Abstract
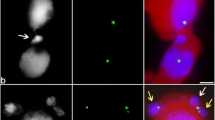
The migration of nuclei between plant cells (cytomixis) is a mysterious cellular phenomenon frequently observable in the male meiosis of higher plants. Cytomixis attracts attention because of unknown cellular mechanisms underlying migration of nuclei and its potential evolutionary significance, since the genetic material is transferred between the cells that form pollen. Although cytomixis was discovered over a century ago, the advance in our understanding of this process has been rather insignificant because of methodological difficulties. The data that allowed for a new insight into this phenomenon were obtained by examining the migrating nuclei with electron and confocal laser microscopy, immunostaining, and fluorescence in situ hybridization. As has been shown, the chromatin migrating between cells is surrounded by an undamaged nuclear membrane. Such chromatin does not undergo heterochromatization and contains normal euchromatin markers. The condensation degree of the migrating chromatin corresponds to the current meiotic stage, and normal structures of synaptonemal complex are present in the migrating part of the nucleus. The cells involved in cytomixis lack any detectable morphological and molecular markers of programmed cell death. It has been shown that individual chromosomes and genomes (in the case of allopolyploids) have no predisposition to the migration between cells, i.e., parts of the nucleus are involved in cytomixis in a random manner. However, the fate of migrating chromatin after it has entered the recipient cell is still vague. A huge amount of indirect data suggests that migrating chromatin is incorporated into the nucleus of the recipient cell; nonetheless, the corresponding direct evidences are still absent. No specific markers of cytomictic chromatin have been yet discovered. Thus, the causes and consequences of cytomixis are still disputable. This review briefs the recent data on the relevant issues, describes the classical and modern methodological approaches to analysis of the intercellular migration of nuclei, and discusses the problems in cytomixis research and its prospects.


Similar content being viewed by others
References
Aksic MF, Cerovic R, Ercisli S, Jensen M (2016) Microsporogenesis and meiotic abnormalities in different ‘Oblacinska’ sour cherry (Prunus cerasus L.) clones. Flora Morphol Distrib Funct Ecol. Plants 219:25–34. https://doi.org/10.1016/j.flora.2015.12.009
Alka AMYK, Bhat TM et al (2012) Induction of polyploid mutant in Linum usitatissimum L. by cytomixis. Golden Res Thoughts 2:1–6. https://doi.org/10.9780/22315063
Arnoldy W (1900) Beiträge zur Morphologie der Gymnospermen. IV. Was sind die ‘Keimbläschen’ oder ‘Hofmeisters-Körperchen. In: der Eizelle der Abietineen? Flora 87, pp 194–204
Bala S, Gupta RC (2011) Effect of secondary associations on meiosis, pollen fertility and pollen size in cape gooseberry (Physalis peruviana L.) Chrom Bot 6(2):25–28. https://doi.org/10.3199/iscb.6.25
Barton DA, Cantrill LC, Law AMK et al (2014) Chilling to zero degrees disrupts pollen formation but not meiotic microtubule arrays in Triticum aestivum L. Plant Cell Environ 37(12):2781–2794. https://doi.org/10.1111/pce
Chicheportiche A, Bernardino-Sgherri J, de Massy B, Dutrillaux B (2007) Characterization of Spo11-dependent and independent phospho-H2AX foci during meiotic prophase I in the male mouse. J Cell Sci 120(10):1733–1742. https://doi.org/10.1242/jcs.004945
De Storme N, Mason A (2014) Plant speciation through chromosome instability and ploidy change: cellular mechanisms, molecular factors and evolutionary relevance. Curr Plant Biol 1:10–33. https://doi.org/10.1016/j.cpb.2014.09.002
