Abstract
Proteins are an important component of living systems and play a crucial role in various physiological functions. Fluorescence imaging of proteins is a powerful tool for monitoring protein dynamics. Fluorescent protein (FP)-based labeling methods are frequently used to monitor the movement and interaction of cellular proteins. However, alternative methods have also been developed that allow the use of synthetic fluorescent probes to target a protein of interest (POI). Synthetic fluorescent probes have various advantages over FP-based labeling methods. They are smaller in size than the fluorescent proteins, offer a wide variety of colors and have improved photochemical properties. There are various chemical recognition-based labeling techniques that can be used for labeling a POI with a synthetic probe. In this review, we focus on the development of protein-labeling systems, particularly the SNAP-tag, BL-tag, and PYP-tag systems, and understanding the fluorescence behavior of the fluorescently labeled target protein in these systems. We also discuss the smart fluorogenic probes for these protein-labeling systems and their applications. The fluorogenic protein labeling will be a useful tool to investigate complex biological phenomena in future work on cell biology.
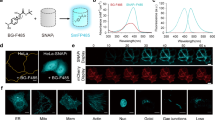
Graphical abstract











Similar content being viewed by others
References
Marks KM, Nolan GP (2006) Chemical labeling strategies for cell biology. Nat Methods 3:591–596
Lippincott-Schwartz J, Patterson GH (2003) Development and use of fluorescent protein markers in living cells. Science 300:87–91
Shimomura O, Johnson FH, Saiga Y (1962) Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Comp Physiol 59:223–239
Prasher DC, Eckenrode VK, Ward WW, Prendergast FG, Cormier MJ (1992) Primary structure of the Aequorea victoria green-fluorescent protein. Gene 111:229–233
Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC (1994) Green fluorescent protein as a marker for gene expression. Science 263:802–805
Shaner NC, Steinbach PA, Tsien RY (2005) A guide to choosing fluorescent proteins. Nat Methods 2:905–909
Zhang J, Campbell RE, Ting AY, Tsien RY (2002) Creating new fluorescent probes for cell biology. Nat Rev Mol Cell Biol 3:906–918
Kremers G-J, Gilbert SG, Cranfill PJ, Davidson MW, Piston DW (2011) Fluorescent proteins at a glance. J Cell Sci 124:157–160
Chudakov DM, Matz MV, Lukyanov S, Lukyanov KA (2010) Fluorescent proteins and their applications in imaging living cells and tissues. Physiol Rev 90:1103–1163
Ormo M, Cubitt AB, Kallio K, Gross LA, Tsien RY, Remington SJ (1996) Crystal structure of the aequorea victoria green fluorescent protein. Science 273:1392–1395
Craggs TD (2009) Green fluorescent protein: structure, folding and chromophore maturation. Chem Soc Rev 38:2865–2875
Kwok EY, Hanson MR (2004) In vivo analysis of interactions between GFP-labeled microfilaments and plastid stromules. BMC Plant Biol 4:2
Naumov GN, Wilson SM, MacDonald IC, Schmidt EE, Morris VL, Groom AC, Hoffman RM, Chambers AF (1999) Cellular expression of green fluorescent protein, coupled with high-resolution in vivo video microscopy, to monitor steps in tumor metastasis. J Cell Sci 112:1835–1842
Giepmans BNG, Adams SR, Ellisman MH, Tsien RY (2006) The fluorescent toolbox for assessing protein location and function. Science 312:217–224
Chen Y, Tsao K, Keillor JW (2015) Fluorogenic protein labelling: a review of photophysical quench mechanisms and principles of fluorogen design. Can J Chem 93:389–398
Jung D, Min K, Jung J, Jang W, Kwon Y (2013) Chemical biology-based approaches on fluorescent labeling of proteins in live cells. Mol BioSyst 9:862–872
Mizukami S, Hori Y, Kikuchi K (2013) Small-molecule-based protein-labeling technology in live cell studies: probe-design concepts and applications. Acc Chem Res 47:247–256
Spicer CD, Davis BG (2014) Selective chemical protein modification. Nat Commun 5:4740
Griffin BA, Adams SR, Tsien RY (1998) Specific covalent labeling of recombinant protein molecules inside live cells. Science 281:269–272
Los GV, Encell LP, McDougall MG, Hartzell DD, Karassina N, Zimprich C, Wood MG, Learish R, Ohana RF, Urh M, Simpson D (2008) HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem Biol 3:373–382
Keppler A, Kindermann M, Gendreizig S, Pick H, Vogel H, Johnsson K (2004) Labeling of fusion proteins of O6-alkylguanine-DNA alkyltransferase with small molecules in vivo and in vitro. Methods 32:437–444
