Summary.
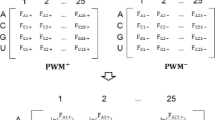
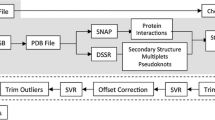
Protein–RNA interactions play a key role in a number of biological processes such as protein synthesis, mRNA processing, assembly and function of ribosomes and eukaryotic spliceosomes. A reliable identification of RNA-binding sites in RNA-binding proteins is important for functional annotation and site-directed mutagenesis. We developed a novel method for the prediction of protein residues that interact with RNA using support vector machine (SVM) and position-specific scoring matrices (PSSMs). Two cases have been considered in the prediction of protein residues at RNA-binding surfaces. One is given the sequence information of a protein chain that is known to interact with RNA; the other is given the structural information. Thus, five different inputs have been tested. Coupled with PSI-BLAST profiles and predicted secondary structure, the present approach yields a Matthews correlation coefficient (MCC) of 0.432 by a 7-fold cross-validation, which is the best among all previous reported RNA-binding sites prediction methods. When given the structural information, we have obtained the MCC value of 0.457, with PSSMs, observed secondary structure and solvent accessibility information assigned by DSSP as input. A web server implementing the prediction method is available at the following URL: http://210.42.106.80/printr/.
Similar content being viewed by others
References
S Ahmad MM Gromiha A Sarai (2003) ArticleTitleReal value prediction of solvent accessibility from amino acid sequence Proteins Struct Funct Genet 50 629–635 Occurrence Handle12577269 Occurrence Handle10.1002/prot.10328 Occurrence Handle1:CAS:528:DC%2BD3sXislOlsb4%3D
S Ahmad MM Gromiha A Sarai (2004) ArticleTitleAnalysis and prediction of DNA-binding proteins and their binding residues based on sequence and structural information Bioinformatics 20 477–486 Occurrence Handle14990443 Occurrence Handle10.1093/bioinformatics/btg432 Occurrence Handle1:CAS:528:DC%2BD2cXhvFSru7k%3D
S Ahmad A Sarai (2005) ArticleTitlePSSM-based prediction of DNA binding sites in proteins BMC Bioinformatics 6 33 Occurrence Handle15720719 Occurrence Handle10.1186/1471-2105-6-33 Occurrence Handle1:CAS:528:DC%2BD2MXivVajt74%3D
J Allers Y Shamoo (2001) ArticleTitleStructure-based analysis of protein-RNA interactions using the program ENTANGLE J Mol Biol 311 75–86 Occurrence Handle11469858 Occurrence Handle10.1006/jmbi.2001.4857 Occurrence Handle1:CAS:528:DC%2BD3MXlsVSjsrg%3D
SF Altschul TL Madden AA Schaffer J Zhang Z Zhang W Miller DJ Lipman (1997) ArticleTitleGapped blast and psi-blast: a new generation of protein databases and search programs Nucleic Acids Res 25 3389–3402 Occurrence Handle9254694 Occurrence Handle10.1093/nar/25.17.3389 Occurrence Handle1:CAS:528:DyaK2sXlvFyhu7w%3D
HM Berman J Westbrook Z Feng G Gilliland TN Bhat H Weissig IN Shindyalov PE Bourne (2000) ArticleTitleThe protein data bank Nucleic Acids Res 28 235–242 Occurrence Handle10592235 Occurrence Handle10.1093/nar/28.1.235 Occurrence Handle1:CAS:528:DC%2BD3cXhvVKjt7w%3D
YD Cai GP Zhou KC Chou (2003) ArticleTitleSupport vector machines for predicting membrane protein types by using functional domain composition Biophys J 84 3257–3263 Occurrence Handle12719255 Occurrence Handle1:CAS:528:DC%2BD3sXjvFGju7o%3D
