d, respectively). Across the GC genomes examined, missense mutations dominated other types of mutations overall. However, a substantial number of frameshift indels were observed for MSI-H GC genomes, especially in common mutation categories (e.g., GC9 and GC10 MSI-H genomes; Fig. 1b). In mutation spectra, the relative enrichment of C > A mutations accompanied by depletion of C > T mutations were observed for LN-specific for half of the cases (Fig. 1c). C > A mutations have been previously reported to be enriched in malignant ascites of advanced gastric cancers [5] and these reports suggest that C > A mutations represent late, region-specific genomic aberrations during gastric carcinogenesis. We further performed mutation signature analyses to classify the mutations according to the trinucleotide contexts and associate them with probable mutagenic events as previously reported [23] (Fig. 1d). The relative proportion of mutation signatures with known associated mutagens or causal events are heterogeneous across the cases and regions. Signature 1A/1B that accounts for C > T mutations and is correlated with patient ages are consistently observed across cases, often comprising the majority of the mutations in given cases (black in Fig. 1d). These mutations may represent those that have arisen before the emergence of tumor-initiating cells [24], and consistently, they are relatively enriched with common mutations. The signature 6 associated with DNA mismatch repair deficiency (MMRd) is enriched for two MSI-H cases of GC9 and GC10 (red in Fig. 1d). Although a substantial number of region-specific mutations are called as signature 6 in both cases, a relative deficit of signature 6 in common mutations of GC9 suggests that MMRd may have occurred just before the divergence of LN metastasis, in this case, constituting a majority of primary- and LN-specific mutations. In the case of GC7 genome with elevated mutation burdens without MMRd, signature 17 mutations comprise the majority of lesion-specific mutations. Although the mutation signature is not well known for their causality, our observation suggests that the signature may be associated with potential hypermutation-associated events.
CNAs in primary GC and matched LN genomes
Using whole-exome sequencing data, we performed genome-wide CNA profiling for 15 pairs of primary GC and lymph node genomes (Fig. 2a). In some GC genomes, the CNAs are largely concordant between the primary vs. metastatic LN genomes. For example, CNAs of GC8 genomes are largely shared between the primary and LN genomes suggesting that these CNAs may have occurred before the emergence of regional LN metastases. The concordance of CNAs between the primary tumor and matched LNs are separately shown (Fig. 2b). Spearman correlation between primary and single-matched lymph node CNAs are shown for GC1–GC10 (black bars; Fig. 2b). Except for a case GC7/MSS with correlation of 0.03, the concordance levels between primary-vs.-lymph node genomes were observed in the range of 0.23 (GC9/MSI-H) to 0.67 (GC8/MSI-H). We also observed that the concordance level of CNAs between primary and LN genomes are highly correlated with the mutation-level concordance (i.e., the ratio of the number of common mutations over total mutations of given case) (r = 0.889 for GC1-GC10) suggesting that the somatic mutations and CNAs are shared between two regional biopsies to a similar extent. This is consistent with our previous report in colorectal cancers [10] where the evolutionary distance and relationship estimated by somatic mutations and CNAs were similar between regional biopsies in a given individual. For GC11–GC15 with 3 matched LN genomes available per case, the extent of CNA-level concordance those between primary-vs.-LN genomes and among LN genomes are shown with green and red whiskers, respectively (Fig. 2b). For these genomes, we also observed that the correlation values among the LN genomes are higher than those between primary and LN genomes. This suggests that the LN genomes are highly related to each other than their matching primary genomes, which is consistent with a late-dissemination model where a single subclone or genetically heterogeneous subclones in the primary tumor are responsible for metastases [10, 25].
