Abstract
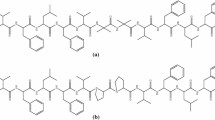
Conformations of three pairs of dehydropeptides with the opposite configuration of the ΔPhe residue, Boc-Gly-ΔZ/EPhe-Phe-p-NA (Z- p -NA and E- p -NA), Boc-Gly-ΔZ/EPhe-Phe-OMe (Z-OMe and E-OMe), and Boc-Gly-ΔZ/EPhe-Phe-OH (Z-OH and E-OH) were compared on the basis of CD and NMR studies in MeOH, TFE, and DMSO. The CD results were used as the additional input data for the NMR-based calculations of the detailed solution conformations of the peptides. It was found that Z- p -NA, E- p -NA, Z-OMe, and Z-OH adopt the β-turn conformations and E-OMe and E-OH are unordered. There are two overlapping type III β-turns in Z- p -NA, type II’ β-turn in E- p -NA, and type II β-turn in Z-OMe and Z-OH. The results obtained indicate that in the case of methyl esters and peptides with a free carboxyl group, ΔZPhe is a much stronger inducer of ordered conformations than ΔEPhe. It was also found that temperature coefficients of the amide protons are not reliable indicators of intramolecular hydrogen bonds donors in small peptides.






Similar content being viewed by others
References
Baldwin JE, Claridge TDW, Hulme C, Rodger A, Schofield CJ (1994) Comments on the use of a dichromophoric circular dichroism assay for the identification of β-turns in peptides. Int J Peptide Protein Res 43:180–183
Cammers-Goodwin A, Allen TJ, Oslick SL, McClure KF, Lee JH, Kemp DS (1996) Mechanism of stabilization of helical conformations of polypeptides by water containing trifluoroethanol. J Am Chem Soc 118:3082–3090
Cierpicki T, Otlewski J (2001) Amide proton temperature coefficients as hydrogen bond indicators in proteins. J Biomol NMR 21:249–261
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multi-dimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Goddard TD, Kneller DG (2003) SPARKY 3. University of California, San Francisco
Gupta M, Chauhan V (2011) De novo design of α, β-didehydrophenylalanine containing peptides: from models to applications. Biopolymers 95:161–173 and references therein
Gupta M, Acharya R, Mishra A, Ramakumar S, Ahmed F, Chauhan VS (2008) Dehydrophenylalanine (ΔPhe) as a β breaker: extended structure terminated by a ΔPhe-induced turn in the pentapeptide Boc-Phe1-Ala2-Ile3-ΔPhe4-Ala5-OMe. Chem Bio Chem 9:1375–1378
Halab L, Gosselin F, Lubell WD (2000) Design, synthesis, and conformational analysis of azacycloalkane amino acids as conformationally constrained probes for mimicry of peptide secondary structures. Biopolymers (Pept Sci) 55:101–122
Jaremko Ł, Jaremko M, Pasikowski P, Cebrat M, Stefanowicz P, Lisowski M, Artym J, Zimecki M, Zhukov I, Szewczuk Z (2009) The immunosuppressive activity and solution structures of ubiquitin fragments. Biopolymers 91:423–431
Jewginski M, Latajka R, Krężel A, Haremza K, Makowski M, Kafarski P (2013) Influence of solvents on conformation of dehydropeptides. J Mol Struct 1035:129–139
Koradi R, Billeter M, Wüthrich K (1996) MolMol: a program for display and analysis of macromolecular structures. J Mol Graphics 14:51–55
Latajka R, Makowski M, Jewgiński M, Pawełczak M, Koroniak H, Kafarski P (2006) Peptide p-nitrophenylanilides containing (E)-dehydrophenylalanine–synthesis, structural studies and evaluation of their activity towards cathepsin C. New J Chem 30:1009–1018
Latajka R, Jewgiński M, Makowski M, Krężel A (2008a) Conformational studies of hexapeptides containing two dehydroamino acid residues in positions 3 and 5 in peptide chain. J Mol Struct 92:446–451
Latajka R, Jewgiński M, Makowski M, Krężel A, Paluch S (2008b) Conformational studies of hexapeptides containing two dehydroamino acid residues in positions 2 and 5 in peptide chain. Biopolymers 89:691–699
Lewis PN, Momany FA, Scheraga HA (1973) Chain reversals in proteins. Biochim Biophys Acta 303:211–229