J Chen H Liu J Yang KC Chou (2007) ArticleTitlePrediction of linear B-cell epitopes using amino acid pair antigenicity scale Amino Acids 33 423–428 Occurrence Handle17252308 Occurrence Handle10.1007/s00726-006-0485-9 Occurrence Handle1:CAS:528:DC%2BD2sXpvVagsrc%3D
KC Chou (2001) ArticleTitlePrediction of protein cellular attributes using pseudo amino acid composition Proteins Struct Funct Genet 43 246–255 Occurrence Handle11288174 Occurrence Handle10.1002/prot.1035 Occurrence Handle1:CAS:528:DC%2BD3MXjtFOls74%3D
KC Chou (2005a) ArticleTitleCoupling interaction between thromboxane A2 receptor and alpha-13 subunit of guanine nucleotide-binding protein J Proteome Res 4 1681–1686 Occurrence Handle10.1021/pr050145a Occurrence Handle1:CAS:528:DC%2BD2MXmsVyitro%3D
KC Chou (2005b) ArticleTitleInsights from modeling the 3D structure of DNA-CBF3b complex J Proteome Res 4 1657–1660 Occurrence Handle10.1021/pr050135+ Occurrence Handle1:CAS:528:DC%2BD2MXmvVCrsLg%3D
KC Chou (2005c) ArticleTitleUsing amphiphilic pseudo amino acid composition to predict enzyme subfamily classes Bioinformatics 21 10–19 Occurrence Handle10.1093/bioinformatics/bth466 Occurrence Handle1:CAS:528:DC%2BD2MXisVWitw%3D%3D
KC Chou YD Cai (2002) ArticleTitleUsing functional domain composition and support vector machines for prediction of protein subcellular location J Biol Chem 277 45765–45769 Occurrence Handle12186861 Occurrence Handle10.1074/jbc.M204161200 Occurrence Handle1:CAS:528:DC%2BD38XovFKjurg%3D
KC Chou HB Shen (2006a) ArticleTitleHum-PLoc: a novel ensemble classifier for predicting human protein subcellular localization Biochem Biophys Res Commun 347 150–157 Occurrence Handle10.1016/j.bbrc.2006.06.059 Occurrence Handle1:CAS:528:DC%2BD28Xmslyrsbc%3D
KC Chou HB Shen (2006b) ArticleTitleLarge-scale predictions of Gram-negative bacterial protein subcellular locations J Proteome Res 5 3420–3428 Occurrence Handle10.1021/pr060404b Occurrence Handle1:CAS:528:DC%2BD28XhtFehurjJ
KC Chou HB Shen (2007a) ArticleTitleEuk-mPLoc: a fusion classifier for large-scale eukaryotic protein subcellular location prediction by incorporating multiple sites J Proteome Res 6 1728–1734 Occurrence Handle1:CAS:528:DC%2BD2sXjs1SrsbY%3D
KC Chou HB Shen (2007b) ArticleTitleLarge-scale plant protein subcellular location prediction J Cell Biochem 100 665–678 Occurrence Handle10.1002/jcb.21096 Occurrence Handle1:CAS:528:DC%2BD2sXht1Slu7c%3D
KC Chou HB Shen (2007c) ArticleTitleMemType-2L: a web server for predicting membrane proteins and their types by incorporating evolution information through Pse-PSSM Biochem Biophys Res Commun 360 339–345 Occurrence Handle10.1016/j.bbrc.2007.06.027 Occurrence Handle1:CAS:528:DC%2BD2sXnslSqtLw%3D
KC Chou HB Shen (2007d) ArticleTitleReview: recent progresses in protein subcellular location prediction Anal Biochem 370 1–16 Occurrence Handle10.1016/j.ab.2007.07.006 Occurrence Handle1:CAS:528:DC%2BD2sXhtVOmur%2FF
KC Chou HB Shen (2007e) ArticleTitleSignal-CF: a subsite-coupled and window-fusing approach for predicting signal peptides Biochem Biophys Res Commun 357 633–640 Occurrence Handle10.1016/j.bbrc.2007.03.162 Occurrence Handle1:CAS:528:DC%2BD2sXkslCju78%3D
KC Chou CT Zhang (1995) ArticleTitlePrediction of protein structural classes Crit Rev Biochem Mol Biol 30 275–349 Occurrence Handle7587280 Occurrence Handle10.3109/10409239509083488 Occurrence Handle1:CAS:528:DyaK2MXosFentb8%3D
