Abstract
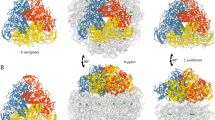
Nitroreductase enzymes are of interest as antibiotic targets and as activators in enzyme prodrug therapy, but the precise substrate binding orientation and reaction mechanism are poorly understood. In order to design more effective antibiotics and improve enzyme prodrug therapy, an atomistic description of nitroaromatic substrate binding in the active Michaelis complex is highly desirable. Here, using an iterative molecular dynamics (MD) simulation protocol, the binding of p-nitrobenzoic acid (p-NBA) in oxidized and reduced Enterobacter cloacae nitroreductase (NR) was investigated. For the oxidized NR, the MD simulations distinguished between the two possible binding orientations of p-NBA in NR from X-ray crystal structure data. For the reduced NR, a distinct active binding orientation of p-NBA was found when the second active site of the NR homodimer was occupied by a NADH analogue. This model of the active Michaelis complex of p-NBA with NR provides a rationale for the reduction of p-NBA by NR via a hydride transfer reaction mechanism suggested by experimental results, and brings the proposed reaction mechanism from experiment and computational models into agreement.





Similar content being viewed by others
References
Lafayette L, Sauter G, Vu L, Meade B (2016) Spartan performance and flexibility: an HPC-cloud chimera. OpenStack Summit, Barcelona, October 27, 2016: https://doi.org/10.4225/4249/4258ead4290dceaaa
Race PR, Lovering, A.L., Green, R.M., Ossor, A., White, S.A., Searle, P.F., Wrighton, C.J., and Hyde E.I. (2005) Structural and mechanistic studies of Escherichia coli nitroreductase with the antibiotic nitrofurazone. J. Biol. Chem. 280 (14):13256–13264
McCalla DR, Kaiser C, Green MHL (1978) Genetics of nitrofurazone resistance in Escherichia-coli. J. Bacteriol. 133(1):10–16
Sastry SS, Jayaraman R (1984) Nitrofurantoin-resistant mutants of Escherichia-coli - isolation and mapping. Mol. Gen. Genet. 196(2):379–380
Weedon SJ, Green NK, McNeish IA, Gilligan MG, Mautner V, Wrighton CJ, Mountain A, Young LS, Kerr DJ, Searle PF (2000) Sensitisation of human carcinoma cells to the prodrug CB1954 by adenovirus vector-mediated expression of E-coli nitroreductase. Int. J. Cancer 86(6):848–854
Grove JI, Searle PF, Weedon SJ, Green NK, McNeish IA, Kerr DJ (1999) Virus-directed enzyme prodrug therapy using CB1954. Anticancer Drug Des. 14(6):461–472
White CL, Menghistu T, Twigger KR, Searle PF, Bhide SA, Vile RG, Melcher AA, Pandha HS, Harrington KJ (2008) Escherichia coli nitroreductase plus CB1954 enhances the effect of radiotherapy in vitro and in vivo. Gene Ther. 15(6):424–433
Denny WA (2002) Nitroreductase-based GDEPT. Curr. Pharm. Des. 8(15):1349–1361. https://doi.org/10.2174/1381612023394584
Anlezark GM, Melton RG, Sherwood RF, Coles B, Friedlos F, Knox RJ (1992) The bioactivation of 5-(aziridin-1-yl)-2,4-dinitrobenzamide (CB1954).1. purification and properties of a nitroreductase enzyme from Escherichia-coli - a potential enzyme for antibody-directed enzyme prodrug therapy (Adept). Biochem. Pharmacol. 44 (12):2289–2295
Lehouritis P, Hogan G, Tangney M (2017) Designer bacteria as intratumoural enzyme biofactories. Adv. Drug Deliv. Rev. 118:8–23. https://doi.org/10.1016/j.addr.2017.09.012
Johansson E, Parkinson GN, Denny WA, Neidle S (2003) Studies on the nitroreductase prodrug-activating system. Crystal structures of complexes with the inhibitor dicoumarol and dinitrobenzamide prodrugs and of the enzyme active form. Journal of Medicinal Chemistry 46:4009–4020
Haynes CA, Koder RL, Miller A-F, Rodgers DW (2002) Structures of nitroreductase in three states. J. Biol. Chem. 277(13):11513–11520. https://doi.org/10.1074/jbc.M111334200
Bryant C, DeLuca M (1991) Purification and characterization of an oxygen-insensitive NAD(P)H nitroreductase from Enterobacter cloacae. J. Biol. Chem. 266(7):4119–4125
Koder RL, Miller A-F (1998) Steady-state kinetic mechanism, stereospecificity, substrate and inhibitor specificity of Enterobacter cloacae nitroreductase. Biochim. Biophys. Acta Protein Struct. Mol. Enzymol. 1387(1):395–405. https://doi.org/10.1016/S0167-4838(98)00151-4
Pitsawong W, Haynes CA, Koder Jr RL, Rodgers DW, Miller A-F (2017) Mechanism-informed refinement reveals altered substrate-binding mode for catalytically competent nitroreductase. Structure 25(7):978–987. https://doi.org/10.1016/j.str.2017.05.002
Barna TM, Khan H, Bruce NC, Barsukov I, Scrutton NS, Moody PCE (2001) Crystal structure of pentaerythritol tetranitrate reductase: “flipped” binding geometries for steroid substrates in different redox states of the enzyme. J. Mol. Biol. 310(2):433–447
Christofferson A, Wilkie J (2009) Mechanism of CB1954 reduction by Escherichia coli nitroreductase. Biochem. Soc. Trans. 37(2):413–418. https://doi.org/10.1042/BST0370413
Isayev O, Crespo-Hernández CE, Gorb L, Hill FC, Leszczynski J (2012) In silico structure–function analysis of E. cloacae nitroreductase. Proteins 80(12):2728–2741. https://doi.org/10.1002/prot.24157