Farooq U, Lovleen, Saggoo MIS (2014) Male meiosis and behaviour of sex chromosomes in different populations of Rumex acetosa L. from the Western Himalayas, India. Plant Syst Evol 300(2):287–294. https://doi.org/10.1007/s00606-013-0881-z
Feijo JA, Pais MS (1989) Cytomixis in meiosis during the microsporogenesis of Ophrys lutea: an ultrastructural study. Caryologia 42(1):37–48. https://doi.org/10.1080/00087114.1989.10796951
Ferreira K, Torres GA, Carvalho IV, Davide LC (2009) Abnormal meiotic behavior in three species of Crotalaria. Pesqui Agropecuária Bras 44(12):1641–1646. https://doi.org/10.1590/S0100-204X2009001200012
Fuchs J, Schubert I (2012) Chromosomal distribution and functional interpretation of epigenetic histone marks in plants. Plant Cytogenetics:231–253
Fuentes I, Stegemann S, Golczyk H, Karcher D, Bock R (2014) Horizontal genome transfer as an asexual path to the formation of new species. Nature 511(7508):232–235. https://doi.org/10.1038/nature13291
Gates RR (1911) Pollen formation in Oenothera gigas. Ann Bot 25:909–940
Gerdes HH, Bukoreshtliev NV, Barroso JF (2007) Tunneling nanotubes: a new route for the exchange of components between animal cells. FEBS Lett 581(11):2194–2201. https://doi.org/10.1016/j.febslet.2007.03.071
Ghaffari SM (2006) Occurrence of diploid and polyploid microspores in Sorghum bicolor (Poaceae) is the result of cytomixis. Afr J Biotech 5:1450–1453
Giorgetti L, Castiglione M, Martini G et al (2007) Methylated DNA sequence extrusion during plant early meiotic prophase. Caryologia 60(3):279–289. https://doi.org/10.1080/00087114.2007.10797949
Goday C, Pigozzi MI (2010) Heterochromatin and histone modifications in the germline-restricted chromosome of the zebra finch undergoing elimination during spermatogenesis. Chromosoma 119(3):325–336. https://doi.org/10.1007/s00412-010-0260-2
Griffiths N, Jaipargas E, Wozny M et al (2016) Photo-convertible fluorescent proteins as tools for fresh insights on subcellular interactions in plants. J Microsc 263(2):148–157. https://doi.org/10.1111/jmi.12383
Guan JZ, Wang JJ, Cheng ZH et al (2012) Cytomixis and meiotic abnormalities during microsporogenesis are responsible for male sterility and chromosome variations in Houttuynia cordata. Genet Mol Res 11:121–130 doi: 0.4238/2012.January.17.2
Gupta H, Gupta RC, Kumar R, Singhal VK (2017) A profile of chromosome counts, male meiosis and pollen fertility in 45 species of Asteraceae from Parvati Valley in Kullu district, Himachal Pradesh. Caryologia 70(2):128–140. https://doi.org/10.1080/00087114.2017.1299976
Gurdona C, Svab Z, Feng Y et al (2016) Cell-to-cell movement of mitochondria in plants. PNAS 113(12):3395–3400. https://doi.org/10.1073/pnas.1518644113
Guzicka M, Wozny A (2005) Cytomixis in shoot apex of Norway spruce [Picea abies (L.) Karst.] Trees 18(6):722–724. https://doi.org/10.1007/s00468-004-0331-1
He S, Yan S, Wang P, Zhu W, Wang X, Shen Y, Shao K, Xin H, Li S, Li L (2014) Comparative analysis of genome-wide chromosomal histone modification patterns in maize cultivars and their wild relatives. PLoS One 9(5):e97364. https://doi.org/10.1371/journal.pone.0097364
He Y, Wang C, Higgins J et al (2016) MEIOTIC F-BOX is essential for male meiotic DNA double strand break repair in rice. Plant Cell 28(8):1879–1893. https://doi.org/10.1105/tpc.16.00108
Hedde PN, Nienhaus GU (2014) Super-resolution localization microscopy with photoactivatable fluorescent marker proteins. Protoplasma 251(2):349–362. https://doi.org/10.1007/s00709-013-0566-z
Houben A, Demidov D, Caperta AD, Karimi R, Agueci F, Vlasenko L (2007) Phosphorylation of histone H3 in plants—a dynamic affair. Biochim Biophys Acta-Gene Struct Expr 1769(5-6):308–315. https://doi.org/10.1016/j.bbaexp.2007.01.002
