Abstract
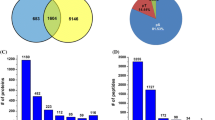
Cytoplasmic male sterility (CMS) is widely used in plant breeding and represents a perfect model to understand cyto-nuclear interactions and pollen development research. Protein phosphorylation is ubiquitous and is involved in the regulation of diverse cellular processes. To reveal the possible mechanism of CMS and pollen development in kenaf, we performed an iTRAQ-based comparative phosphoproteome analysis in the anthers of a CMS line and wild-type plant (Wt). Whole transcriptome unigenes of kenaf as the reference genome, we identified a total of 3045 phosphorylated sites on 1640 peptides corresponding to 974 unique proteins. 292 of the peptides which corresponding to 247 unique proteins were differentially phosphorylated (fold change ≥ 1.20 with P value< 0.05) between these two materials. 113 and 134 proteins were characterized as up-regulated or down-regulated phosphorylated, respectively. An evaluation of the phosphoproteome and proteomic results indicated that the most significantly phosphorylated proteins were not associated with abundant changes at the protein level. Bioinformatics analysis demonstrated that many of these proteins were involved in various biological processes which may play key roles in pollen development, including carbohydrate metabolism, energy metabolism, transport, gene expression regulation, signal transduction, and cell cycle control. Our results provide insight into the CMS mechanism and pollen development in kenaf from a protein phosphorylation perspective.







Similar content being viewed by others
References
Baxter CJ, Foyer CH, Turner J, Rolfe SA, Quick WP (2003) Elevated sucrose-phosphate synthase activity in transgenic tobacco sustains photosynthesis in older leaves and alters development. J Exp Bot 54(389):1813–1820. https://doi.org/10.1093/jxb/erg196
Baxter A, Mittler R, Suzuki N (2014) ROS as key players in plant stress signalling. J Exp Bot 65(5):1229–1240. https://doi.org/10.1093/jxb/ert375
Bots M, Vergeldt F, Wolters-Arts M, Weterings K, van As H, Mariani C (2005) Aquaporins of the PIP2 class are required for efficient anther dehiscence in tobacco. Plant Physiol 137(3):1049
Bourbon HM, Aguilera A, Ansari AZ, Asturias FJ, Berk AJ, Bjorklund S, Blackwell TK, Borggrefe T, Carey M, Carlson M, Conaway JW, Conaway RC, Emmons SW, Fondell JD, Freedman LP, Fukasawa T, Gustafsson CM, Han M, He X, Herman PK, Hinnebusch AG, Holmberg S, Holstege FC, Jaehning JA, Kim Y-J, Kuras L, Leutz A, Lis JT, Meisterernest M, Naar AM, Nasmyth K, Parvin JD, Ptashne M, Reinberg D, Ronne H, Sadowski I, Sakurai H, Sipiczki M, Sternberg PW, Stillman DJ, Strich R, Struhl K, Svejstrup JQ, Tuck S, Winston F, Roeder RG, Kornberg RD (2004) A unified nomenclature for protein subunits of mediator complexes linking transcriptional regulators to RNA polymerase II. Mol Cell 14(5):553–557. https://doi.org/10.1016/j.molcel.2004.05.011
Chao Q, Gao ZF, Wang YF, Li Z, Huang XH, Wang YC, Mei YC, Zhao BG, Li L, Jiang YB, Wang BC (2016) The proteome and phosphoproteome of maize pollen uncovers fertility candidate proteins. Plant Mol Biol 91(3):287–304. https://doi.org/10.1007/s11103-016-0466-7