CNA of GC genomes. a For GC genomes, genome-wide CNA profiles are shown as snapshots of IGV browser. The genomic amplifications and deletions are shown as red and blue, respectively. Primary- and LN genomes in given case are shown in upper and lower lanes (GC1–GC10) and in four lanes in order of primary and LN1-3 genomes (GC11-GC15). b The correlation of the CNAs between the primary and LN genomes are shown as bar plots (black; GC1-GC10) and green whiskers for GC11-15. The correlation values among the three LN genomes are shown in red whiskers (GC11-15)
Interpretation of recurrent mutations with respect to their regional presentation
Table 2 lists the recurrent non-silent mutations in known cancer-related genes of Cancer Census Genes [21] observed in no less than 3 cases among 15 GC genomes. Most of TP53 mutations were observed as common mutations (7 out of 10 TP53 non-silent mutations) suggesting that the TP53 mutations represent early genomic aberrations in gastric carcinogenesis before the divergence of LN metastasis from primary tumors. Among the 10 TP53 mutations observed, 5 out of 8 missense mutations have occurred in known mutation hotspots (i.e., amino acids 125–300 corresponding to the DNA binding domain) [26] and the other 2 mutations were truncating events (one frameshifting indel and one nonsense mutation) suggesting that most of TP53 mutations are functional inactivating events for the corresponding cases. Common PIK3CA mutations were observed in three cases often accompanied by additional primary- and LN-specific PIK3CA mutations. All three PIK3CA common mutations occurred in known hotspots (p.E545K/GC1, p.E542K/GC7, p.H1047R/GC10) but region-specific mutations occurred outside of known hotspots (p.M1043I/GC1, p.N756D/GC7, p.L15V/GC7, and D129N/GC15) [27]. This suggests that PIK3CA hotspot mutations as potential cancer drivers may have occurred earlier in the carcinogenesis like TP53 common mutations. And the functional implications of the additionally acquired non-hotspot, biopsy-specific PIK3CA mutations require further investigation since the occurrence of two PIK3CA mutations in independent clones may represent the potential functional divergence [27]. Recurrent RNF43 indel/MSI (p.G659fs) have been recently identified in gastric cancers [28] and we observed the corresponding mutations as common events for two MSI-H genomes (GC9 and GC10). Three primary-specific RNF43 mutations were identified as one nonsense and two missense mutations suggesting that activating RNF43 mutations in addition to known DNA slippage errors may be also common and often region-specific. It is of interests that POLE mutations are solely observed as region-specific (two primary- and three LN-specific mutations, respectively). As a proofreading DNA polymerase, POLE mutations have been associated with hypermutated phenotype in some tumor types including GC [29, 30]. Of interest, one MSS genome with comparable mutation abundance (GC7) harbored one primary-specific missense mutation (p.N875Y) and LN-specific mutation (p.E584D) that may be responsible for the elevated region-specific mutation rates compared to that of common mutations in this case. Three additional mutations (two missense and one frameshift indel) were also observed as either primary- or LN-specific mutations. One splicing CDH1 mutation was identified as a common mutation along with two additional primary-specific, missense mutations. CDH1 mutations in GC genomes showed unique preference for a GC subtype called a genomically stable subtype [31] and our study further suggests that CDH1 mutations may be often late, primary-specific events.
Among the cancer-related genes with recurrent mutations, the regional distribution of mutations (e.g., common, primary- and LN-specific) may provide clues on the relative timing or temporal order of somatic mutations acquired. Thus, we next investigated the somatic mutations with regional biases. Table 3 lists non-silent mutations of cancer-related genes significantly enriched to any of three regions (common, primary- and LN-specific; P < 0.1, Fisher’s exact test). Consistent with Table 2, TP53 and LRP1B mutations were relatively specific to common mutations. Nonsilent mutations on CSMD3 and ERBB3 were observed in three cases and one case as common and primary-specific mutations, respectively and thus called as common-enriched mutations (see Tables 2, 3). A total of 10 primary-specific mutations were observed. Among them, SMARCA4 mutations have been identified as region-specific mutations in the multiregional biopsies of gastric adenomas in our previous study suggesting that SMARCA4 mutations are late events in primary tumors [32], which is consistent with our observation. LN-specific mutations include CTNNB1. Two missense mutations of CTNNB1 were both observed in MSS cases and occurred in known hotspots of the gene (p.S33p/GC7 and p.K335T/GC2) suggesting these LN-specific mutations are likely functional activating events. However, it requires further investigation whether the observed CTNNB1 mutations lead to LN-specific activation of Wnt/β-catenin signaling.
Mutation abundance-based clonal analyses
Mutation allele frequency (MAF) has been used to distinguish clonal from subclonal mutations, and also to infer the evolutionary relationship between the regional biopsies obtained from a single individual [33]. For example, the subclonal architectures reflected by the distribution of MAF of common mutations can be compared between two regional biopsies and to select mutations under selective forces (e.g., mutations that are subclonal in primary tumors but became clonally fixed in LN metastases). For 10 cases (GC1–GC10) with single-matched LN genomes, we compared the MAF of mutations in primary and LN metastasis genomes (Fig. 3). Correlation values were also calculated for the MAF of common mutations between primary and LN genomes. Except for two cases (GC3 and GC9) with relatively small number of common mutations, significant level of correlation was observed between the MAF of common mutations shared by primary and LN genomes. This suggests that the subclonal structures of somatic mutations, at least for the common mutations, are relatively conserved between two separate regional biopsies. This evolutionary pattern is also consistent with the paralleled evolution of multiple regional biopsies we have proposed for the development of synchronous colorectal adenomas and carcinomas [7].