Y Diao M Li Z Feng J Yin Y Pan (2007a) ArticleTitleThe community structure of human cellular signaling network J Theor Biol 247 608–615 Occurrence Handle10.1016/j.jtbi.2007.04.007
Diao Y, Ma D, Wen Z, Yin J, Xiang J, Li M (2007b) Using pseudo amino acid composition to predict transmembrane regions in protein: cellular automata and Lempel-Ziv complexity. Amino Acids, DOI: 10.1007/s00726-007-0550-z
YS Ding TL Zhang KC Chou (2007) ArticleTitlePrediction of protein structure classes with pseudo amino acid composition and fuzzy support vector machine network Protein Peptide Lett 14 811–815 Occurrence Handle10.2174/092986607781483778 Occurrence Handle1:CAS:528:DC%2BD2sXhtlWiur7J
DE Draper (1994) ArticleTitleProtein-RNA recognition Annu Rev Biochem 64 593–620 Occurrence Handle10.1146/annurev.bi.64.070195.003113
DE Draper (1999) ArticleTitleThemes in RNA-protein recognition J Mol Biol 293 255–270 Occurrence Handle10550207 Occurrence Handle10.1006/jmbi.1999.2991 Occurrence Handle1:CAS:528:DyaK1MXms12rt7k%3D
Y Gao SH Shao X Xiao YS Ding YS Huang ZD Huang KC Chou (2005) ArticleTitleUsing pseudo amino acid composition to predict protein subcellular location: approached with Lyapunov index, Bessel function, and Chebyshev filter Amino Acids 28 373–376 Occurrence Handle15889221 Occurrence Handle10.1007/s00726-005-0206-9 Occurrence Handle1:CAS:528:DC%2BD2MXlt1Kmurw%3D
YZ Guo M Li M Lu Z Wen K Wang G Li J Wu (2006) ArticleTitleClassifying G protein-coupled receptors and nuclear receptors based on protein power spectrum from fast Fourier transform Amino Acids 30 397–402 Occurrence Handle16773242 Occurrence Handle10.1007/s00726-006-0332-z Occurrence Handle1:CAS:528:DC%2BD28Xls1egs7o%3D
SJ Hua Z Sun (2001) ArticleTitleA novel method of protein secondary structure prediction with high segment overlap measure: support vector machine approach J Mol Biol 308 397–407 Occurrence Handle11327775 Occurrence Handle10.1006/jmbi.2001.4580 Occurrence Handle1:CAS:528:DC%2BD3MXjtVShs7k%3D
SJ Hua Z Sun (2001) ArticleTitleSupport vector machine approach for protein subcellular location prediction Bioinformatics 17 721–728 Occurrence Handle11524373 Occurrence Handle10.1093/bioinformatics/17.8.721 Occurrence Handle1:CAS:528:DC%2BD3MXntFKjsb0%3D
S Jahandideh P Abdolmaleki M Jahandideh EB Asadabadi (2007) ArticleTitleNovel two-stage hybrid neural discriminant model for predicting proteins structural classes Biophys Chem 128 87–93 Occurrence Handle17467878 Occurrence Handle10.1016/j.bpc.2007.03.006 Occurrence Handle1:CAS:528:DC%2BD2sXkvFGku7g%3D
E Jeong IF Chung S Miyano (2004) ArticleTitleA neural network method for identification of RNA-interacting residues in proteins Genome Inform Ser Workshop Genome Inform 15 105–116 Occurrence Handle1:CAS:528:DC%2BD2MXps1Cmurk%3D
Jeong E, Miyano S (2006) A weighted profile based method for Protein-RNA interacting residues prediction. Trans Comput Syst Biol IV: 123–139
T Joachims (1999) Making large-scale SVM learning practical B Schőlkopf C Burges A Smola (Eds) Advances in kernel methods-support vector learning MIT Press Cambridge, MA, USA
S Jones JM Thornton (1997) ArticleTitlePrediction of protein-protein interaction sites using patch analysis J Mol Biol 272 133–143 Occurrence Handle9299343 Occurrence Handle10.1006/jmbi.1997.1233 Occurrence Handle1:CAS:528:DyaK2sXmt1Wgtrc%3D
DT Jones (1999) ArticleTitleProtein secondary structure prediction based on position-specific scoring matrices J Mol Biol 292 195–202 Occurrence Handle10493868 Occurrence Handle10.1006/jmbi.1999.3091 Occurrence Handle1:CAS:528:DyaK1MXlvFyksb0%3D