Chase CD (2007) Cytoplasmic male sterility: a window to the world of plant mitochondrial–nuclear interactions. Trends Genet 23(2):81–90. https://doi.org/10.1016/j.tig.2006.12.004
Chen P, Ran SM, Li R, Huang ZP, Qian JH, Yu ML, Zhou RY (2014) Transcriptome de novo assembly and differentially expressed genes related to cytoplasmic male sterility in kenaf (Hibiscus cannabinus L.). Mol Breed 34(4):1879–1891. https://doi.org/10.1007/s11032-014-0146-8
Chou MF, Schwartz D (2002) Biological sequence motif discovery using motif-x. Curr Protoc Bioinform. https://doi.org/10.1002/0471250953.bi1315s35
Christensen AR, Malcomber ST (2012) Duplication and diversification of the LEAFY HULL STERILE1 and Oryza sativa MADS5 SEPALLATA lineages in graminoid Poales. EvoDevo 3(1):4. https://doi.org/10.1186/2041-9139-3-4
Clement C, Burrus M, Audran JC (1996) Floral organ growth and carbohydrate content during pollen development in lilium. Am J Bot 83(4):459–469. https://doi.org/10.2307/2446215
Di Giorgio JAP, Bienert GP, Ayub ND, Yaneff A, Barberini ML, Mecchia MA, Amodeo G, Soto GC, Muschietti JP (2016) Pollen-specific aquaporins NIP4;1 and NIP4;2 are required for pollen development and pollination in Arabidopsis thaliana. Plant Cell 28(5):1053
Dorion S, Lalonde S, Saini HS (1996) Induction of male sterility in wheat by meiotic-stage water deficit is preceded by a decline in invertase activity and changes in carbohydrate metabolism in anthers. Plant Physiol 111(1):137
Dowd PE, Coursol S, Skirpan AL, Kao TH, Gilroy S (2006) Petunia phospholipase C1 is involved in pollen tube growth. Plant Cell 18(6):1438
Fíla J, Radau S, Matros A, Hartmann A, Scholz U, Feciková J, Mock H-P, Čapková V, Zahedi RP, Honys D (2016) Phosphoproteomics profiling of tobacco mature pollen and pollen activated in vitro. Mol Cell Proteom 15(4):1338–1350. https://doi.org/10.1074/mcp.m115.051672
Fujii S, Toriyama K (2008) DCW11, down-regulated gene 11 in CW-type cytoplasmic male sterile rice, encoding mitochondrial protein phosphatase 2C is related to cytoplasmic male sterility. Plant Cell Physiol 49(4):633–640. https://doi.org/10.1093/pcp/pcn036
Guan Y, Meng X, Khanna R, LaMontagne E, Liu Y, Zhang S (2014) Phosphorylation of a WRKY transcription factor by MAPKs is required for pollen development and function in Arabidopsis. PLoS Genet 10(5):e1004384. https://doi.org/10.1371/journal.pgen.1004384
Hepler PK, Kunkel JG, Rounds CM, Winship LJ (2012) Calcium entry into pollen tubes. Trends Plant Sci 17(1):32–38. https://doi.org/10.1016/j.tplants.2011.10.007
Hirakawa T, Hasegawa J, White CI, Matsunaga S (2017) RAD54 forms DNA repair foci in response to DNA damage in living plant cells. Plant J 90(2):372–382. https://doi.org/10.1111/tpj.13499
Holmes-Davis R, Tanaka CK, Vensel WH, Hurkman WJ, McCormick S (2005) Proteome mapping of mature pollen of Arabidopsis thaliana. Proteomics 5(18):4864–4884. https://doi.org/10.1002/pmic.200402011
Hubbard MJ, Cohen P (1993) On target with a new mechanism for the regulation of protein phosphorylation. Trends Biochem Sci 18(5):172–177. https://doi.org/10.1016/0968-0004(93)90109-Z
Hutvagner G, Simard MJ (2008) Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol 9(1):22