Lisowski M, Latajka R, Picur B, Lis T, Bryndal I, Rospenk M, Makowski M, Kafarski P (2008) Combined effect of the ΔPhe or ΔAla residue and the p-nitroanilide group on a didehydropeptides conformation. Biopolymers 89:220–234
Lisowski M, Jaremko Ł, Jaremko M, Mazur A, Latajka R, Makowski M (2010) Effect of the ΔPhe residue configuration on a didehydropeptides conformation: a combined CD and NMR study. Biopolymers 93:1055–1064
Luo P, Baldwin RL (1997) Mechanism of helix induction by trifluoroethanol: a framework for extrapolating the helix-forming properties of peptides from trifluoroethanol/water mixtures back to water. Biochemistry 36:8413–8421
Makowski M, Rzeszotarska B, Kubica Z, Pietrzyński G, Hetper J (1986) Coupling experiments with C-terminal dehydrophenylalanine and dehydroalanine residues. Liebigs Ann Chem 1986:980–991
Makowski M, Pawełczak M, Latajka R, Nowak K, Kafarski P (2001) Synthesis of tetrapeptide p-nitrophenylanilides containing dehydroalanine and dehydrophenylalanine and their influence on cathepsin C activity. J Peptide Sci 7:141–145
Makowski M, Brzuszkiewicz A, Lisowski M, Lis T (2005) N-[tert-Butoxycarbonylglycyl-(Z)-α,β-dehydrophenylalanylglycyl-(E)-α,β-dehydrophenylalanylphenylalanyl]-4-nitroaniline ethanol solvate. Acta Cystallogr Sect C 61:o424–o426
Makowski M, Lisowski M, Mikołajczyk I, Lis T (2007) N-[tert-Butoxycarbonylglycyl-(E)-α,β-dehydrophenylalanylglycylglycyl-(E)-α,β-dehydrophenylalanyl]glycine. Acta Crystallogr Sect E 63:o19–o21
Makowski M, Lisowski M, Maciąg A, Wiktor M, Szlachcic A, Lis T (2010) Two pentadehydropeptides with different configurations of the ΔPhe residues. Acta Crystallogr Sect C 66:o119–o123
Mathur P, Ramakumar S, Chauhan VS (2004) Peptide design using α, β-dehydro amino acids: from β-turns to helical hairpins. Biopolymers 76:150–161 and references therein
Pieroni O, Fissi A, Pratesi C, Temussi PA, Ciardelli F (1993) Solution structure of peptides containing two dehydrophenylalanine residues: a CD investigation. Biopolymers 33:1–10
Pieroni O, Fissi A, Jain RM, Chauhan VS (1996) Solution structure of dehydropeptides: a CD investigation. Biopolymers 38:97–108
Rajan R, Balaram P (1996) A model for the interaction of trifluoroethanol with peptides and proteins. Int J Peptide Protein Res 48:328–336
Ramagopal UA, Ramakumar S, Sahal D, Chauhan VS (2001) De novo design and characterization of an apolar helical hairpin peptide at atomic resolution: compaction mediated by weak interactions. Proc Natl Acad Sci USA 98:870–874
Ramagopal UA, Ramakumar S, Mathur P, Joshi RM, Chauhan VS (2002) Dehydrophenylalanine zippers: strong helix–helix clamping through a network of weak interactions. Protein Eng 15:331–335
Sato K, Kawai M, Nagai U (1981) β-Turn preference of tetrapeptide sequences as analyzed by CD spectra of their dnp-pNA derivatives. Biopolymers 20:1921–1927
Schwieters CD, Kuszewski JJ, Tjandra N, Clore GM (2003) The Xplor-NIH NMR molecular structure determination package. J Magn Reson 160:65–73
Stammer CH (1982) Dehydroamino acids and peptides. Chem Biochem Amino Acids Peptides Proteins 6:33–74
Wüthrich K (1986) NMR of proteins and nucleic acids. John Wiley and Sons, New York
Acknowledgments
We would like to thank prof. Andrzej Ejchart (Institute of Biochemistry and Biophysics, Polish Academy of Sciences) for making the 500 MHz Varian spectrometer available to us. The syntheses of peptides, performed by M. Makowski, were supported by the Wrocław Research Center EIT + under the project “Biotechnologies and advanced medical technologies––BioMed” (POIG 01.01.02-02-003/08-00 financed from the European Regional Development Fund (Operational Programme Innovative Economy, 1.1.2).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Jaremko, M., Jaremko, Ł., Mazur, A. et al. Enhanced β-turn conformational stability of tripeptides containing ΔPhe in cis over trans configuration. Amino Acids 45, 865–875 (2013). https://doi.org/10.1007/s00726-013-1534-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-013-1534-9