Abstract
Hemoproteins are commonly found in nature, and involved in many important cellular processes such as oxygen transport, electron transfer, and catalysis. Rational design of hemoproteins can not only inspire novel biocatalysts but will also lead to a better understanding of structure–function relationships in native hemoproteins. Here, the heme nitric oxide/oxygen-binding protein from Caldanaerobacter subterraneus subsp. tengcongensis (TtH-NOX) is used as a novel scaffold for oxidation biocatalyst design. We show that signaling protein TtH-NOX can be reengineered to catalyze H2O2 decomposition and oxidation of 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) by H2O2. In addition, the role of the distal tyrosine (Tyr140) in catalysis is investigated. The mutation of Tyr140 to alanine hinders the catalysis of the oxidation reactions. On the other hand, the mutation of Tyr140 to histidine, which is commonly observed in peroxidases, leads to a significant increase of the catalytic activity. Taken together, these results show that, while the distal histidine plays an important role in hemoprotein reactions with H2O2, it is not always essential for oxidation activity. We show that TtH-NOX protein can be used as an alternative scaffold for the design of novel biocatalysts with desired reactivity or functionality.
Graphical abstract
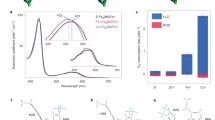
H-NOX proteins are homologous to the nitric oxide sensor soluble guanylate cyclase. Here, we show that the gas sensor protein TtH-NOX shows limited capacity for catalysis of redox reactions and it can be used as a novel scaffold in biocatalysis design.










Similar content being viewed by others
Abbreviations
- H-NOX:
-
Heme nitric oxide/oxygen binding
- TtH-NOX:
-
Heme domain of the methyl-accepting chemotaxis protein from Caldanaerobacter subterraneus subsp. tengcongensis
- Mb:
-
Myoglobin
- HRP:
-
Horseradish peroxidase
- ABTS:
-
2,2′-Azino-bis(3-ethylbenzothiazoline-6-sulphonic acid)
- WT:
-
Wild type
- SDM:
-
Site-directed mutagenesis
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- ALA:
-
5-Aminolevulinic acid hydrochloride
References
Poulos TL (2014) Heme enzyme structure and function. Chem Rev 114:3919–3962. https://doi.org/10.1021/cr400415k
Olson JS, Eich RF, Smith LP et al (1997) Protein engineering strategies for designing more stable hemoglobin-based blood substitutes. Artif Cells Blood Substit Biotechnol 25:227–241. https://doi.org/10.3109/10731199709118912
Isaac IS, Dawson JH (1999) Haem iron-containing peroxidases. Essays Biochem 34:51–69
Park SY, Shimizu H, Adachi S et al (1997) Crystal structure of nitric oxide reductase from denitrifying fungus Fusarium oxysporum. Nat Struct Biol 4:827–832
Pellequer J-L, Brudler R, Getzoff ED (1999) Biological sensors: more than one way to sense oxygen. Curr Biol 9:R416–R418. https://doi.org/10.1016/S0960-9822(99)80257-7
Jain R, Chan MK (2003) Mechanisms of ligand discrimination by heme proteins. J Biol Inorg Chem 8:1–11. https://doi.org/10.1007/s00775-002-0405-8
Ozaki S, Matsui T, Roach MP, Watanabe Y (2000) Rational molecular design of a catalytic site: engineering of catalytic functions to the myoglobin active site framework. Coord Chem Rev 198:39–59. https://doi.org/10.1016/S0010-8545(00)00234-4
Lu Y, Berry SM, Pfister TD (2001) Engineering novel metalloproteins: design of metal-binding sites into native protein scaffolds. Chem Rev 101:3047–3080. https://doi.org/10.1021/cr0000574
Colonna S, Gaggero N, Richelmi C, Pasta P (1999) Recent biotechnological developments in the use of peroxidases. Trends Biotechnol 17:163–168. https://doi.org/10.1016/S0167-7799(98)01288-8