Ischebeck T, Valledor L, Lyon D, Gingl S, Nagler M, Meijón M, Egelhofer V, Weckwerth W (2014) Comprehensive cell-specific protein analysis in early and late pollen development from diploid microsporocytes to pollen tube growth. Mol Cell Proteomics 13(1):295–310. https://doi.org/10.1074/mcp.m113.028100
Jagadish SVK, Cairns J, Lafitte R, Wheeler TR, Price AH, Craufurd PQ (2010) Genetic Analysis of Heat Tolerance at Anthesis in Rice. Crop Sci 50(5):1633–1641. https://doi.org/10.2135/cropsci2009.09.0516
Jorrin-Novo JV (2014) Plant proteomics methods and protocols. In: Jorrin-Novo JV, Komatsu S, Weckwerth W, Wienkoop S (eds) Plant proteomics: methods and protocols. Humana Press, Totowa, pp 3–13. https://doi.org/10.1007/978-1-62703-631-3_1
Kassavetis GA, Soragni E, Driscoll R, Geiduschek EP (2005) Reconfiguring the connectivity of a multiprotein complex: fusions of yeast TATA-binding protein with Brf1, and the function of transcription factor IIIB. Proc Natl Acad Sci 102(43):15406
Kersten B, Agrawal GK, Durek P, Neigenfind J, Schulze W, Walther D, Rakwal R (2009) Plant phosphoproteomics: an update. Proteomics 9(4):964–988. https://doi.org/10.1002/pmic.200800548
Kuromori T, Ito T, Sugimoto E, Shinozaki K (2011) Arabidopsis mutant of AtABCG26, an ABC transporter gene, is defective in pollen maturation. J Plant Physiol 168(16):2001–2005. https://doi.org/10.1016/j.jplph.2011.05.014
Laporte MM, Galagan JA, Prasch AL, Vanderveer PJ, Hanson DT, Shewmaker CK, Sharkey TD (2001) Promoter strength and tissue specificity effects on growth of tomato plants transformed with maize sucrose-phosphate synthase. Planta 212(5):817–822. https://doi.org/10.1007/s004250000433
Lemoine R, Bürkle L, Barker L, Sakr S, Kühn C, Regnacq M, Gaillard C, Delrot S, Frommer WB (1999) Identification of a pollen-specific sucrose transporter-like protein NtSUT3 from tobacco. FEBS Lett 454(3):325–330. https://doi.org/10.1016/S0014-5793(99)00843-1
Li WQ, Zhang XQ, Xia C, Deng Y, Ye D (2010) Male gametophyte defective 1, encoding the FAd subunit of mitochondrial F1F0-ATP synthase, is essential for pollen formation in Arabidopsis thaliana. Plant Cell Physiol 51(6):923–935. https://doi.org/10.1093/pcp/pcq066
Liu J, Pang C, Wei H, Song M, Meng Y, Ma J, Fan S, Yu S (2015) iTRAQ-facilitated proteomic profiling of anthers from a photosensitive male sterile mutant and wild-type cotton (Gossypium hirsutum L.). J Proteom 126:68–81. https://doi.org/10.1016/j.jprot.2015.05.031
Ma H (2005) Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu Rev Plant Biol 56(1):393–434. https://doi.org/10.1146/annurev.arplant.55.031903.141717
Mann M, Ong SE, Grønborg M, Steen H, Jensen ON, Pandey A (2002) Analysis of protein phosphorylation using mass spectrometry: deciphering the phosphoproteome. Trends Biotechnol 20(6):261–268. https://doi.org/10.1016/S0167-7799(02)01944-3
Mascarenhas JP (1990) Gene activity during pollen development. Ann Rev Plant Physiol Mol Biol 41:317–338
Maurel C, Javot H, Lauvergeat V, Gerbeau P, Tournaire C, Santoni V, Heyes J (2002) Molecular physiology of aquaporins in plants. Int Rev Cytol 215:105–148. https://doi.org/10.1016/S0074-7696(02)15007-8
Mayank P, Grossman J, Wuest S, Boisson-Dernier A, Roschitzki B, Nanni P, Nühse T, Grossniklaus U (2012) Characterization of the phosphoproteome of mature Arabidopsis pollen. Plant J 72(1):89–101. https://doi.org/10.1111/j.1365-313X.2012.05061.x
