Abstract
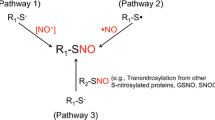
Protein S-nitrosylation is the covalent redox-related modification of cysteine sulfhydryl groups with nitric oxide, creating a regulatory impact similar to phosphorylation. Recent studies have reported a growing number of proteins to be S-nitrosylated in vivo resulting in altered functions. These studies support S-nitrosylation as a critical regulatory mechanism, fine-tuning protein activities within diverse cellular processes and biochemical pathways. In addition, S-nitrosylation appears to have key roles in the etiology of a broad range of human diseases. In this review, we discuss recent advances in proteomic approaches for the enrichment, identification, and quantitation of cysteine S-nitrosylated proteins and peptides. These advances have provided analytical tools with the power to interpret the impact of S-nitrosylation at the system level, providing a new platform for drug discovery and the identification of diagnostic markers for human diseases.


Similar content being viewed by others
References
Bogdan C (2001) Nitric oxide and the immune response. Nat Immunol 2(10):907–916
Camerini S, Polci ML, Restuccia U, Usuelli V, Malgaroli A, Bachi A (2007) A novel approach to identify proteins modified by nitric oxide: the HIS-TAG switch method. J Proteome Res 6(8):3224–3231
Chen YY, Huang YF, Khoo KH, Meng TC (2007) Mass spectrometry-based analyses for identifying and characterizing S-nitrosylation of protein tyrosine phosphatases. Methods 42(3):243–249
Chen YJ, Ku WC, Lin PY, Chou HC, Khoo KH, Chen YJ (2010) S-alkylating labeling strategy for site-specific identification of the s-nitrosoproteome. J Proteome Res 9(12):6417–6439
Chung KK (2006) Say NO to neurodegeneration: role of S-nitrosylation in neurodegenerative disorders. Neurosignals 15(6):307–313. doi:000109071.10.1159/000109071
Cleland WW (1964) Dithiothreitol, a new protective reagent for Sh groups. Biochemistry 3:480–482
Derakhshan B, Wille PC, Gross SS (2007) Unbiased identification of cysteine S-nitrosylation sites on proteins. Nat Protoc 2(7):1685–1691. doi:10.10.1038/nprot.2007.210
Doulias PT, Greene JL, Greco TM, Tenopoulou M, Seeholzer SH, Dunbrack RL, Ischiropoulos H (2010) Structural profiling of endogenous S-nitrosocysteine residues reveals unique features that accommodate diverse mechanisms for protein S-nitrosylation. Proc Natl Acad Sci USA 107(39):16958–16963
Faccenda A, Bonham CA, Vacratsis PO, Zhang X, Mutus B (2010) Gold nanoparticle enrichment method for identifying S-nitrosylation and S-glutathionylation sites in proteins. J Am Chem Soc 132(33):11392–11394. doi:10.1021/ja103591v
Forrester MT, Foster MW, Benhar M, Stamler JS (2009a) Detection of protein S-nitrosylation with the biotin-switch technique. Free Radic Biol Med 46(2):119–126
Forrester MT, Thompson JW, Foster MW, Nogueira L, Moseley MA, Stamler JS (2009b) Proteomic analysis of S-nitrosylation and denitrosylation by resin-assisted capture. Nat Biotechnol 27(6):557–559
Foster MW, McMahon TJ, Stamler JS (2003) S-nitrosylation in health and disease. Trends Mol Med 9(4):160–168
Foster MW, Forrester MT, Stamler JS (2009a) A protein microarray-based analysis of S-nitrosylation. Proc Natl Acad Sci USA 106(45):18948–18953. doi:10.1073/pnas.0900729106
Foster MW, Hess DT, Stamler JS (2009b) Protein S-nitrosylation in health and disease: a current perspective. Trends Mol Med 15(9):391–404. doi:10.1016/j.molmed.2009.06.007
Fukumura D, Kashiwagi S, Jain RK (2006) The role of nitric oxide in tumour progression. Nat Rev Cancer 6(7):521–534