Alvarado B, Torres E (2009) Recents patents in the use of peroxidases. Recent Pat Biotechnol 3:88–102
Schonbaum GR, Chance B (1976) The enzymes. Catalase 7:363–408
Matsui T, Ozaki S, Watanabe Y (1999) Formation and catalytic roles of compound I in the hydrogen peroxide-dependent oxidations by His64 myoglobin mutants. J Am Chem Soc 121:9952–9957. https://doi.org/10.1021/ja9914846
Battistuzzi G, Bellei M, Bortolotti CA, Sola M (2010) Redox properties of heme peroxidases. Arch Biochem Biophys 500:21–36. https://doi.org/10.1016/j.abb.2010.03.002
Yu F, Cangelosi VM, Zastrow ML et al (2014) Protein design: toward functional metalloenzymes. Chem Rev 114:3495–3578. https://doi.org/10.1021/cr400458x
Chand S, Ray S, Wanigasekara E et al (2018) Improved rate of substrate oxidation catalyzed by genetically-engineered myoglobin. Arch Biochem Biophys 639:44–51. https://doi.org/10.1016/j.abb.2017.12.014
Matsuo T, Fukumoto K, Watanabe T, Hayashi T (2011) Precise design of artificial cofactors for enhancing peroxidase activity of myoglobin: myoglobin mutant H64D reconstituted with a “single-winged cofactor” is equivalent to native horseradish peroxidase in oxidation activity. Chemistry 6:2491–2499. https://doi.org/10.1002/asia.201100107
Sato H, Hayashi T, Ando T et al (2004) Hybridization of modified-heme reconstitution and distal histidine mutation to functionalize sperm whale myoglobin. J Am Chem Soc 126:436–437. https://doi.org/10.1021/ja038798k
Iyer LM, Anantharaman V, Aravind L (2003) Ancient conserved domains shared by animal soluble guanylyl cyclases and bacterial signaling proteins. BMC Genom 4:5. https://doi.org/10.1186/1471-2164-4-5
Schmidt PM, Schramm M, Schröder H et al (2004) Identification of residues crucially involved in the binding of the heme moiety of soluble guanylate cyclase. J Biol Chem 279:3025–3032. https://doi.org/10.1074/jbc.M310141200
Karow DS, Pan D, Tran R et al (2004) Spectroscopic characterization of the soluble guanylate cyclase-like heme domains from Vibrio cholerae and Thermoanaerobacter tengcongensis. Biochemistry 43:10203–10211. https://doi.org/10.1021/bi049374l
Pellicena P, Karow DS, Boon EM et al (2004) Crystal structure of an oxygen-binding heme domain related to soluble guanylate cyclases. Proc Natl Acad Sci USA 101:12854–12859. https://doi.org/10.1073/pnas.0405188101
Martin E, Berka V, Bogatenkova E et al (2006) Ligand selectivity of soluble guanylyl cyclase: effect of the hydrogen-bonding tyrosine in the distal heme pocket on binding of oxygen, nitric oxide, and carbon monoxide. J Biol Chem 281:27836–27845. https://doi.org/10.1074/jbc.M601078200
Olea C, Boon EM, Pellicena P et al (2008) Probing the function of heme distortion in the H-NOX family. ACS Chem Biol 3:703–710. https://doi.org/10.1021/cb800185h
Winter MB, McLaurin EJ, Reece SY et al (2010) Ru-porphyrin protein scaffolds for sensing O2. J Am Chem Soc 132:5582–5583. https://doi.org/10.1021/ja101527r
Poulos TL, Sheriff S, Howard AJ (1987) Cocrystals of yeast cytochrome c peroxidase and horse heart cytochrome c. J Biol Chem 262:13881–13884
Dawson JH (1988) Probing structure-function relations in heme-containing oxygenases and peroxidases. Science 240:433–439
Poulos T, Kraut J (1980) A hypothetical model of the cytochrome c peroxidase cytochrome c electron transfer complex. J Biol Chem 255:10322–10330
Gajhede M, Schuller DJ, Henriksen A et al (1997) Crystal structure of horseradish peroxidase C at 2.15 A resolution. Nat Struct Biol 4:1032–1038