Mei S, Fei W, Zhou S (2011) Gene ontology based transfer learning for protein subcellular localization. BMC Bioinform 12(1):44. https://doi.org/10.1186/1471-2105-12-44
Meyuhas O (2008) Physiological roles of ribosomal protein S6: one of its kind. Int Rev Cell Mol Biol 268:1–37. https://doi.org/10.1016/S1937-6448(08)00801-0
Muñoz-Bertomeu J, Cascales-Miñana B, Irles-Segura A, Mateu I, Nunes-Nesi A, Fernie AR, Segura J, Ros R (2010) The plastidial glyceraldehyde-3-phosphate dehydrogenase is critical for viable pollen development in Arabidopsis. Plant Physiol 152(4):1830
Ngounou Wetie AG, Sokolowska I, Woods AG, Roy U, Deinhardt K, Darie CC (2014) Protein–protein interactions: switch from classical methods to proteomics and bioinformatics-based approaches. Cell Mol Life Sci 71(2):205–228. https://doi.org/10.1007/s00018-013-1333-1
Ozlu N, Akten B, Timm W, Haseley N, Steen H, Steen JA (2010) Phosphoproteomics. Wiley interdisciplinary reviews systems biology and medicine 2(3):255–276. https://doi.org/10.1002/wsbm.41
Pacini E, Guarnieri M, Nepi M (2006) Pollen carbohydrates and water content during development, presentation, and dispersal: a short review. Protoplasma 228(1):73. https://doi.org/10.1007/s00709-006-0169-z
Palancade B, Bensaude O (2003) Investigating RNA polymerase II carboxyl-terminal domain (CTD) phosphorylation. Eur J Biochem 270(19):3859–3870. https://doi.org/10.1046/j.1432-1033.2003.03794.x
Park JY, Canam T, Kang KY, Ellis DD, Mansfield SD (2008) Over-expression of an Arabidopsis family A sucrose phosphate synthase (SPS) gene alters plant growth and fibre development. Transgenic Res 17(2):181–192. https://doi.org/10.1007/s11248-007-9090-2
Qin P, Tu B, Wang Y, Deng L, Quilichini TD, Li T, Wang H, Ma B, Li S (2013) ABCG15 encodes an ABC transporter protein, and is essential for post-meiotic anther and pollen exine development in rice. Plant Cell Physiol 54(1):138–154. https://doi.org/10.1093/pcp/pcs162
Quévillon HM, Zeitler DM, Hauptmann J, Bruckmann A, Fressigné L, Danner J, Piquet S, Strieder N, Engelmann JC, Jannot G, Deutzmann R, Simard MJ, Meister G (2017) Phosphorylation of argonaute proteins affects mRNA binding and is essential for microRNA-guided gene silencing. EMBO J. https://doi.org/10.15252/embj.201696386
Ramesh M (2016) Kenaf (Hibiscus cannabinus L.) fibre based bio-materials: a review on processing and properties. Prog Mater Sci 78–79:1–92. https://doi.org/10.1016/j.pmatsci.2015.11.001
Ravagnan L, Gurbuxani S, Susin SA, Maisse C, Daugas E, Zamzami N, Mak T, Jäättelä M, Penninger JM, Garrido C, Kroemer G (2001) Heat-shock protein 70 antagonizes apoptosis-inducing factor. Nat Cell Biol 3:839. https://doi.org/10.1038/ncb0901-839
Sakamoto T, Inui YT, Uraguchi S, Yoshizumi T, Matsunaga S, Mastui M, Umeda M, Fukui K, Fujiwara T (2011) Condensin II alleviates DNA damage and is essential for tolerance of boron overload stress in Arabidopsis. Plant Cell 23(9):3533
Sarhadi E, Bazargani MM, Sajise AG, Abdolahi S, Vispo NA, Arceta M, Nejad GM, Singh RK, Salekdeh GH (2012) Proteomic analysis of rice anthers under salt stress. Plant Physiol Biochem 58:280–287. https://doi.org/10.1016/j.plaphy.2012.07.013
