Abstract
In the present work we investigated the differential interactions of the antimicrobial peptides (AMPs) aurein 1.2 and maculatin 1.1 with a bilayer composed of a mixture of the lipids 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (POPG) and 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE). We carried out molecular dynamics (MD) simulations using a coarse-grained approach within the MARTINI force field. The POPE/POPG mixture was used as a simple model of a bacterial (prokaryotic cell) membrane. The results were compared with our previous findings for structures of 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), a representative lipid of mammalian cells. We started the simulations of the peptide–lipid system from two different initial conditions: peptides in water and peptides inside the hydrophobic core of the membrane, employing a pre-assembled lipid bilayer in both cases. Our results show similarities and differences regarding the molecular behavior of the peptides in POPE/POPG in comparison to their behavior in a POPC membrane. For instance, aurein 1.2 molecules can adopt similar pore-like structures on both POPG/POPE and POPC membranes, but the peptides are found deeper in the hydrophobic core in the former. Maculatin 1.1 molecules, in turn, achieve very similar structures in both kinds of bilayers: they have a strong tendency to form clusters and induce curvature. Therefore, the results of this study provide insight into the mechanisms of action of these two peptides in membrane leakage, which allows organisms to protect themselves against potentially harmful bacteria.

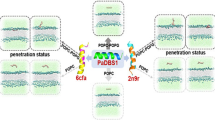
Aurein pore structure (green) in a lipid bilayer composed by POPE (blue) and POPG (red) mixture. It is possible to see water beads (light blue) inside the pore.






Similar content being viewed by others
References
Wang G, Li X, Wang Z (2016) APD3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res 44:D1087–D1093. https://doi.org/10.1093/nar/gkv1278
Reddy KVR, Yedery RD, Aranha C (2004) Antimicrobial peptides: Premises and promises. Int J Antimicrob Agents 24:536–547
Martin E, Ganz T, Lehrer RI (1995) Defensins and other endogenous peptide antibiotics of vertebrates. J Leukoc Biol 58:128–136
Rozek T, Wegener KL, Bowie JH et al (2000) The antibiotic and anticancer active aurein peptides from the Australian bell frogs Litoria aurea and Litoria raniformis: the solution structure of aurein 1.2. Eur J Biochem 267:5330–5341. https://doi.org/10.1046/j.1432-1327.2000.01536.x
Hoskin DW, Ramamoorthy A (2008) Studies on anticancer activities of antimicrobial peptides. Biochim Biophys Acta Biomembr 1778:357–375
de la Fuente-Núñez C, Cardoso MH, de Souza CE et al (2016) Synthetic antibiofilm peptides. Biochim Biophys Acta Biomembr 1858:1061–1069. https://doi.org/10.1016/j.bbamem.2015.12.015
Hilchie AL, Wuerth K, Hancock REW (2013) Immune modulation by multifaceted cationic host defense (antimicrobial) peptides. Nat Chem Biol 9:761–768. https://doi.org/10.1038/nchembio.1393
Mansour SC, Pena OM, Hancock REW (2014) Host defense peptides: Front-line immunomodulators. Trends Immunol 35:443–450
Guilhelmelli F, Vilela N, Albuquerque P et al (2013) Antibiotic development challenges: the various mechanisms of action of antimicrobial peptides and of bacterial resistance. Front Microbiol 4:353
Dathe M, Wieprecht T (1999) Structural features of helical antimicrobial peptides: their potential to modulate activity on model membranes and biological cells. Biochim Biophys Acta Biomembr 1462:71–87. https://doi.org/10.1016/S0005-2736(99)00201-1
Huang Y, Huang J, Chen Y (2010) Alpha-helical cationic antimicrobial peptides: relationships of structure and function. Protein Cell 1:143–152. https://doi.org/10.1007/s13238-010-0004-3
REW H, Rozek A (2002) Role of membranes in the activities of antimicrobial cationic peptides. FEMS Microbiol Lett 206:143–149
Jenssen H, Hamill P, Hancock REW (2006) Peptide antimicrobial agents. Clin Microbiol Rev 19:491–511
Tossi A, Sandri L, Giangaspero A (2000) Amphipathic, alpha-helical antimicrobial peptides. Biopolymers 55:4–30. https://doi.org/10.1002/1097-0282(2000)55:1<4::AID-BIP30>3.0.CO;2-M
Zelezetsky I, Tossi A (2006) Alpha-helical antimicrobial peptides—using a sequence template to guide structure–activity relationship studies. Biochim Biophys Acta Biomembr 1758:1436–1449
