Abstract
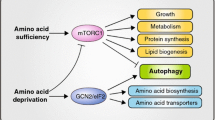
Amino acids, especially glutamine (GLN) have been known for many years to stimulate the growth of small intestinal mucosa. Polyamines are also required for optimal mucosal growth, and the inhibition of ornithine decarboxylase (ODC), the first rate-limiting enzyme in polyamine synthesis, blocks growth. Certain amino acids, primarily asparagine (ASN) and GLN stimulate ODC activity in a solution of physiological salts. More importantly, their presence is also required before growth factors and hormones such as epidermal growth factor and insulin are able to increase ODC activity. ODC activity is inhibited by antizyme-1 (AZ) whose synthesis is stimulated by polyamines, thus, providing a negative feedback regulation of the enzyme. In the absence of amino acids mammalian target of rapamycin complex 1 (mTORC1) is inhibited, whereas, mTORC2 is stimulated leading to the inhibition of global protein synthesis but increasing the synthesis of AZ via a cap-independent mechanism. These data, therefore, explain why ASN or GLN is essential for the activation of ODC. Interestingly, in a number of papers, AZ has been shown to inhibit cell proliferation, stimulate apoptosis, or increase autophagy. Each of these activities results in decreased cellular growth. AZ binds to and accelerates the degradation of ODC and other proteins shown to regulate proliferation and cell death, such as Aurora-A, Cyclin D1, and Smad1. The correlation between the stimulation of ODC activity and the absence of AZ as influenced by amino acids is high. Not only do amino acids such as ASN and GLN stimulate ODC while inhibiting AZ synthesis, but also amino acids such as lysine, valine, and ornithine, which inhibit ODC activity, increase the synthesis of AZ. The question remaining to be answered is whether AZ inhibits growth directly or whether it acts by decreasing the availability of polyamines to the dividing cells. In either case, evidence strongly suggests that the regulation of AZ synthesis is the mechanism through which amino acids influence the growth of intestinal mucosa. This brief article reviews the experiments leading to the information presented above. We also present evidence from the literature that AZ acts directly to inhibit cell proliferation and increase the rate of apoptosis. Finally, we discuss future experiments that will determine the role of AZ in the regulation of intestinal mucosal growth by amino acids.








Similar content being viewed by others
References
Alpers DH (1972) Protein synthesis in intestinal mucosa: the effect of route of administration of precursor amino acids. J Clin Invest 51:167–173
Burrin DG, Shulman RJ, Langston C, Storm MC (1994) Supplemental alanyl glutamine, organ growth and nitrogen metabolism in neonatal pigs fed by total parenteral nutrition. J Parenter Enteral Nutr 18:313–319
Chen KY, Canellakis ES (1977) Enzyme regulation in neuroblastoma cells in a salts-glucose medium: induction of ornithine decarboxylase by asparagine and glutamine. Proc Natl Acad Sci USA 74:3791–3795
Clarke RM (1977) “Luminal nutrition” versus “functional work-load” as controllers of mucosal morphology and epithelial replacement in the rat small intestine. Digestion 15:411–412
Coleman CS, Stanley BA, Viswanath R, Pegg AE (1994) Rapid exchange of subunits of mammalian ornithine decarboxylase. J Biol Chem 269:3155–3158
Deng W, Viar MJ, Johnson LR (2005) Polyamine depletion inhibits irradiation-induced apoptosis in intestinal epithelia. Am J Physiol 289:G599–G606
Dulloo I, Gopalan G, Melino G, Sabapathy K (2010) The antiapoptotic delta Np73 is degraded in a c-Jun-dependent manner upon genotoxic stress through the antizyme-mediated pathway. Proc Natl Acad Sci USA 107:4902–4917
Fausto N (1971) The control of ornithine decarboxylase activity during liver regeneration. Biochim Biophys Acta 238:116–128