Sheoran IS, Olson DJH, Ross ARS, Sawhney VK (2005) Proteome analysis of embryo and endosperm from germinating tomato seeds. Proteomics 5(14):3752–3764. https://doi.org/10.1002/pmic.200401209
Sheoran IS, Ross ARS, Olson DJH, Sawhney VK (2009) Differential expression of proteins in the wild type and 7B-1 male-sterile mutant anthers of tomato (Solanum lycopersicum): a proteomic analysis. J Proteom 71(6):624–636. https://doi.org/10.1016/j.jprot.2008.10.006
Shin SB, Golovkin M, Reddy ASN (2014) A pollen-specific calmodulin-binding protein, NPG1, interacts with putative pectate lyases. Sci Rep 4:5263. https://doi.org/10.1038/srep05263
Shumyantseva VV, Suprun EV, Bulko TV, Archakov AI (2014) Electrochemical methods for detection of post-translational modifications of proteins. Biosens Bioelectr 61:131–139. https://doi.org/10.1016/j.bios.2014.05.001
Silva-Sanchez C, Li H, Chen S (2015) Recent advances and challenges in plant phosphoproteomics. Proteomics 15(5–6):1127–1141. https://doi.org/10.1002/pmic.201400410
Sjogren CA, Bolaris SC, Larsen PB (2015) Aluminum-dependent terminal differentiation of the Arabidopsis root tip is mediated through an ATR-, ALT2-, and SOG1-regulated transcriptional response. Plant Cell 27(9):2501
Song L, Liu Z, Tong J, Xiao L, Ma H, Zhang H (2015) Comparative proteomics analysis reveals the mechanism of fertility alternation of thermosensitive genic male sterile rice lines under low temperature inducement. Proteomics 15(11):1884–1905. https://doi.org/10.1002/pmic.201400103
Thingholm TE, Jorgensen TJD, Jensen ON, Larsen (2006) Highly selective enrichment of phosphorylated peptides using titanium dioxide. Nat Protoc 1(4):1929
Truernit E, Stadler R, Baier K, Sauer N (1999) A male gametophyte-specific monosaccharide transporter in Arabidopsis. Plant J 17(2):191–201. https://doi.org/10.1046/j.1365-313x.1999.00372.x
van Wijk KJ, Friso G, Walther D, Schulze WX (2014) Meta-analysis of Arabidopsis thaliana phospho-proteomics data reveals compartmentalization of phosphorylation motifs. Plant Cell 26(6):2367–2389
Wang K, Zhao Y, Li M, Gao F, M-k Yang, Wang X, Li S, Yang P (2014) Analysis of phosphoproteome in rice pistil. Proteomics 14(20):2319–2334. https://doi.org/10.1002/pmic.201400004
Waterworth WM, Drury GE, Bray CM, West CE (2011) Repairing breaks in the plant genome: the importance of keeping it together. New Phytol 192(4):805–822. https://doi.org/10.1111/j.1469-8137.2011.03926.x
Williams LE, Lemoine R, Sauer N (2000) Sugar transporters in higher plants—a diversity of roles and complex regulation. Trends Plant Sci 5(7):283–290. https://doi.org/10.1016/S1360-1385(00)01681-2
Wisniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Meth 6(5):359–362. http://www.nature.com/nmeth/journal/v6/n5/suppinfo/nmeth.1322_S1.html
Xie HT, Wan ZY, Li S, Zhang Y (2014) Spatiotemporal production of reactive oxygen species by NADPH oxidase is critical for tapetal programmed cell death and pollen development in Arabidopsis. Plant Cell 26(5):2007–2023. https://doi.org/10.1105/tpc.114.125427
Yang CW, González-Lamothe R, Ewan RA, Rowland O, Yoshioka H, Shenton M, Ye H, Donnell E, Jones JDG, Sadanandom A (2006) The E3 ubiquitin ligase activity of Arabidopsis; plant U-BOX17 and its functional tobacco homolog ACRE276 are required for cell death and defense. Plant Cell 18(4):1084