Sato H, Feix JB (2006) Peptide–membrane interactions and mechanisms of membrane destruction by amphipathic α-helical antimicrobial peptides. Biochim Biophys Acta Biomembr 1758:1245–1256. https://doi.org/10.1016/j.bbamem.2006.02.021
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3:238–250. https://doi.org/10.1038/nrmicro1098
Bahar AA, Ren D (2013) Antimicrobial peptides. Pharmaceuticals 6:1543–1575. https://doi.org/10.3390/ph6121543
Mihajlovic M, Lazaridis T (2010) Antimicrobial peptides in toroidal and cylindrical pores. Biochim Biophys Acta Biomembr 1798:1485–1493. https://doi.org/10.1016/j.bbamem.2010.04.004
Dowhan W (1997) Molecular basis for membrane phospholipid diversity: why are there so many lipids? Annu Rev Biochem 66:199–232. https://doi.org/10.1146/annurev.biochem.66.1.199
Goldfine H (1972) Comparative aspects of bacterial lipids. Adv Microb Physiol 8:1–58. https://doi.org/10.1016/S0065-2911(08)60187-3
Randle CL, Albro PW, Dittmer JC (1969) The phosphoglyceride composition of gram-negative bacteria and the changes in composition during growth. Biochim Biophys Acta Lipids Lipid Metab 187:214–220. https://doi.org/10.1016/0005-2760(69)90030-7
Staudegger E, Prenner EJ, Kriechbaum M et al (2000) X-ray studies on the interaction of the antimicrobial peptide gramicidin S with microbial lipid extracts: evidence for cubic phase formation. Biochim Biophys Acta Biomembr 1468:213–230. https://doi.org/10.1016/S0005-2736(00)00260-1
Von Deuster CIE, Knecht V (2011) Competing interactions for antimicrobial selectivity based on charge complementarity. Biochim Biophys Acta Biomembr 1808:2867–2876. https://doi.org/10.1016/j.bbamem.2011.08.005
Chia BCS, Carver JA, Mulhern TD, Bowie JH (2000) Maculatin 1.1, an anti-microbial peptide from the Australian tree frog, Litoria genimaculata. Solution structure and biological activity. Eur J Biochem 267:1894–1908. https://doi.org/10.1046/j.1432-1327.2000.01089.x
Balatti GE, Ambroggio EE, Fidelio GD et al (2017) Differential interaction of antimicrobial peptides with lipid structures studied by coarse-grained molecular dynamics simulations. Molecules 22:1775. https://doi.org/10.3390/molecules22101775
Hopp TP, Woods KR (1981) Prediction of protein antigenic determinants from amino acid sequences. Immunology 78:3824–3828. https://doi.org/10.1073/pnas.78.6.3824
Ambroggio EE, Separovic F, Bowie JH et al (2005) Direct visualization of membrane leakage induced by the antibiotic peptides: maculatin, citropin, and aurein. Biophys J 89:1874–1881. https://doi.org/10.1529/biophysj.105.066589
Gehman JD, Luc F, Hall K et al (2008) Effect of antimicrobial peptides from Australian tree frogs on anionic phospholipid membranes. Biochemistry 47:8557–8565. https://doi.org/10.1021/bi800320v
Marcotte I, Wegener KL, Lam YH et al (2003) Interaction of antimicrobial peptides from Australian amphibians with lipid membranes. Chem Phys Lipids 122:107–120. https://doi.org/10.1016/S0009-3084(02)00182-2
Bond PJ, Parton DL, Clark JF, Sansom MSP (2008) Coarse-grained simulations of the membrane-active antimicrobial peptide maculatin 1.1. Biophys J 95:3802–3815. https://doi.org/10.1529/biophysj.108.128686
Matsuzaki K, Sugishita KI, Ishibe N et al (1998) Relationship of membrane curvature to the formation of pores by magainin 2. Biochemistry 37:11856–11863. https://doi.org/10.1021/bi980539y
Schmidt NW, Wong GCL (2013) Antimicrobial peptides and induced membrane curvature: geometry, coordination chemistry, and molecular engineering. Curr Opin Solid State Mater Sci 17:151–163. https://doi.org/10.1016/j.cossms.2013.09.004
da Hora GCA, Archilha NL, Lopes JLS et al (2016) Membrane negative curvature induced by a hybrid peptide from pediocin PA-1 and plantaricin 149 as revealed by atomistic molecular dynamics simulations. Soft Matter 12:8884–8898. https://doi.org/10.1039/C6SM01714B
Shahmiri M, Enciso M, Mechler A (2015) Controls and constrains of the membrane disrupting action of Aurein 1.2. Sci Rep 5:16378. https://doi.org/10.1038/srep16378
Lorenzón EN, Sanches PRS, Nogueira LG et al (2013) Dimerization of aurein 1.2: effects in structure, antimicrobial activity and aggregation of Candida albicans cells. Amino Acids 44:1521–1528. https://doi.org/10.1007/s00726-013-1475-3
Lorenzón EN, Piccoli JP, Cilli EM (2014) Interaction between the antimicrobial peptide Aurein 1.2 dimer and mannans. Amino Acids 46:2627–2631. https://doi.org/10.1007/s00726-014-1832-x