S Jones DT Daley NM Luscombe HM Berman JM Thornton (2001) ArticleTitleProtein-RNA interaction: a structural analysis Nucleic Acids Res 29 943–954 Occurrence Handle11160927 Occurrence Handle10.1093/nar/29.4.943 Occurrence Handle1:CAS:528:DC%2BD3MXhs1GjsLg%3D
W Kabsch C Sander (1983) ArticleTitleDictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features Biopolymers 22 2577–2637 Occurrence Handle6667333 Occurrence Handle10.1002/bip.360221211 Occurrence Handle1:CAS:528:DyaL2cXkslegtQ%3D%3D
KD Kedarisetti L Kurgan S Dick (2006) ArticleTitleClassifier ensembles for protein structural class prediction with varying homology Biochem Biophys Res Commun 348 981–988 Occurrence Handle16904630 Occurrence Handle10.1016/j.bbrc.2006.07.141 Occurrence Handle1:CAS:528:DC%2BD28XosVOitL4%3D
A Koike T Takagi (2004) ArticleTitlePrediction of protein–protein interaction sites using support vector machines Protein Eng 17 165–173 Occurrence Handle10.1093/protein/gzh020 Occurrence Handle1:CAS:528:DC%2BD2cXisF2lur4%3D
DQ Liu H Liu HB Shen J Yang KC Chou (2007) ArticleTitlePredicting secretory protein signal sequence cleavage sites by fusing the marks of global alignments Amino Acids 32 493–496 Occurrence Handle17103116 Occurrence Handle10.1007/s00726-006-0466-z Occurrence Handle1:CAS:528:DC%2BD2sXlsVGnsL8%3D
S Mondal R Bhavna R Mohan Babu S Ramakumar (2006) ArticleTitlePseudo amino acid composition and multi-class support vector machines approach for conotoxin superfamily classification J Theor Biol 243 252–260 Occurrence Handle16890961 Occurrence Handle10.1016/j.jtbi.2006.06.014 Occurrence Handle1:CAS:528:DC%2BD28XhtVygtbzM
Morik K, Brockhausen P, Joachims T (1999) Combining statistical learning with a knowledge-based approach – a case study in intensive care monitoring. In: Proceedings of the 16th International Conference on Machine Learning (ICML-99)
P Mundra M Kumar KK Kumar VK Jayaraman BD Kulkarni (2007) ArticleTitleUsing pseudo amino acid composition to predict protein subnuclear localization: approached with PSSM source Pattern Recogn Lett 28 1610–1615 Occurrence Handle10.1016/j.patrec.2007.04.001
B Niu YD Cai WC Lu GY Zheng KC Chou (2006) ArticleTitlePredicting protein structural class with AdaBoost learner Protein Peptide Lett 13 489–492 Occurrence Handle10.2174/092986606776819619 Occurrence Handle1:CAS:528:DC%2BD28XlsVGqs7o%3D
Y Ofran B Rost (2003) ArticleTitlePredicted protein–protein interaction sites from local sequence information FEBS Lett 544 236–239 Occurrence Handle12782323 Occurrence Handle10.1016/S0014-5793(03)00456-3 Occurrence Handle1:CAS:528:DC%2BD3sXktFWksro%3D
HB Shen KC Chou (2005) ArticleTitlePredicting protein subnuclear location with optimized evidence-theoretic K-nearest classifier and pseudo amino acid composition Biochem Biophys Res Commun 337 752–756 Occurrence Handle16213466 Occurrence Handle1:CAS:528:DC%2BD2MXhtFCjs7%2FI Occurrence Handle10.1016/j.bbrc.2005.09.117
HB Shen KC Chou (2006) ArticleTitleUsing ensemble classifier to identify membrane protein types Amino Acids 32 483–488 Occurrence Handle17031474 Occurrence Handle10.1007/s00726-006-0439-2 Occurrence Handle1:CAS:528:DC%2BD2sXlsVGnsLY%3D
HB Shen KC Chou (2007a) ArticleTitleEzyPred: a top-down approach for predicting enzyme functional classes and subclasses Biochem Biophys Res Commun 364 53–59 Occurrence Handle10.1016/j.bbrc.2007.09.098 Occurrence Handle1:CAS:528:DC%2BD2sXht1aktbvF