Houben A, Demidov D, Rutten T, Scheidtmann KH (2005) Novel phosphorylation of histone H3 at threonine 11 that temporally correlates with condensation of mitotic and meiotic chromosomes in plant cells. Cytogenet Genome Res 109(1-3):148–155. https://doi.org/10.1159/000082394
Hunter N, Börner GV, Lichten M, Kleckner N (2001) Gamma-H2AX illuminates meiosis. Nat Genet 27(3):236–238. https://doi.org/10.1038/85781
Kalinka A, Achrem M, Rogalska SM (2010) Cytomixis-like chromosomes/chromatin elimination from pollen mother cells (PMCs) in wheat-rye allopolyploids. Nucleus 53(3):69–83. https://doi.org/10.1007/s13237-010-0002-0
Kawashima SA, Yamagishi Y, Honda T, Ishiguro K, Watanabe Y (2010) Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing shugoshin. Science 327(5962):172–177. https://doi.org/10.1126/science.1180189
Kolczyk J, Tuleja M, Płachno BJ (2015) Histological and cytological analysis of microsporogenesis and microgametogenesis of the invasive species Galinsoga quadriradiata Ruiz & Pav. (Asteraceae). Acta Biol Cracoviensia Ser Bot 57(2):89–97. https://doi.org/10.1515/abcsb-2015-0018
Koyanagi M, Brandes RP, Haendeler J, Zeiher AM, Dimmeler S (2005) Cell-to-cell connection of endothelial progenitor cells with cardiac myocytes by nanotubes: a novel mechanism for cell fate changes? Circ Res 96(10):1039–1041. https://doi.org/10.1161/01.RES.0000168650.23479.0c
Kravets E (2013) Cytomixis and its role in the regulation of plant fertility. Russ J Devel Biol 44(3):113–128. https://doi.org/10.1134/S1062360413030028
Kravets E (2011) The role of cell selection for pollen grain fertility after treatment of barley sprouts (Hordeum distichum L.) with UV-B irradiation. Acta Biol Slov 54:31–41
Kumar G, Srivastava N (2013) Induced cytomictic variations in pollen mother cells of Sesbania cannabina Poir. J Cent Eur Agric 14(3):19–27. https://doi.org/10.5513/JCEA01/14.3.1280
Kumar P, Singhal V (2016a) Nucleoli migration coupled with cytomixis. Biologia 71(6):651–659. https://doi.org/10.1515/biolog-2016-0076
Kumar P, Singhal V (2016b) Morphological and ecological adaptations, and cytological studies in Astragalus rhizanthus Royle ex Benth. (Papilionaceae), an endemic to Himalayas. Cytologia 81(2):155–160. https://doi.org/10.1508/cytologia.81.155
Kumar P, Singhal VK, Kaur D, Kaur S (2010) Cytomixis and associated meiotic abnormalities affecting pollen fertility in Clematis orientalis. Biol Plant 54(1):181–184. https://doi.org/10.1007/s10535-010-0031-1
Kumar P, Singhal V, Srivastava S (2017) First detection of cytomixis and its consequences in Thalictrum cultratum Wall. (Ranunculaceae). Cytol Genet 51(5):384–390. https://doi.org/10.3103/S0095452717050061
Kumar S, Jeelani SM, Rani S, Gupta RC, Kumari S (2013) Cytology of five species of subfamily Papaveroideae from the Western Himalayas. Protoplasma 250(1):307–316. https://doi.org/10.1007/s00709-012-0413-7
Kuras M, Nowakowska J, Sliwinska E et al (2006) Changes in chromosome structure, mitotic activity and nuclear DNA content from cells of Allium Test induced by bark water extract of Uncaria tomentosa (Willd.) DC. J Ethnopharmacol 107(2):211–221. https://doi.org/10.1016/j.jep.2006.03.018
Lavia GI, Ortiz AM, Robledo G, Fernández A, Seijo G (2011) Origin of triploid Arachis pintoi (Leguminosae) by autopolyploidy evidenced by FISH and meiotic behaviour. Ann Bot 108(1):103–111. https://doi.org/10.1093/aob/mcr108
Li XF, Song ZQ, Feng DS, Wang HG (2009) Cytomixis in Thinopyrum intermedium, Thinopyrum ponticum and its hybrids with wheat. Cer Res Comm 37(3):353–361. https://doi.org/10.1556/CRC.37.2009.3.4