Feith DJ, Shantz LM, Pegg AE (2001) Targeted antizyme expression in the skin of transgenic mice reduces tumor promoter induction of ornithine decarboxylase and decreases sensitivity to chemical carcinogenesis. Cancer Res 61:6073–6081
Fong LYY, Feith DJ, Pegg AE (2003) Antizyme over expression in transgenic mice reduces cell proliferation, increases apoptosis, and reduces N-nitrosomethylbenzylamide-induced forestomach carcinogenesis. Cancer Res 63:3945–3954
Hara K, Maruki Y, Long X, Yoshino K, Oshiro N, Hidayat S, Tokunaga C, Avruch J, Yonezawa K (2002) Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110:177–189
Heller JS, Fong WF, Canellakis ES (1976) Induction of a protein inhibitor to ornithine decarboxylase by the end products of its reaction. Proc Natl Acad Sci USA 73:1858–1862
Higashiguchi T, Hasselgren P-O, Wagner K, Fischer JE (1993) Effect of glutamine on protein synthesis in isolated intestinal epithelial cells. J Parenter Enteral Nutr 17:307–314
Hirschfield JS, Kern F (1969) Protein starvation and the small intestine. III. Incorporation of orally and intraperitoneally administered l-leucine 4,55-3H into intestinal protein of protein-deprived rats. J Clin Invest 48:1224–1229
Hogan BLM, Murden S (1974) Effect of growth conditions on the activity of ornithine decarboxylase in cultured hepatoma cells: I. Effect of amino acid supply. J Cell Physiol 83:345–352
Jacinto E, Loewith R, Schmidt A, Liu S, Ruegg MA, Hall A, Hall MN (2004) Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat Cell Biol 6:1122–1128
Jacobs LR, Taylor BR, Dowling RH (1975) Effect of luminal nutrition on intestinal adaptation following Thiry-Vella bypass in the dog. Clin Sci Mol Med 49:26
Johnson LR, McCormack SA (1994) Regulation of gastrointestinal mucosal growth. In: Johnson LR (ed) Physiology of the gastrointestinal tract, 3rd edn. Raven, New York, pp 611–642
Kasbek C, Yang CH, Fisk HA (2010) Antizyme restrains centrosome amplification by regulating the accumulation of Mps1 at centrosomes. Mol Biol Cell 21:3878–3889
Kay JE, Lindsay VJ, Cooke A (1972) Ornithine decarboxylase in phytohaemagglutinin stimulated lymphocytes: control of degradation by amino acids. FEBS Lett 21:123–126
Li X, Coffino P (1992) Regulated degradation of ornithine decarboxylase requires interaction with the polyamine-inducible protein antizyme. Mol Cell Biol 12:3556–3562
Lim SK, Gopalan G (2007) Antizyme mediates AURKA1P1-dependent degradation of Aurora-A. Oncogene 26:6593–6603
Lin Y, Martin J, Gruendler C, Farley J, Meng X, Li BY, Lechleider R, Huff C, Kim RH, Grasser WA, Paralker V, Wang T (2002) A novel link between the proteasome pathway and the signal transduction pathway of the bone morphogenetic proteins (BMPs). BMC Cell Biol 3:15
Lin KW, Nam SY, Toh WH, Dulloo I, Sabapathy K (2004) Multiple stress signals induce p73 beta accumulation. Neoplasia 6:546–557
Mitchell JL, Chen HJ (1990) Conformational changes in ornithine decarboxylase enable recognition by antizyme. Biochim Biophys Acta 1037:115–121
Newman RM, Mosbacher A, Mangold U, Koike C, Diah S, Schmidt M, Finley D, Zetter BR (2004) Antizyme targets cyclin DI for degradation. A novel mechanism for cell growth repression. J Biol Chem 279:41504–41511
Packham G, Cleveland JL (1994) Ornithine decarboxylase is a mediator of c-Myc induced apoptosis. Mol Cell Biol 14:5741–5747
Pegg AE (2006) Regulation of ornithine decarboxylase. J Biol Chem 281:14529–14532
Pegg AE, McCann PP (1982) Polyamine metabolism and function. Am J Physiol 243:C212–C221
Pegg AE, McGovern KA, Weist L (1987) Decarboxylation of alpha-difluoromethylamine by ornithine decarboxylase. Biochem J 241:305–307
Pegg AE, Feith DJ, Fong LY, Coleman CS, O’Brien TG, Shantz LM (2003) Transgenic mouse models for studies of the role of polyamines in normal, hypertrophic and neoplastic growth. Biochem Soc Trans 31:356–360
Pouline R, Pelletier G, Pegg AE (1995) Induction of apoptosis by excessive polyamine accumulation in ornithine decarboxylase-overproducing L1210 cells. Biochem J 311:723–727