Dean KM, Palmer AE (2014) Advances in fluorescence labeling strategies for dynamic cellular imaging. Nat Chem Biol 10:512–523
Zhang Y, Park K-Y, Suazo KF, Distefano MD (2018) Recent progress in enzymatic protein labelling techniques and their applications. Chem Soc Rev 47:9106–9136
Griffin BA, Adams SR, Jones J, Tsien RY (2000) Fluorescent labeling of recombinant proteins in living cells with FlAsH. Methods Enzymol 327:565–578
Sletten EM, Bertozzi CR (2009) Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew Chem Int Ed 48:6974–6998
Miller LW, Cai Y, Sheetz MP, Cornish VW (2005) In vivo protein labeling with trimethoprim conjugates: a flexible chemical tag. Nat Methods 2:255–257
Bonger KM, Chen L-C, Liu CW, Wandless TJ (2011) Small-molecule displacement of a cryptic degron causes conditional protein degradation. Nat Chem Biol 7:531–537
Cheeseman IM, Desai A (2005) A combined approach for the localization and tandem affinity purification of protein complexes from metazoans. Sci STKE 2005:pl–pl5
Nesbeth D, Williams SL, Chan L, Brain T, Slater NK, Farzaneh F, Darling D (2006) Metabolic biotinylation of lentiviral pseudotypes for scalable paramagnetic microparticle-dependent manipulation. Mol Ther 13:814–822
Kosa NM, Haushalter RW, Smith AR, Burkart MD (2012) Reversible labeling of native and fusion-protein motifs. Nat Methods 9:981–984
Gautier A, Juillerat A, Heinis C, Corrêa IR Jr, Kindermann M, Beaufils F, Johnsson K (2008) An engineered protein tag for multiprotein labeling in living cells. Chem Biol 15:128–136
Hori Y, Ueno H, Mizukami S, Kikuchi K (2009) Photoactive yellow protein-based protein labeling system with turn-on fluorescence intensity. J Am Chem Soc 131:16610–16611
Mizukami S, Watanabe S, Hori Y, Kikuchi K (2009) Covalent protein labeling based on noncatalytic β-lactamase and a designed FRET substrate. J Am Chem Soc 131:5016–5017
Pegg AE (2000) Repair of O 6-alkylguanine by alkyltransferases. Mutat Res 462:83–100
Damoiseaux R, Keppler A, Johnsson K (2001) Synthesis and applications of chemical probes for human O 6-alkylguanine-DNA alkyltransferase. Chem Bio Chem 2:285–287
Keppler A, Gendreizig S, Gronemeyer T, Pick H, Vogel H, Johnsson K (2003) A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat Biotechnol 21:86–89
Kalderon D, Roberts BL, Richardson WD, Smith AE (1984) A short amino acid sequence able to specify nuclear location. Cell 39:499–509
Juillerat A, Gronemeyer T, Keppler A, Gendreizig S, Pick H, Vogel H, Johnsson K (2003) Directed evolution of O 6-alkylguanine-DNA alkyltransferase for efficient labeling of fusion proteins with small molecules in vivo. Chem Biol 10:313–317
Wibley JE, Pegg AE, Moody PC (2000) Crystal structure of the human O(6)-alkylguanine-DNA alkyltransferase. Nucleic Acids Res 28:393–401
Keppler A, Pick H, Arrivoli C, Vogel H, Johnsson K (2004) Labeling of fusion proteins with synthetic fluorophores in live cells. Proc Natl Acad Sci USA 101:9955–9959
Stöhr K, Siegberg D, Ehrhard T, Lymperopoulos K, Öz S, Schulmeister S, Pfeifer AC, Bachmann J, Klingmüller U, Sourjik V, Herten DP (2010) Quenched substrates for live-cell labeling of SNAP-tagged fusion proteins with improved fluorescent background. Anal Chem 82:8186–8193
Zhang C-J, Li L, Chen GYJ, Xu Q-H, Yao SQ (2011) One- and two-photon live cell imaging using a mutant SNAP-tag protein and its FRET substrate pairs. Org Lett 13:4160–4163
Komatsu T, Johnsson K, Okuno H, Bito H, Inoue T, Nagano T, Urano Y (2011) Real-time measurements of protein dynamics using fluorescence activation-coupled protein labeling method. J Am Chem Soc 133:6745–6751
Sun X, Zhang A, Baker B, Sun L, Howard A, Buswell J, Maurel D, Masharina A, Johnsson K, Noren CJ, Xu MQ, CorrÞa IR Jr (2011) Development of SNAP-tag fluorogenic probes for wash free fluorescence imaging. Chem Bio Chem 12:2217–2226
Lukinavičius G, Umezawa K, Olivier N, Honigmann A, Yang G, Plass T, Mueller V, Reymond L, Corrêa IR Jr, Luo Z-G, Schultz C, Lemke EA, Heppenstall P, Eggeling C, Manley S, Johnsson K (2013) A near-infrared fluorophore for live-cell superresolution microscopy of cellular proteins. Nat Chem 5:132–139
Westphal V, Rizzoli SO, Lauterbach MA, Kamin D, Jahn R, Hell SW (2008) Video-rate far-field optical nanoscopy dissects synaptic vesicle movement. Science 320:246–249
Sadhu KK, Mizukami S, Hori Y, Kikuchi K (2011) Switching modulation for protein labeling with activatable fluorescent probes. Chem Bio Chem 12:1299–1308