Giustarini D, Dalle-Donne I, Colombo R, Milzani A, Rossi R (2008) Is ascorbate able to reduce disulfide bridges? A cautionary note. Nitric Oxide 19(3):252–258
Greco TM, Hodara R, Parastatidis I, Heijnen HF, Dennehy MK, Liebler DC, Ischiropoulos H (2006) Identification of S-nitrosylation motifs by site-specific mapping of the S-nitrosocysteine proteome in human vascular smooth muscle cells. Proc Natl Acad Sci USA 103(19):7420–7425
Han P, Chen C (2008) Detergent-free biotin switch combined with liquid chromatography/tandem mass spectrometry in the analysis of S-nitrosylated proteins. Rapid Commun Mass Spectrom 22(8):1137–1145. doi:10.1002/rcm.3476
Hao G, Gross SS (2006) Electrospray tandem mass spectrometry analysis of S- and N-nitrosopeptides: facile loss of NO and radical-induced fragmentation. J Am Soc Mass Spectrom 17(12):1725–1730
Hao G, Xie L, Gross SS (2004) Argininosuccinate synthetase is reversibly inactivated by S-nitrosylation in vitro and in vivo. J Biol Chem 279(35):36192–36200
Hao G, Derakhshan B, Shi L, Campagne F, Gross SS (2006) SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc Natl Acad Sci USA 103(4):1012–1017
Hess DT, Matsumoto A, Kim SO, Marshall HE, Stamler JS (2005) Protein S-nitrosylation: purview and parameters. Nat Rev Mol Cell Biol 6(2):150–166
Huang B, Chen C (2010) Detection of protein S-nitrosation using irreversible biotinylation procedures (IBP). Free Radic Biol Med 49(3):447–456. doi:S0891-5849(10)00285-6.10.1016/j.freeradbiomed.2010.05.001
Huerta S, Chilka S, Bonavida B (2008) Nitric oxide donors: novel cancer therapeutics (review). Int J Oncol 33(5):909–927
Jaffrey SR, Snyder SH (2001) The biotin switch method for the detection of S-nitrosylated proteins. Sci STKE 2001 (86):pl1
Kaneko R, Wada Y (2003) Decomposition of protein nitrosothiolsin matrix-assisted laser desorption/ionization and electrospray ionization mass spectrometry. J Mass Spectrom 38(5):526–530
Knott AB, Bossy-Wetzel E (2009) Nitric oxide in health and disease of the nervous system. Antioxid Redox Signal 11(3):541–554. doi:10.1089/ARS.2008.2234
Kohr MJ, Aponte AM, Sun J, Wang G, Murphy E, Gucek M, Steenbergen C (2011) Characterization of potential S-nitrosylation sites in the myocardium. Am J Physiol Heart Circ Physiol 300(4):H1327–H1335. doi:10.1152/ajpheart.00997.2010
Lim KH, Ancrile BB, Kashatus DF, Counter CM (2008) Tumour maintenance is mediated by eNOS. Nature 452(7187):646–649. doi:10.1038/nature06778
Liu M, Hou J, Huang L, Huang X, Heibeck TH, Zhao R, Pasa-Tolic L, Smith RD, Li Y, Fu K, Zhang Z, Hinrichs SH, Ding SJ (2010) Site-specific proteomics approach for study protein S-nitrosylation. Anal Chem 82(17):7160–7168
Mannick JB, Schonhoff CM (2008) Measurement of protein S-nitrosylation during cell signaling. Methods Enzymol 440:231–242. doi:10.1016/S0076-6879(07)00814-2
Mirza UA, Chait BT, Lander HM (1995) Monitoring reactions of nitric oxide with peptides and proteins by electrospray ionization-mass spectrometry. J Biol Chem 270(29):17185–17188
Murray CI, Kane LA, Uhrigshardt H, Wang SB, Van Eyk JE (2011a) Site-mapping of in vitro S-nitrosation in cardiac mitochondria: implications for cardioprotection. Mol Cell Proteomics 10(3):M110 004721. doi:10.1074/mcp.M110.004721
Murray CI, Uhrigshardt H, O’Meally RN, Cole RN, Van Eyk JE (2011b) Identification and quantification of S-nitrosylation by cysteine reactive tandem mass tag switch assay. Mol Cell Proteomics. doi:10.1074/mcp.M111.013441
Ravi K, Brennan LA, Levic S, Ross PA, Black SM (2004) S-nitrosylation of endothelial nitric oxide synthase is associated with monomerization and decreased enzyme activity. Proc Natl Acad Sci USA 101(8):2619–2624. doi:10.1073/pnas.0300464101