Richard JP (2019) Protein flexibility and stiffness enable efficient enzymatic catalysis. J. Am. Chem. Soc. https://doi.org/10.1021/jacs.8b10836
Lovering AL, Hyde EI, Searle PF, Scott AW (2001) The structure of Escherichia coli nitroreductase complexed with nicotinic acid: three crystal forms at 1.7 Å, 1.8 Å and 2.4 Å resolution. J. Mol. Biol. 309(1):203–213
Christofferson A, Zhao L, Pei Q (2012) Dynamic simulations as a complement to experimental studies of enzyme mechanisms. In: Christov C, Karabencheva-Christova T (eds) Structural and mechanistic enzymology - bringing together experiments and computing, vol 87. Advances in protein chemistry and structural biology. Academic press (Elsevier), San Diego, CA, USA, pp 293-335. https://doi.org/10.1016/B978-0-12-398312-1.00010-X
Jakalian A, Bush BL, Jack DB, Bayly CI (2000) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. method. J. Comput. Chem. 21:132–146
Sviatenko LK, Gorb L, Okovytyy SI, Leszczynski J (2016) Two-electron reduction of nitroaromatic compounds by flavin mononucleotide. DFT computational study. Bulletin of Dnipropetrovsk University series chemistry 24 (1). dx.doi.org/10.15421/081601
Wan S, Knapp B, Wright DW, Deane CM, Coveney PV (2015) Rapid, precise, and reproducible prediction of peptide–MHC binding affinities from molecular dynamics that correlate well with experiment. J. Chem. Theory Comput. 11(7):3346–3356. https://doi.org/10.1021/acs.jctc.5b00179
Bhati AP, Wan S, Hu Y, Sherborne B, Coveney PV (2018) Uncertainty quantification in alchemical free energy methods. J. Chem. Theory Comput. https://doi.org/10.1021/acs.jctc.7b01143
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res. 28:235–242
Case D, Darden T, Cheatham III T, Simmerling C, Wang J, Duke R, Luo R, Walker R, Zhang W, Merz K (2012) AMBER 12. University of California, San Francisco 1 (2):3
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery Jr JA, Peralta JE, Ogliaro F, Bearpark MJ, Heyd J, Brothers EN, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam NJ, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09. Gaussian, Inc., Wallingford, CT, USA
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J. Comput. Chem. 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78(8):1950–1958. https://doi.org/10.1002/prot.22711
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4(3):435–447. https://doi.org/10.1021/ct700301q
Sousa da Silva AW, Vranken WF (2012) ACPYPE - antechamber python parser interface. BMC Research Notes 5(1):367. https://doi.org/10.1186/1756-0500-5-367
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J. Mol. Graph. 14(1):33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Miller BR, McGee TD, Swails JM, Homeyer N, Gohlke H, Roitberg AE (2012) MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8(9):3314–3321. https://doi.org/10.1021/ct300418h
Herpoldt K-L, Artzy-Schnirman A, Christofferson AJ, Makarucha AJ, de la Rica R, Yarovsky I, Stevens MM (2015) Designing fluorescent peptide sensors with dual specificity for the detection of HIV-1 protease. Chem. Mater. 27(20):7187–7195. https://doi.org/10.1021/acs.chemmater.5b03651
Pitsawong W, Hoben JP, Miller A-F (2014) Understanding the broad substrate repertoire of nitroreductase based on its kinetic mechanism. J. Mol. Biol
Zimanyi Christina M, Ando N, Brignole Edward J, Asturias Francisco J, Stubbe J, Drennan Catherine L (2012) Tangled up in knots: structures of inactivated forms of E. coli class Ia ribonucleotide reductase. Structure 20(8):1374–1383. https://doi.org/10.1016/j.str.2012.05.009
Cianci M, Folli C, Zonta F, Florio P, Berni R, Zanotti G (2015) Structural evidence for asymmetric ligand binding to transthyretin. Acta Crystallographica section D 71 (8):1582-1592. https://doi.org/10.1107/S1399004715010585
Acknowledgments
This research was undertaken using the LIEF HPC-GPGPU Facility (resource grant pRMIT0007) hosted at the University of Melbourne [1], which was established with the assistance of LIEF Grant LE170100200. This research was also undertaken with the assistance of resources and services (resource grant nr3) from the National Computational Infrastructure (NCI), which is supported by the Australian Government, and Melbourne Bioinformatics (resource grant RMIT0003). Drs. E. Hyde, S. White, P. Searle, J. Wilkie, P. Charchar, M. Penna, and Q. Besford are gratefully acknowledged for useful discussions, and Drs. E. Hyde, Q. Besford, and Prof. C. F. McConville for their comments on the manuscript. Prof. I. Yarovsky and Prof. C. F. McConville are gratefully acknowledged for their continued support. The anonymous reviewers are also thanked for helpful and constructive comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1691 kb)
Rights and permissions
About this article
Cite this article
Christofferson, A.J. Asymmetric ligand binding in homodimeric Enterobacter cloacae nitroreductase yields the Michaelis complex for nitroaromatic substrates. J Mol Model 26, 28 (2020). https://doi.org/10.1007/s00894-020-4288-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-020-4288-9