Abundance of somatic mutations in primary and LN genomes. For 10 GC cases, the MAF are shown for mutations as observed in primary genomes (x-axis) and LN genomes (y-axis). Except for two cases with a relative deficit of common mutations (GC3 and GC9), significant level of correlation between the MAFs were observed for common mutations suggesting that the subclonal mutational architecture of primary genomes are relatively conserved in LN genomes
Mutation-based phylogenetic analysis
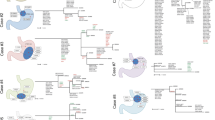
We inferred mutation-based evolutionary relationship of multiple regional biopsies from the same individuals for five cases with primary GC and three metastatic LN genomes available (GC11–GC15; Fig. 4). Of note, a common phylogeny pattern was observed across five cases in which the primary genome is branched from a trunk while all the three LN genomes appear in a cluster of a separate branch. We also observed that three LN samples were most highly related to each other while retaining substantial level of mutation similarities to the primary genome, which is consistent with our previous observation in colorectal cancers [10]. The genomic similarities among multiple metastatic biopsies have been considered as evidence supporting a model of late dissemination where metastatic dissemination has occurred from a single progenitor with primary tumor cells [25]. Thus, we assume that a single subclone or genetically homogeneous subclones in primary tumor mass are responsible for the LN metastases. Of note, we found that the numbers of biopsy-specific mutations for LN genomes are relatively constant in given cases. If the rate of mutation acquisition is relatively constant in given cases [34], it can be assumed that the dissemination of locoregional LN metastases may have occurred in a limited time interval instead of a continuous shedding of metastatic cells from the primary tumor mass.
Discussion
To better understand the mutational processes associated with the tumor invasion and metastases during gastric carcinogenesis, we performed whole-exome sequencing of 15 pairs of primary GC genomes and their matched LN metastases. The evolutionary history of tumors is encrypted in the genomes. Thus, the comparison of genome or exome sequencing data between separate regional biopsies (e.g., primary and LN metastatic lesions) may reveal the evolutionary history or mutational processes that have been operative during carcinogenesis [35, 36].
In terms of the mutation abundance, the primary tumors and their matched LN metastatic GC genomes were comparable for the mutation burdens overall in spite of a substantial level of heterogeneity across the samples and between the matched regions (i.e., the extent of overlap between the primary and LN genomes). The mutation abundance has been proposed as a molecular clock to infer the timing of the development of colorectal and pancreatic tumor progression [37, 38]. Given the abundance of common and region-specific mutations, it can be assumed that the divergence of LN metastatic lesions from primary tumors have occurred earlier for some tumors (e.g., GC7, GC9 and GC14 with relatively abundance region-specific mutations) compared to others. Comparable abundance of primary- and LN-specific mutations also suggests that the after divergence, two lesions have acquired additional region-specific mutations at a similar rate without particular genome events that may have accelerated the mutation acquisition.
Mutation spectra analysis revealed that C > A transversions were frequently observed in LN-specific mutations for a majority of GC cases examined. A previous study reported that the ascitic tumor cells in the GC carcinomatosis peritonei are enriched with C > A transversions [5]. Thus, the C > A transversions may represent the mutations occurring in the LN metastases or malignant ascites after divergence from the primary tumor. This suggests that the mutation forces that are operative in the primary tumors and metastatic lesions are distinct.