Quaroni A, Wands J, Trelstad RL, Isselbacher KJ (1979) Epithelioid cell culture from rat small intestine. Characterization by morphologic and immunologic criteria. J Cell Biol 80:248–265
Ray RM, Viar MJ, Patel TB, Johnson LR (1999) Interaction of asparagine and EGF in the regulation of ornithine decarboxylase in IEC-6 cells. Am J Physiol 276:G773–G780
Ray RM, Viar MJ, Yuan Q, Johnson LR (2000) Polyamine depletion delays apoptosis of rat intestinal epithelial cells. Am J Physiol 278:C400–C489
Ray RM, Viar MJ, Johnson LR (2012) Amino acids regulate the expression of antizyme-1 to modulate ornithine decarboxylase activity. J Biol Chem 287:3674–3690
Rinehart CA Jr, Canellakis ES (1985) Induction of ornithine decarboxylase activity by insulin and growth factors is mediated by amino acids. Proc Natl Acad Sci USA 82:4365–4368
Rinehart CA Jr, Viceps-Madore D, Fong WF, Ortiz JG, Canellakis ES (1985) The effect of transport system A and N amino acids and nerve and epidermal growth factors on the induction of ornithine decarboxylase activity. J Cell Physiol 123:435–441
Russell DH (1985) Ornithine decarboxylase: a key regulatory enzyme in normal and neoplastic growth. Drug Met Rev 16:1–88
Ryan KK, Seeley RJ (2013) Food as a hormone. Science 339:918–919
Sarbassov DD, Ali SM, Kim DH, Guertin DA, Latek RR, Erdjument-Bromage H, Tempst P, Sabitini DM (2004) Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr Biol 14:1296–1302
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM (2005) Phosphorylation and regulation of Akt/PKB by Rictor mTOR complex. Science 307:1098–1101
Suzuki J, Murakami Y, Samejima K, Ontani KK, Oka T (2009) Antizyme is necessary for conversion of pancreatic tumor cells into glucagon-producing differentiated cells. Endocr Relat Cancer 16:649–659
Tato I, Bartons R, Ventura F, Rosa JL (2011) Amino acids activate mammalian target of rapamycin complex 2 (mTORC2) via P13K/Akt signaling. J Biol Chem 272:26457–26463
Tobias KE, Kahana C (1995) Exposure to ornithine results in excessive accumulation of putrescine and apoptotic cell death in ornithine decarboxylase overproducing mouse myeloma cells. Cell Growth Differ 10:1279–1285
Viceps-Madore D, Chen KY, Tsou HR, Canellakis ES (1982) Studies on the role of protein synthesis and of sodium on the regulation of ornithine decarboxylase activity. Biochim Biophys Acta 717:305–315
Wang J-Y, Johnson LR (1990) Role of ornithine decarboxylase in the repair process of gastric mucosal stress ulcers. Am J Physiol 258:G78–G85
Wang J-Y, Johnson LR (1991) Polyamines and ornithine decarboxylase during repair of duodenal mucosa after stress in rats. Gastroenterology 100:333–343
Wang X, Proud CG (2006) The mTOR pathway in the control of protein synthesis. Physiology 21:362–369
Woo RA, Poon RY (2003) Cyclin-dependent kinases and S phase control in mammalian cells. Cell Cycle 2:316–324
Yoshida S, Leskiw MJ, Schulter MD et al (1992) Effect of total parenteral nutrition, systemic sepsis, and glutamine on gut mucosa in rats. Am J Physiol 263:E368–E373
Zimmerman BJ, McCormack SA, Israel M, Johnson LR (1995) Polyamine-mediated cell migration and growth: structural requirement for diamine analogues to substitute for putrescine. Cell Pharmacol 2:109–113
Zoncu R, Efeyan A, Sabitini DM (2011) From growth signal integration to cancer, diabetes and aging. Nature Rev Mol Cell Biol 12:21–35
Acknowledgments
This publication was made possible by Grants (Number DK-16505 and DK-052784) from the National Institute of Diabetes and Digestive and Kidney Disease (NIDDK) and by support from the Thomas A. Gerwin endowment. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the National Institute of Health.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ray, R.M., Johnson, L.R. Regulation of intestinal mucosal growth by amino acids. Amino Acids 46, 565–573 (2014). https://doi.org/10.1007/s00726-013-1565-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-013-1565-2