Retamal MA, Cortes CJ, Reuss L, Bennett MV, Saez JC (2006) S-nitrosylation and permeation through connexin 43 hemichannels in astrocytes: induction by oxidant stress and reversal by reducing agents. Proc Natl Acad Sci USA 103(12):4475–4480. doi:10.1073/pnas.0511118103
Shi Q, Xiong Q, Wang B, Le X, Khan NA, Xie K (2000) Influence of nitric oxide synthase II gene disruption on tumor growth and metastasis. Cancer Res 60(10):2579–2583
Sinha V, Wijewickrama GT, Chandrasena RE, Xu H, Edirisinghe PD, Schiefer IT, Thatcher GR (2010) Proteomic and mass spectroscopic quantitation of protein S-nitrosation differentiates NO-donors. ACS Chem Biol 5(7):667–680. doi:10.1021/cb100054m
Snyder SH (1992) Nitric oxide: first in a new class of neurotransmitters. Science 257(5069):494–496
Sun J, Murphy E (2010) Protein S-nitrosylation and cardioprotection. Circ Res 106(2):285–296. doi:10.1161/CIRCRESAHA.109.209452
Sun J, Steenbergen C, Murphy E (2006) S-nitrosylation: NO-related redox signaling to protect against oxidative stress. Antioxid Redox Signal 8(9–10):1693–1705. doi:10.1089/ars.2006.8.1693
Taldone FS, Tummala M, Goldstein EJ, Ryzhov V, Ravi K, Black SM (2005) Studying the S-nitrosylation of model peptides and eNOS protein by mass spectrometry. Nitric Oxide 13(3):176–187
Torta F, Usuelli V, Malgaroli A, Bachi A (2008) Proteomic analysis of protein S-nitrosylation. Proteomics 8(21):4484–4494
Tripathi P, Kashyap L, Singh V (2007) The role of nitric oxide in inflammatory reactions. FEMS Immunol Med Microbiol 51(3):443–452. doi:10.1111/j.1574-695X.2007.00329.x
Tuteja N, Chandra M, Tuteja R, Misra MK (2004) Nitric oxide as a unique bioactive signaling messenger in physiology and pathophysiology. J Biomed Biotechnol 4:227–237. doi:10.1155/S1110724304402034
Wang Y, Liu T, Wu C, Li H (2008) A strategy for direct identification of protein S-nitrosylation sites by quadrupole time-of-flight mass spectrometry. J Am Soc Mass Spectrom 19(9):1353–1360
Wiktorowicz JE, Stafford S, Rea H, Urvil P, Soman K, Kurosky A, Perez-Polo JR, Savidge TC (2011) Quantification of cysteinyl S-nitrosylation by fluorescence in unbiased proteomic studies. Biochemistry 50(25):5601–5614. doi:10.1021/bi200008b
Witze ES, Old WM, Resing KA, Ahn NG (2007) Mapping protein post-translational modifications with mass spectrometry. Nat Methods 4(10):798–806. doi:10.1038/nmeth1100
Xie K, Fidler IJ (1998) Therapy of cancer metastasis by activation of the inducible nitric oxide synthase. Cancer Metastasis Rev 17(1):55–75
Zhao R, Ding SJ, Shen Y, Camp DG 2nd, Livesay EA, Udseth H, Smith RD (2009) Automated metal-free multiple-column nanoLC for improved phosphopeptide analysis sensitivity and throughput. J Chromatogr B Analyt Technol Biomed Life Sci 877(8–9):663–670
Zhou X, Han P, Li J, Zhang X, Huang B, Ruan HQ, Chen C (2010) ESNOQ, proteomic quantification of endogenous S-nitrosation. PLoS ONE 5(4):e10015. doi:10.1371/journal.pone.0010015
Acknowledgments
We thank Dr. Lawrence M. Schopfer and Stites M. Kirsten for editing this manuscript. This work was supported by the Department of Pathology and Microbiology at the University of Nebraska Medical Center (UNMC), NEHHS LB606 to S.J.D., Nebraska Research Initiative to S.J.D. and J.E.T., M.L. is supported by a fellowship from the College of Medicine at UNMC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, M., Talmadge, J.E. & Ding, SJ. Development and application of site-specific proteomic approach for study protein S-nitrosylation. Amino Acids 42, 1541–1551 (2012). https://doi.org/10.1007/s00726-012-1279-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-012-1279-x