Among the recurrent mutations, we observed that the TP53 were specifically observed as common mutations indicative of their early occurrences. As the most common mutations observed in GC genomes, their spatial nature is consistent with the relative timing of TP53 mutation acquisition in gastric carcinogenesis (i.e., adenoma to early differentiated cancers) [39]. The acquisition of region-specific mutations as shown in the example of PIK3CA with available targeted agents, may have clinical impacts because the late-occurring, region-specific mutations may be responsible for the development of drug-resistant clones [40]. In our cases, the region-specific POLE mutations were often observed for those with excessive region-specific mutation rates suggesting that the mutation abundance, as the potential predictors for immune blockage treatment responsiveness [41], may be variable across the regional biopsies. RNF43 is a negative regulator of Wnt signaling, which reduces Frizzled receptor by ubiquitination [42]. RNF43 inactivating mutations by frameshift indels is frequently observed in MSI-H cases and known to be associated with early gastric carcinogenesis [28], which is consistent with our observation that RNF43 mutations were largely observed as common or primary-specific mutations. GNAS mutations were observed in gastric and duodenal benign lesions, such as pyloric gland adenoma and gastric foveolar metaplasia, and gastric and duodenal malignancies [43]. We note that GNAS mutations were observed as common mutations as well as LN-specific mutations, suggesting that GNAS mutations may play a role as early cancer drivers, but may be also associated with the spreading of LN metastases. Further analysis using metastatic lymphatic tissues would be needed to clarify the exact function of GNAS mutations.
Our study has proposed two potential evolutionary models associated with the locoregional LN metastases. First, subclonal mutation architecture for common mutations was conserved between primary and LN genomes. This is suggestive of parallel evolution of multiple tumor regions. This pattern has been previously observed in the comparison of synchronous colorectal adenomas and carcinomas [7]. It is still not clear how primary and LN genomes share subclonal mutations and it is worthy of further investigation for a number of hypotheses previously made (e.g., polyseeding and oligoclonal origin for metastasis) [25]. Second, our phylogenetic analysis using multiple LN genomes may provide some clues on the origins of LN metastases. Because the three LN genomes examined were highly related with each other also showing comparable mutation burdens, it is likely that the LN metastases have occurred in a limited time period from a single subclone (or at least, genetically homogeneous subclones) in the primary mass. However, this assumption should be confirmed in a larger cohort and additional genomic features such as the gene expression and epigenetic profiling, beyond the mutation profiles, should be taken into account. Recent report on colorectal cancers proposed the diverse heterogeneity in terms of DNA methylation highlighting the extent and nature of epigenomic diversity within individual tumors [44].
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Tan P, Yeoh KG. Genetics and molecular pathogenesis of gastric adenocarcinoma. Gastroenterology. 2015;149:1153–62. e3.
Meyerson M, Gabriel S, Getz G. Advances in understanding cancer genomes through second-generation sequencing. Nat Rev Genet. 2010;11:685–96.
Chen K, Yang D, Li X, Sun B, Song F, Cao W, et al. Mutational landscape of gastric adenocarcinoma in Chinese: implications for prognosis and therapy. Proc Natl Acad Sci USA. 2015;112:1107–12.
Lim B, Kim C, Kim JH, Kwon WS, Lee WS, Kim JM, et al. Genetic alterations and their clinical implications in gastric cancer peritoneal carcinomatosis revealed by whole-exome sequencing of malignant ascites. Oncotarget. 2016;7:8055–66.
Zhang J, Huang JY, Chen YN, Yuan F, Zhang H, Yan FH, et al. Whole genome and transcriptome sequencing of matched primary and peritoneal metastatic gastric carcinoma. Sci Rep. 2015;5:13750.
Kim TM, An CH, Rhee JK, Jung SH, Lee SH, Baek IP, et al. Clonal origins and parallel evolution of regionally synchronous colorectal adenoma and carcinoma. Oncotarget. 2015;6:27725–35.
Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K, et al. Clonal architecture of secondary acute myeloid leukemia. N Engl J Med. 2012;366:1090–8.
Choi YJ, Rhee JK, Hur SY, Kim MS, Lee SH, Chung YJ, et al. Intraindividual genomic heterogeneity of high-grade serous carcinoma of the ovary and clinical utility of ascitic cancer cells for mutation profiling. J Pathol. 2017;241:57–66.
Kim TM, Jung SH, An CH, Lee SH, Baek IP, Kim MS, et al. Subclonal genomic architectures of primary and metastatic colorectal cancer based on intratumoral genetic heterogeneity. Clin Cancer Res. 2015;21:4461–72.
Weinberg RA. Mechanisms of malignant progression. Carcinogenesis. 2008;29:1092–5.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–60.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Kim TM, Laird PW, Park PJ. The landscape of microsatellite instability in colorectal and endometrial cancer genomes. Cell. 2013;155:858–68.
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568–76.