Sutcliffe JG (1978) Nucleotide sequence of the ampicillin resistance gene of Escherichia coli plasmid pBR322. Proc Natl Acad Sci USA 75:3737–3741
Zlokarnik G, Negulescu PA, Knapp TE, Mere L, Burres N, Feng L, Whitney M, Roemer K, Tsien RY (1998) Quantitation of transcription and clonal selection of single living cells with β-lactamase as reporter. Science 279:84–88
Adachi H, Ohta T, Matsuzawa H (1991) Site-directed mutants, at position 166, of RTEM-1 β-lactamase that form a stable acyl-enzyme intermediate with penicillin. J Biol Chem 266:3186–3191
Guillaume G, Vanhove M, Lamotte-Brasseur J, Ledent P, Jamin M, Joris B, Frére J-M (1997) Site-directed mutagenesis of glutamate 166 in two β-lactamases. J Biol Chem 272:5438–5444
Sadhu KK, Mizukami S, Watanabe S, Kikuchi K (2010) Turn-on fluorescence switch involving aggregation and elimination processes for b-lactamase-tag. Chem Commun 46:7403–7405
Mizukami S, Watanabe S, Akimoto Y, Kikuchi K (2012) No-wash protein labeling with designed fluorogenic probes and application to real-time pulse-chase analysis. J Am Chem Soc 134:1623–1629
Watanabe S, Mizukami S, Hori Y, Kikuchi K (2010) Multicolor protein labeling in living cells using mutant β-lactamase-tag technology. Bioconjugate Chem 21:2320–2326
Watanabe S, Mizukami S, Akimoto Y, Hori Y, Kikuchi K (2011) Intracellular Protein labeling with prodrug-like probes using a mutant β-lactamase tag. Chem Eur J 17:8342–8349
Sato R, Kozuka J, Ueda M, Mishima R, Kumagai Y, Yoshimura A, Minoshima M, Mizukami S, Kikuchi K (2017) Intracellular protein-labeling probes for multicolor single-molecule imaging of immune receptor–adaptor molecular dynamics. J Am Chem Soc 139:17397–17404
Zhou Z, Koglin A, Wang Y, McMahon AP, Walsh CT (2008) An eight residue fragment of an acyl carrier protein suffices for post-translational introduction of fluorescent pantetheinyl arms in protein modification in vitro and in vivo. J Am Chem Soc 130:9925–9930
Kamiuchi M, Hara MT, Stalcup P, Xie A, Hoff WD (2008) Identification of six new photoactive yellow proteins—diversity and structure-function relationships in a bacterial blue light photoreceptor. Photochem Photobiol 84:956–969
Kyndt JA, Meyer TE, Cusanovich MA, Van Beeumen JJ (2002) Characterization of a bacterial tyrosine ammonia lyase, a biosynthetic enzyme for the photoactive yellow protein. FEBS Lett 512:240–244
Hori Y, Nakaki K, Sato M, Mizukami S, Kikuchi K (2012) Development of protein-labeling probes with a redesigned fluorogenic switch based on intramolecular association for no-wash live-cell imaging. Angew Chem Int Ed 51:5611–5614
Hori Y, Norinobu T, Sato M, Arita K, Shirakawa M, Kikuchi K (2013) Development of fluorogenic probes for quick no-wash live-cell imaging of intracellular proteins. J Am Chem Soc 135:12360–12365
Hori Y, Hirayama S, Sato M, Kikuchi K (2015) Redesign of a fluorogenic labeling system to improve surface charge, brightness, and binding kinetics for imaging the functional localization of bromodomains. Angew Chem Int Ed 54:14368–14371
Hirayama S, Hori Y, Benedek Z, Suzuki T, Kikuchi K (2016) Fluorogenic probes reveal a role of GLUT4N-glycosylation in intracellular trafficking. Nat Chem Biol 12:853–859
Kumar N, Hori Y, Kikuchi K (2018) Live-cell imaging of DNA methylation based on synthetic-molecule/protein hybrid probe. Chem Rec 18:1672–1680
Hori Y, Otomura N, Nishida A, Nishiura M, Umeno M, Suetake I, Kikuchi K (2018) Synthetic-molecule/protein hybrid probe with fluorogenic switch for live-cell imaging of DNA methylation. J Am Chem Soc 140:1686–1690
Acknowledgements
This research was partly supported by the Grant-in-Aid for Scientific Research (Grant no. JP16K01933 to M.M., JP18H03935 to K.K. JP17H02210, JP18K19402 to Y. H.), Innovative Areas “Frontier Research on Chemical Communications” (No. JP17H06409) and “Resonance Bio” (No. JP18H04735) of Ministry of Education, Culture, Sports, Science, and Technology (MEXT) of Japan; AMED (Nos. 18he0902005h0004, 17ae0101041h9902, 18fm0208018h0002 to K.K.); JSPS A3 Foresight Program; JSPS CORE-to-CORE Program “Asian Chemical Biology Initiative”; SICORP from JST, and the Asahi Glass Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Reja, S.I., Minoshima, M., Hori, Y. et al. Development of an effective protein-labeling system based on smart fluorogenic probes. J Biol Inorg Chem 24, 443–455 (2019). https://doi.org/10.1007/s00775-019-01669-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01669-y