HB Shen KC Chou (2007b) ArticleTitleGpos-PLoc: an ensemble classifier for predicting subcellular localization of Gram-positive bacterial proteins Protein Eng Des Sel 20 39–46 Occurrence Handle10.1093/protein/gzl053 Occurrence Handle1:CAS:528:DC%2BD2sXhvFWmtr8%3D
HB Shen KC Chou (2007c) ArticleTitleHum-mPLoc: an ensemble classifier for large-scale human protein subcellular location prediction by incorporating samples with multiple sites Biochem Biophys Res Commun 355 1006–1011 Occurrence Handle10.1016/j.bbrc.2007.02.071 Occurrence Handle1:CAS:528:DC%2BD2sXivVahur0%3D
Shen HB, Chou KC (2007d) Nuc-PLoc: a new web-server for predicting protein subnuclear localization by fusing PseAA composition and PsePSSM. Protein Eng Des Sel (DOI: 1093/protein/hzm057)
HB Shen KC Chou (2007e) ArticleTitleSignal-3L: a 3-layer approach for predicting signal peptide Biochem Biophys Res Commun 363 297–303 Occurrence Handle10.1016/j.bbrc.2007.08.140 Occurrence Handle1:CAS:528:DC%2BD2sXhtFWgsrbJ
HB Shen KC Chou (2007f) ArticleTitleUsing ensemble classifier to identify membrane protein types Amino Acids 32 483–488 Occurrence Handle10.1007/s00726-006-0439-2 Occurrence Handle1:CAS:528:DC%2BD2sXlsVGnsLY%3D
HB Shen KC Chou (2007g) ArticleTitleVirus-PLoc: a fusion classifier for predicting the subcellular localization of viral proteins within host and virus-infected cells Biopolymers 85 233–240 Occurrence Handle10.1002/bip.20640 Occurrence Handle1:CAS:528:DC%2BD2sXhvFWhs70%3D
HB Shen J Yang KC Chou (2007a) ArticleTitleEuk-PLoc: an ensemble classifier for large-scale eukaryotic protein subcellular location prediction Amino Acids 33 57–67 Occurrence Handle10.1007/s00726-006-0478-8 Occurrence Handle1:CAS:528:DC%2BD2sXotVWru7Y%3D
HB Shen J Yang KC Chou (2007b) ArticleTitleReview: methodology development for predicting subcellular localization and other attributes of proteins Expert Rev Proteomics 4 453–463 Occurrence Handle10.1586/14789450.4.4.453 Occurrence Handle1:CAS:528:DC%2BD2sXptFyrtLo%3D
JY Shi SW Zhang Q Pan YM Cheng J Xie (2007) ArticleTitlePrediction of protein subcellular localization by support vector machines using multi-scale energy and pseudo amino acid composition Amino Acids 33 69–74 Occurrence Handle17235454 Occurrence Handle10.1007/s00726-006-0475-y Occurrence Handle1:CAS:528:DC%2BD2sXotVWru7g%3D
XD Sun RB Huang (2006) ArticleTitlePrediction of protein structural classes using support vector machines Amino Acids 30 469–475 Occurrence Handle16622605 Occurrence Handle10.1007/s00726-005-0239-0 Occurrence Handle1:CAS:528:DC%2BD28Xls1ehu7c%3D
Tan F, Feng X, Fang Z, Li M, Guo Y, Jiang L (2007) Prediction of mitochondrial proteins based on genetic – algorithm partial least squares and support vector machine. Amino Acids (DOI: 10.1007/s00726-006-0465-0)
M Terribilini JH Lee C Yan RL Jernigan V Honavar D Dobbs (2006) ArticleTitlePrediction of RNA binding sites in proteins from amino acid sequence RNA 12 1–13 Occurrence Handle10.1261/rna.2197306 Occurrence Handle1:CAS:528:DC%2BD28XnslCmtrY%3D
M Treger E Westhof (2001) ArticleTitleStatistical analysis of atomic contacts at RNA-protein interfaces J Mol Recogn 14 199–214 Occurrence Handle10.1002/jmr.534 Occurrence Handle1:CAS:528:DC%2BD3MXmslSmsLs%3D
V Vapnik (1998) The nature of statistical learning theory Springer New York