Olshen AB, Venkatraman ES, Lucito R, Wigler M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics. 2004;5:557–72.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–6.
Felsenstein J. PHYLIP—phylogeny inference package (version 3.2). Cladistics. 1989;5:164–166.
Kim TM, Jung SH, Baek IP, Lee SH, Choi YJ, Lee JY, et al. Regional biases in mutation screening due to intratumoural heterogeneity of prostate cancer. J Pathol. 2014;233:425–35.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Tomasetti C, Vogelstein B, Parmigiani G. Half or more of the somatic mutations in cancers of self-renewing tissues originate prior to tumor initiation. Proc Natl Acad Sci USA. 2013;110:1999–2004.
Hunter KW, Amin R, Deasy S, Ha NH, Wakefield L. Genetic insights into the morass of metastatic heterogeneity. Nat Rev Cancer. 2018;18:211–23.
Olivier M, Hollstein M, Hainaut P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol. 2010;2:a001008.
Chang MT, Asthana S, Gao SP, Lee BH, Chapman JS, Kandoth C, et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat Biotechnol. 2016;34:155–63.
Min BH, Hwang J, Kim NK, Park G, Kang SY, Ahn S, et al. Dysregulated Wnt signalling and recurrent mutations of the tumour suppressor RNF43 in early gastric carcinogenesis. J Pathol. 2016;240:304–14.
TCGA consortium. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Rayner E, van Gool IC, Palles C, Kearsey SE, Bosse T, Tomlinson I, et al. A panoply of errors: polymerase proofreading domain mutations in cancer. Nat Rev Cancer. 2016;16:71–81.
TCGA consortium. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202–9.
Lim CH, Cho YK, Kim SW, Choi MG, Rhee JK, Chung YJ, et al. The chronological sequence of somatic mutations in early gastric carcinogenesis inferred from multiregion sequencing of gastric adenomas. Oncotarget. 2016;7:39758–67.
Miller CA, White BS, Dees ND, Griffith M, Welch JS, Griffith OL, et al. SciClone: inferring clonal architecture and tracking the spatial and temporal patterns of tumor evolution. PLoS Comput Biol. 2014;10:e1003665.
Shibata D. Mutation and epigenetic molecular clocks in cancer. Carcinogenesis. 2011;32:123–8.
Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366:883–92.
Swanton C. Intratumor heterogeneity: evolution through space and time. Cancer Res. 2012;72:4875–82.
Jones S, Chen WD, Parmigiani G, Diehl F, Beerenwinkel N, Antal T, et al. Comparative lesion sequencing provides insights into tumor evolution. Proc Natl Acad Sci USA. 2008;105:4283–8.
Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B, et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature. 2010;467:1114–7.
Grabsch HI, Tan P. Gastric cancer pathology and underlying molecular mechanisms. Dig Surg. 2013;30:150–8.
Turke AB, Zejnullahu K, Wu YL, Song Y, Dias-Santagata D, Lifshits E, et al. Preexistence and clonal selection of MET amplification in EGFR mutant NSCLC. Cancer Cell. 2010;17:77–88.
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371:2189–99.
Koo BK, Spit M, Jordens I, Low TY, Stange DE, van de Wetering M, et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature. 2012;488:665–9.
Matsubara A, Sekine S, Kushima R, Ogawa R, Taniguchi H, Tsuda H, et al. Frequent GNAS and KRAS mutations in pyloric gland adenoma of the stomach and duodenum. J Pathol. 2013;229:579–87.
Roerink SF, Sasaki N, Lee-Six H, Young MD, Alexandrov LB, Behjati S, et al. Intra-tumour diversification in colorectal cancer at the single-cell level. Nature. 2018;556:457–62.
Acknowledgements
This work was supported by the Korea Health Technology R&D Project via the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant no. HI15C1578 and HI15C3224) and supported by the National Research Foundation of Korea (nos. 2012R1A1A1043576, 2015R1A1A1A05028000 and 2018R1D1A1B07045486); the Research Fund of Seoul St. Mary’s Hospital, The Catholic University of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Human rights statement
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions.
Informed consent
Informed consent or an appropriate substitute for it was obtained from all patients included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, H.H., Kim, S.Y., Jung, E.S. et al. Mutation heterogeneity between primary gastric cancers and their matched lymph node metastases. Gastric Cancer 22, 323–334 (2019). https://doi.org/10.1007/s10120-018-0870-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-018-0870-6