Abraham MJ, Murtola T, Schulz R et al (2015) Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Van Der Spoel D, Lindahl E, Hess B et al (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Lindahl E, Hess B, van der Spoel D (2001) GROMACS 3.0: a package for molecular simulation and trajectory analysis. J Mol Model 7:306–317. https://doi.org/10.1007/s008940100045
Pronk S, Páll S, Schulz R et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854. https://doi.org/10.1093/bioinformatics/btt055
Hess B, Kutzner C, Van Der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447. https://doi.org/10.1021/ct700301q
Marrink SJ, de Vries AH, Mark AE (2004) Coarse grained model for semiquantitative lipid simulations. J Phys Chem B 108:750–760. https://doi.org/10.1021/jp036508g
Marrink SJ, Risselada HJ, Yefimov S et al (2007) The MARTINI forcefield: coarse grained model for biomolecular simulations. J Phys Chem B 111:7812–7824
De Jong DH, Singh G, Bennett WFD et al (2013) Improved parameters for the MARTINI coarse-grained protein force field. J Chem Theory Comput 9:687–697. https://doi.org/10.1021/ct300646g
Yesylevskyy SO, Schäfer LV, Sengupta D, Marrink SJ (2010) Polarizable water model for the coarse-grained MARTINI force field. PLoS Comput Biol 6:1–17. https://doi.org/10.1371/journal.pcbi.1000810
Berman HM, Westbrook J, Feng Z et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Wang G, Li Y, Li X (2005) Correlation of three-dimensional structures with the antibacterial activity of a group of peptides designed based on a nontoxic bacterial membrane anchor. J Biol Chem 280:5803–5811. https://doi.org/10.1074/jbc.M410116200
Uggerhøj LE, Munk JK, Hansen PR et al (2014) Structural features of peptoid-peptide hybrids in lipid–water interfaces. FEBS Lett 588:3291–3297. https://doi.org/10.1016/j.febslet.2014.07.016
Schrödinger, LLC (2015) The PyMol molecular graphics system, version 1.8. Schrödinger, LLC, New York
Joosten RP, Te Beek TAH, Krieger E et al (2011) A series of PDB related databases for everyday needs. Nucleic Acids Res 39(Suppl 1):D411–D419. https://doi.org/10.1093/nar/gkq1105
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. https://doi.org/10.1002/bip.360221211
Lee J, Jung SW, Cho AE (2016) Molecular insights into the adsorption mechanism of human β-defensin-3 on bacterial membranes. Langmuir 32:1782–1790. https://doi.org/10.1021/acs.langmuir.5b04113
Wassenaar TA, Ingólfsson HI, Böckmann RA et al (2015) Computational lipidomics with insane: a versatile tool for generating custom membranes for molecular simulations. J Chem Theory Comput 11:2144–2155. https://doi.org/10.1021/acs.jctc.5b00209
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126(1):014101. https://doi.org/10.1063/1.2408420
Parrinello M (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182. https://doi.org/10.1063/1.328693
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Stambulchik E (2018) Grace homepage. http://plasma-gate.weizmann.ac.il/Grace/
Marrink SJ, Tieleman DP (2013) Perspective on the MARTINI model. Chem Soc Rev 42:6801. https://doi.org/10.1039/c3cs60093a
Bennett WFD, Tieleman DP (2011) Water defect and pore formation in atomistic and coarse-grained lipid membranes: pushing the limits of coarse graining. J Chem Theory Comput 7:2981–2988. https://doi.org/10.1021/ct200291v
Hugo AA, Tymczyszyn EE, Gómez-Zavaglia A, Pérez PF (2012) Effect of human defensins on lactobacilli and liposomes. J Appl Microbiol 113:1491–1497. https://doi.org/10.1111/j.1365-2672.2012.05433.x
Acknowledgements
This work was developed with financial support from Universidad de Buenos Aires (no. UBACYT 20020130200096BA), CONICET (nos. 0131-2014), and ANPCyT (nos. PICT-2014-3653, 2013-1205). M.F.M. and M.P. are career members of CONICET.
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper belongs to Topical Collection XIX - Brazilian Symposium of Theoretical Chemistry (SBQT2017)
Electronic supplementary material
ESM 1
(PDF 3355 kb)
Rights and permissions
About this article
Cite this article
Balatti, G.E., Martini, M.F. & Pickholz, M. A coarse-grained approach to studying the interactions of the antimicrobial peptides aurein 1.2 and maculatin 1.1 with POPG/POPE lipid mixtures. J Mol Model 24, 208 (2018). https://doi.org/10.1007/s00894-018-3747-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3747-z