Phillips GN, Arduini RM, Springer BA, Sligar SG (1990) Crystal structure of myoglobin from a synthetic gene. Proteins 7:358–365. https://doi.org/10.1002/prot.340070407
Weinert EE, Plate L, Whited CA et al (2010) Determinants of ligand affinity and heme reactivity in H-NOX domains. Angew Chemie Int Ed 49:720–723. https://doi.org/10.1002/anie.200904799
Barr I, Guo F (2015) Pyridine hemochromagen assay for determining the concentration of heme in purified protein solutions. Bio Protoc 5:7–20. https://doi.org/10.21769/BioProtoc.1594
Re R, Pellegrini N, Proteggente A et al (1999) Antioxidant activity applying an improved ABTS radical cation decolorization assay. Free Radic Biol Med 26:1231–1237
Chance B, Maehly AC (1955) [136] Assay of catalases and peroxidases. Methods Enzymol 2:764–775. https://doi.org/10.1016/S0076-6879(55)02300-8
Boon EME, Huang SH, Marletta MAMMA (2005) A molecular basis for NO selectivity in soluble guanylate cyclase. Nat Chem Biol 1:53–59. https://doi.org/10.1038/nchembio704
Boon EM, Marletta MA (2005) Ligand discrimination in soluble guanylate cyclase and the H-NOX family of heme sensor proteins. Curr Opin Chem Biol 9:441–446. https://doi.org/10.1016/j.cbpa.2005.08.015
Dai Z, Boon EM (2010) Engineering of the Heme pocket of an H-NOX domain for direct cyanide detection and quantification. J Am Chem Soc 132:11496–11503. https://doi.org/10.1021/ja101674z
Richards MP (2013) Redox reactions of myoglobin. Antioxid Redox Signal 18:2342–2351. https://doi.org/10.1089/ars.2012.4887
Childs BRE, Bardsley WG (1975) The steady-state kinetics of peroxidase with 2,2′-azino-di-(3-ethylbenzothiazoline- 6-sulphonic acid) as chromogen. Biochem J 145:93–103. https://doi.org/10.1042/bj1450093
Hespen CW, Bruegger JJ, Phillips-Piro CM, Marletta MA (2016) Structural and functional evidence indicates selective oxygen signaling in Caldanaerobacter subterraneus H-NOX. ACS Chem Biol 11:2337–2346. https://doi.org/10.1021/acschembio.6b00431
Weinert EE, Phillips-Piro CM, Marletta MA (2013) Porphyrin π-stacking in a heme protein scaffold tunes gas ligand affinity. J Inorg Biochem 127:7–12. https://doi.org/10.1016/j.jinorgbio.2013.06.004
Winter MB, Klemm PJ, Phillips-Piro CM et al (2013) Porphyrin-substituted H-NOX proteins as high-relaxivity MRI contrast agents. Inorg Chem 52:2277–2279. https://doi.org/10.1021/ic302685h
Winter MB, Herzik MA, Kuriyan J, Marletta MA (2011) Tunnels modulate ligand flux in a heme nitric oxide/oxygen binding (H-NOX) domain. Proc Natl Acad Sci 108:E881–E889. https://doi.org/10.1073/pnas.1114038108
Winter MB, Woodward JJ, Marletta MA (2013) An Escherichia coli expression-based approach for porphyrin substitution in heme proteins. Methods Mol Biol 987:95–106. https://doi.org/10.1007/978-1-62703-321-3-8
Kosowicz JG, Boon EM (2013) Insights into the distal heme pocket of H-NOX using fluoride as a probe for H-bonding interactions. J Inorg Biochem 126:91–95. https://doi.org/10.1016/j.jinorgbio.2013.05.012
Tran R, Weinert EE, Boon EM et al (2011) Determinants of the heme–CO vibrational modes in the H-NOX family. Biochemistry 50:6519–6530
Brantley RE, Smerdon SJ, Wilkinson AJ et al (1993) The mechanism of autooxidation of myoglobin. J Biol Chem 268:6995–7010
Yeh L-H, Alayash AI (2003) Redox side reactions of haemoglobin and cell signalling mechanisms. J Intern Med 253:518–526
Yonetani T, Schleyer H (1969) Studies on cytochrome c peroxidase IX. The reaction of ferrimyoglobin with hydroperoxides and a comparison of peroxide-induced compounds of ferrimyoglobin and cytochrome c peroxidase. J Biol Chem 242:1947–1979
