Abstract
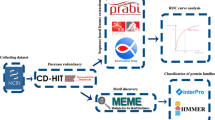
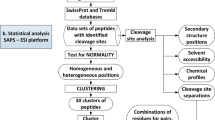
Seryl-histidine dipeptide (Ser-His) has been recognized as the shortest peptide with hydrolysis cleavage activity; however, its protein cleavage spectrum has not yet been fully explored. Here, four differently folded proteins were treated with Ser-His, and the digestion products were evaluated with high-resolution mass spectrometry. The cleavage efficiency and cleavage propensity of Ser-His against these protein substrates were calculated at both the primary and secondary sequence levels. The above experiments show that Ser-His cleaves a broad spectrum of substrate proteins of varying secondary structures. Moreover, Ser-His could cleave at all 20 amino acids with different efficiencies according to the protein, which means that Ser-His has the original digestion function of serine proteases. Furthermore, we collected and compared the catalytic sites and cleavage sites of 340 extant serine proteases derived from 17 representative organisms. A consensus motif Ser-[X]-His was identified as the major pattern at the catalytic sites of serine proteases from all of the organisms represented except Danio rerio, which uses Ser-Lys instead. This finding indicates that Ser-His is the core component of the serine protease catalytic site. Moreover, our analysis revealed that the cleavage sites of modern serine proteases have become more specific over the evolutionary history of this family. Based on the above analysis results, it could be found that Ser-His is likely the original serine protease and maybe the evolutionary core of modern serine proteases.




Similar content being viewed by others
References
Adamala K, Szostak JW (2013) Competition between model protocells driven by an encapsulated catalyst. Nat Chem 5(7):495–501
Andersson MK, Karlson U, Hellman L (2008) The extended cleavage specificity of the rodent beta-chymases rMCP-1 and mMCP-4 reveal major functional similarities to the human mast cell chymase. Mol Immunol 45(3):766–775
Bairoch A, Apweiler R (2000) The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res 28(1):45–48
Barker WC, Dayhoff MO (1980) Evolutionary and functional-relationships of homologous physiological-mechanisms. Bioscience 30(9):593–600
Bartlett GJ, Porter CT, Borkakoti N et al (2002) Analysis of catalytic residues in enzyme active sites. J Mol Biol 324(1):105–121
Brady L, Brzozowski AM, Derewenda ZS et al (1990) A serine protease triad forms the catalytic center of a triacylglycerol lipase. Nature 343(6260):767–770
Chen J, Wan R, Liu H et al (2000) Cleavage of BSA by a dipeptide seryl-histidine. Lett Pept Sci 7(6):325–329
Chen J, Wan R, Liu H et al (2001) Studies on the cleavage of bovine serum albumin by Ser-His. Chem J Chin Univ Chin 22(8):1349–1351
Contreras JA, Karlsson M, Osterlund T et al (1996) Hormone-sensitive lipase is structurally related to acetylcholinesterase, bile salt-stimulated lipase, and several fungal lipases—building of a three-dimensional model for the catalytic domain of hormone-sensitive lipase. J Biol Chem 271(49):31426–31430
Deng WK, Wang YB, Liu ZX et al (2014) HemI: a toolkit for Illustrating Heatmaps. Plos One 9(11):e111988
Du HL, Wang YT, Yang LF et al (2002) Appraisal of green fluorescent protein as a model substrate for seryl-histidine dipeptide cleaving agent. Lett Pept Sci 9(1):5–10
Gorlero M, Wieczorek R, Adamala K et al (2009) Ser-His catalyses the formation of peptides and PNAs. FEBS Lett 583(1):153–156
Hedstrom L (2002) Serine protease mechanism and specificity. Chem Rev 102(12):4501–4523
Jakubke HD, Kuhl P, Konnecke A (1985) Basic principles of protease-catalyzed peptide-bond formation. Angew Chem Int Edn Engl 24(2):85–93
Li YS, Zhao YF, Hatfield S et al (2000) Dipeptide seryl-histidine and related oligopeptides cleave DNA, protein, and a carboxyl ester. Bioorg Med Chem 8(12):2675–2680