Yao SG, Ohmori S, Kimizu M, Yoshida H (2008) Unequal genetic redundancy of rice PISTILLATA orthologs, OsMADS2 and OsMADS4, in lodicule and stamen development. Plant Cell Physiol 49(5):853–857. https://doi.org/10.1093/pcp/pcn050
Ye J (2015) Proteomic and phosphoproteomic analyses reveal extensive phosphorylation of regulatory proteins in developing rice anthers. Plant J 84(3):527–544. https://doi.org/10.1111/tpj.13019
Ye J, Zhang Z, Long H, Zhang Z, Hong Y, Zhang X, You C, Liang W, Ma H, Lu P (2015) Proteomic and phosphoproteomic analyses reveal extensive phosphorylation of regulatory proteins in developing rice anthers. Plant J 84(3):527–544. https://doi.org/10.1111/tpj.13019
Ye J, Zhang Z, You C, Zhang X, Lu J, Ma H (2016) Abundant protein phosphorylation potentially regulates Arabidopsis anther development. J Exp Bot 67(17):4993–5008. https://doi.org/10.1093/jxb/erw293
Yin LL, Xue HW (2012) The MADS29 transcription factor regulates the degradation of the nucellus and the nucellar projection during rice seed development. Plant Cell 24(3):1049
Yoshiyama K, Sakaguchi K, Kimura S (2013) DNA damage response in plants: conserved and variable response compared to animals. Biology 2(4):1338
Zhai Q, Yan L, Tan D, Chen R, Sun J, Gao L, Dong M-Q, Wang Y, Li C (2013) Phosphorylation-coupled proteolysis of the transcription factor MYC2 is important for jasmonate-signaled plant immunity. PLoS Genet 9(4):e1003422. https://doi.org/10.1371/journal.pgen.1003422
Zhang H, Liang W, Yang X, Luo X, Jiang N, Ma H, Zhang D (2010) Carbon starved anther encodes a MYB domain protein that regulates sugar partitioning required for rice pollen development. Plant Cell 22(3):672
Zhang M, Lv D, Ge P, Bian Y, Chen G, Zhu G, Li X, Yan Y (2014) Phosphoproteome analysis reveals new drought response and defense mechanisms of seedling leaves in bread wheat (Triticum aestivum L.). J Proteom 109:290–308. https://doi.org/10.1016/j.jprot.2014.07.010
Zhao Z, Assmann SM (2011) The glycolytic enzyme, phosphoglycerate mutase, has critical roles in stomatal movement, vegetative growth, and pollen production in Arabidopsis thaliana. J Exp Bot 62(14):5179–5189. https://doi.org/10.1093/jxb/err223
Zhou RY, Zhang X, Zhang JQ, Wei HX (2008) A breakthrough in kenaf cytoplasmic male sterile lines breeding and heterosis utilization. Sci Agric Sin 41(1):314 (in Chinese)
Zhu LM, Ai SY, Zhou RY (2007) A cytological study on microsporogenesis of cytoplasmic male sterile lines in kenaf (Hibiscus cannabinus L.). Acta Agronom Sinica 33(6):999–1003 (in Chinse)
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant no. 31560341) and the earmarked fund for Modern Agro-industry Technology Research System (CARS-16-E14). We thank LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they do not have any commercial or associative interest that represents a conflict of interest in connection with the work submitted.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, P., Li, R. & Zhou, R. Comparative phosphoproteomic analysis reveals differentially phosphorylated proteins regulate anther and pollen development in kenaf cytoplasmic male sterility line. Amino Acids 50, 841–862 (2018). https://doi.org/10.1007/s00726-018-2564-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-018-2564-0