Khan KK, Mondal MS, Padhy L, Mitra S (1998) The role of distal histidine in peroxidase activity of myoglobin. Eur J Biochem 257:547–555. https://doi.org/10.1046/j.1432-1327.1998.2570547.x
Edwards SL, Raag R, Wariishi H et al (1993) Crystal structure of lignin peroxidase. Proc Natl Acad Sci USA 90:750–754. https://doi.org/10.1006/jmbi.1998.2507
Erman JE, Vitello LB, Miller MA et al (1993) Histidine 52 is a critical residue for rapid formation of cytochrome c peroxidase compound I. Biochemistry 32:9798–9806. https://doi.org/10.1021/bi00088a035
Newmyer SL, Ortiz de Montellano PR (1995) Horseradish peroxidase His-42 → Ala, His-42 → Val, and Phe-41 → Ala mutants. Histidine catalysis and control of substrate access to the heme iron. J Biol Chem 270:19430–19438. https://doi.org/10.1074/jbc.270.33.19430
Matsui T, Ozaki SI, Liong E et al (1999) Effects of the location of distal histidine in the reaction of myoglobin with hydrogen peroxide. J Biol Chem 274:2838–2844. https://doi.org/10.1074/jbc.274.5.2838
Giulivi C, Davies KJA (1994) [30] Hydrogen peroxide-mediated ferrylhemoglobin generation in vitro and in red blood cells. Methods Enzymol 231:490–496. https://doi.org/10.1016/0076-6879(94)31032-7
Savenkova MI, Kuo JM, Ortiz de Montellano PR (1998) Improvement of peroxygenase activity by relocation of a catalytic histidine within the active site of horseradish peroxidase. Biochemistry 37:10828–10836. https://doi.org/10.1021/bi9725780
Li QS, Ogawa J, Shimizu S (2001) Critical role of the residue size at position 87 in H2O2-dependent substrate hydroxylation activity and H2O2 inactivation of cytochrome P450BM-3. Biochem Biophys Res Commun 280:1258–1261. https://doi.org/10.1006/bbrc.2001.4261
Cirino PC, Arnold FH (2003) A self-sufficient peroxide-driven hydroxylation biocatalyst. Angew Chem Int Ed Engl 42:3299–3301. https://doi.org/10.1002/anie.200351434
Carlsen CU, Skovgaard IM, Skibsted LH (2003) Pseudoperoxidase activity of myoglobin: kinetics and mechanism of the peroxidase cycle of myoglobin with H2O2 and 2,2-azino-bis(3-ethylbenzothiazoline-6-sulfonate) as substrates. J Agric Food Chem 51:5815–5823. https://doi.org/10.1021/jf030067g
George P, Hanania G (1952) The Ionization of acidic metmyoglobin. Biochem J 52:517–523. https://doi.org/10.1042/bj0650756
Svistunenko DA, Sharpe MA, Nicholls P et al (2000) The pH dependence of naturally occurring low-spin forms of methaemoglobin and metmyoglobin: an EPR study. Biochem J 351:595–605. https://doi.org/10.1042/0264-6021:3510595
Olea C, Kuriyan J, Marletta MA (2010) Modulating heme redox potential through protein-induced porphyrin distortion. J Am Chem Soc 132:12794–12795. https://doi.org/10.1021/ja106252b
Acknowledgements
This work was supported by The Scientific and Technological Research Council of Turkey (TUBİTAK, 115C134). We thank Dr. Michael A. Marletta for the generous gift of pET20b plasmid encoding WT TtH-NOX. We thank Dr. Eric S. Underbakke for critical reading of the manuscript. J.E.A-F. received financial support from Turkiye Scholarships. We thank Belgin Tunçel Kırkar and the Department of Chemical Engineering at Izmir Institute of Technology for their help with the glovebox instrument. We are thankful to Biotechnology and Bioengineering Application and Research Center at Izmir Institute of Technology for DNA sequencing analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aggrey-Fynn, J.E., Surmeli, N.B. A novel thermophilic hemoprotein scaffold for rational design of biocatalysts. J Biol Inorg Chem 23, 1295–1307 (2018). https://doi.org/10.1007/s00775-018-1615-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-018-1615-z