Li YS, Hatfield S, Li J et al (2002) Seryl-histidine as an alternative DNA nicking agent in nick translation yields superior DNA probes and hybridizations. Bioorg Med Chem 10(3):667–673
Liu XX, Zhang JX, Ni F et al (2010a) Genome wide exploration of the origin and evolution of amino acids. BMC Evol Biol 10(1):1–11
Liu Y, Shi Y, Liu X et al (2010b) Evaluation of non-covalent interaction between Seryl-Histidine dipeptide and cyclophilin A using NMR and molecular modeling. Sci China Chem 53(9):1987–1993
Luthi P, Luisi PL (1984) Enzymatic-synthesis of hydrocarbon-soluble peptides with reverse micelles. J Am Chem Soc 106(23):7285–7286
Ma Y, Zhao Y (1997) Different function of cysteine and serine residue on RNA and DNA. Phosphorus Sulfur Silicon 120(1):447–448
Magrane M, Consortium U (2011) UniProt knowledgebase: a hub of integrated protein data. Database (Oxford). 2011:bar009. doi:10.1093/database/bar009
Otte KB, Hauer B (2015) Enzyme engineering in the context of novel pathways and products. Curr Opin Biotechnol 35C:16–22
Perkins DN, Pappin DJC, Creasy DM et al (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20(18):3551–3567
Rawlings ND, Waller M, Barrett AJ et al (2014) MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res 42(D1):D503–D509
Reimer JM, Enoksson M, Samollow PB et al (2008) Extended substrate specificity of opossum chymase—implications for the origin of mast cell chymases. Mol Immunol 45(7):2116–2125
Schagger H, Vonjagow G (1987) Tricine sodium dodecyl-sulfate polyacrylamide-gel electrophoresis for the separation of proteins in the range from 1-kDa to 100-Kda. Anal Biochem 166(2):368–379
Schrag JD, Li Y, Wu S et al (1991) Ser-His-Glu triad forms the catalytic site of the lipase from geotrichum-candidum. Nature 351(6329):761–764
Sternlicht MD, Werb Z (2001) How matrix metalloproteinases regulate cell behavior. Annu Rev Cell Dev Biol 17:463–516
Sussman JL, Harel M, Frolow F et al (1991) Atomic-structure of acetylcholinesterase from Torpedo-Californica—a prototypic acetylcholine-binding protein. Science 253(5022):872–879
Wieczorek R, Dorr M, Chotera A et al (2013) Formation of RNA phosphodiester bond by histidine-containing dipeptides. ChemBioChem 14(2):217–223
Acknowledgements
The authors acknowledge the National Basic Research Program of China (2013CB910700), the Chinese National Science Foundation (No. 21232005, No. 21375113, and No. 31271405) and the Fundamental Research Funds for the Central Universities (No. 20720150049). We also thank Prof. Zhongzhou Chen in China Agricultural University for his helpful manuscript modification.
Author information
Authors and Affiliations
Contributions
Yan Liu designed the experiment, performed the MS analysis of CyPA and GFP, and wrote the manuscript. Yin-bo Li performed the related data analysis and calculation. Yan Liu and Yin-bo Li contributed equally to this work. Xiang Gao performed the MS analysis of BSA and Mb. Yong-fei Yu performed one part of the mass data analysis. Zhiliang Ji designed the calculation experiments, contributed to data interpretation and provided critical. Yuan Ma and Yan-mei Li contributed to the overall concept and experiment preparation. Yu-fen Zhao designed experiments and contributed to data analysis and interpretation as well as manuscript writing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The article does not contain or refer to any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: P. R. Jungblut.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Y., Li, Yb., Gao, X. et al. Evolutionary relationships between seryl-histidine dipeptide and modern serine proteases from the analysis based on mass spectrometry and bioinformatics. Amino Acids 50, 69–77 (2018). https://doi.org/10.1007/s00726-017-2487-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-017-2487-1