Wang LJ, Brown SJ (2006) BindN: a web-based tool for efficient prediction of DNA and RNA binding sites in amino acid sequences. Nucleic Acids Res Web Server Issue: W243–W248
M Wang J Yang GP Liu ZJ Xu KC Chou (2004) ArticleTitleWeighted-support vector machines for predicting membrane protein types based on pseudo-amino acid composition Protein Eng Des Sel 17 509–516 Occurrence Handle15314209 Occurrence Handle10.1093/protein/gzh061 Occurrence Handle1:CAS:528:DC%2BD2cXos1GisLY%3D
M Wang J Yang KC Chou (2005) ArticleTitleUsing string kernel to predict signal peptide cleavage site based on subsite coupling model Amino Acids 28 395–402 Occurrence Handle15838592 Occurrence Handle10.1007/s00726-005-0189-6 Occurrence Handle1:CAS:528:DC%2BD2MXlt1KmtbY%3D
Z Wen M Li Y Li Y Guo K Wang (2006) ArticleTitleDelaunay triangulation with partial least squares projection to latent structures: a model for G-protein coupled receptors classification and fast structure recognition Amino Acids 32 277–283 Occurrence Handle16729188 Occurrence Handle10.1007/s00726-006-0341-y Occurrence Handle1:CAS:528:DC%2BD2sXhtFyhtLY%3D
X Xiao KC Chou (2007) ArticleTitleDigital coding of amino acids based on hydrophobic index Protein Peptide Lett 14 871–875 Occurrence Handle10.2174/092986607782110293 Occurrence Handle1:CAS:528:DC%2BD2sXhsVWrs7jO
X Xiao SH Shao YS Ding ZD Huang Y Huang KC Chou (2005) ArticleTitleUsing complexity measure factor to predict protein subcellular location Amino Acids 28 57–61 Occurrence Handle15611847 Occurrence Handle10.1007/s00726-004-0148-7 Occurrence Handle1:CAS:528:DC%2BD2MXhsVKqsro%3D
X Xiao SH Shao ZD Huang KC Chou (2006) ArticleTitleUsing cellular automata images and pseudo amino acid composition to predict protein subcellular location Amino Acids 30 49–54 Occurrence Handle16044193 Occurrence Handle10.1007/s00726-005-0225-6 Occurrence Handle1:CAS:528:DC%2BD28XhsFCksrk%3D
X Xiao SH Shao ZD Huang KC Chou (2006) ArticleTitleUsing pseudo amino acid composition to predict protein structural classes: approached with complexity measure factor J Comput Chem 27 478–482 Occurrence Handle16429410 Occurrence Handle10.1002/jcc.20354 Occurrence Handle1:CAS:528:DC%2BD28XitFyqsr4%3D
ZR Yang KC Chou (2004) ArticleTitleBio-support vector machines for computational proteomics Bioinformatics 20 735–741 Occurrence Handle14751989 Occurrence Handle10.1093/bioinformatics/btg477 Occurrence Handle1:CAS:528:DC%2BD2cXitlyhs7w%3D
SW Zhang Q Pan HC Zhang ZC Shao JY Shi (2006) ArticleTitlePrediction protein homo-oligomer types by pseudo amino acid composition: approached with an improved feature extraction and naive Bayes feature fusion Amino Acids 30 461–468 Occurrence Handle16773245 Occurrence Handle10.1007/s00726-006-0263-8 Occurrence Handle1:CAS:528:DC%2BD28Xls1egsr0%3D
Zhang TL, Ding YS (2007) Using pseudo amino acid composition and binary-tree support vector machines to predict protein structural classes. Amino Acids (DOI: 10.1007/s00726-007-0496-1)
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors’ address: Yan Wang, Institute of Biophysics and Biochemistry, School of Life Science, Huazhong University of Science and Technology, Wuhan City 430074, China
Rights and permissions
About this article
Cite this article
Wang, Y., Xue, Z., Shen, G. et al. PRINTR: Prediction of RNA binding sites in proteins using SVM and profiles. Amino Acids 35, 295–302 (2008). https://doi.org/10.1007/s00726-007-0634-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-007-0634-9