Lim KY, Matyásek R, Lichtenstein CP, Leitch AR (2000) Molecular cytogenetic analyses and phylogenetic studies in the Nicotiana section Tomentosae. Chromosoma 109(4):245–258. https://doi.org/10.1007/s004120000074
Lone FA, Lone S (2013) Cytomixis – a well known but less understood phenomenon in plants. Int J Recent Sci Res 4:347–352
Malik R, Gupta R, Singh V et al (2017) New chromosome reports in Lamiaceae of Kashmir (Northwest Himalaya), India. Protoplasma 254(2):971–985. https://doi.org/10.1007/s00709-016-1006-7
Mandal A, Datta AK, Gupta S, Paul R, Saha A, Ghosh BK, Bhattacharya A, Iqbal M (2013) Cytomixis—a unique phenomenon in animal and plant. Protoplasma 250(5):985–996. https://doi.org/10.1007/s00709-013-0493-z
Mandal D, Nandi A (2017) Cytomixis with associated chromosomal anomalies and the reproduction of Chlorophytum borivilianum Santapau & R. R Fern Taiwania 62:211–215. https://doi.org/10.6165/tai.2017.62.211
Manzanero S, Arana P, Puertas MJ, Houben A (2000) The chromosomal distribution of phosphorylated histone H3 differs between plants and animals at meiosis. Chromosoma 109(5):308–317. https://doi.org/10.1007/s004120000087
Mursalimov S, Deineko E (2017) Cytomixis in tobacco microsporogenesis: are there any genome parts predisposed to migration? Protoplasma 254(3):1379–1384. https://doi.org/10.1007/s00709-016-1028-1
Mursalimov S, Permyakova N, Deineko E, Houben A, Demidov D (2015) Cytomixis doesn’t induce obvious changes in chromatin modifications and programmed cell death in tobacco male meiocytes. Front Plant Sci 6:1–13. https://doi.org/10.3389/fpls.2015.00846
Mursalimov S, Sidorchuk Y, Deineko E (2013a) The role of spherosome-like vesicles in formation of cytomictic channels between tobacco microsporocytes. Biol Plant 57(2):291–297. https://doi.org/10.1007/s10535-012-0276-y
Mursalimov S, Sidorchuk Y, Deineko E (2017a) Analysis of cytomixis in tobacco microsporocytes with confocal laser scanning microscopy. Protoplasma 254(1):539–545. https://doi.org/10.1007/s00709-016-0973-z
Mursalimov S, Sidorchuk Y, Demidov D, Meister A, Deineko E (2016) A rise of ploidy level influences the rate of cytomixis in tobacco male meiosis. Protoplasma 253(6):1583–1588. https://doi.org/10.1007/s00709-015-0907-1
Mursalimov S, Zagorskaya A, Deineko E (2017b) Evaluation of DNA damage in tobacco male meiocytes involved in cytomixis using comet assay. Protoplasma. https://doi.org/10.1007/s00709-017-1144-6
Mursalimov SR, Baiborodin SI, Sidorchuk YV, Shumny VK, Deineko EV (2010) Characteristics of the cytomictic channel formation in Nicotiana tabacum L. pollen mother cells. Cytol Genet 44(1):14–18. https://doi.org/10.3103/S0095452710010032
Mursalimov SR, Deineko EV (2015) How cytomixis can form unreduced gametes in tobacco. Plant Syst Evol 301(4):1293–1297. https://doi.org/10.1007/s00606-014-1150-5
Mursalimov SR, Deineko EV (2011) An ultrastructural study of cytomixis in tobacco pollen mother cells. Protoplasma 248(4):717–724. https://doi.org/10.1007/s00709-010-0234-5
Mursalimov SR, Sidorchuk YV, Deineko EV (2013b) New insights into cytomixis: specific cellular features and prevalence in higher plants. Planta 238(3):415–423. https://doi.org/10.1007/s00425-013-1914-0
Negron-Ortiz V (2007) Chromosome numbers, nuclear DNA content, and polyploidy in Consolea (Cactaceae), an endemic cactus of the Caribbean Islands. Am J Bot 94(8):1360–1370. https://doi.org/10.3732/ajb.94.8.1360
Ogienko A, Karagodin D, Pavlova N et al (2008) Molecular and genetic description of a new hypomorphic mutation of Trithorax-like gene and analysis of its effect on Drosophila melanogaster oogenesis. Russ J Dev Biol 39(2):108–115. https://doi.org/10.1134/S1062360408020070
Papini A, Tani G, Di Falco P, Brighigna L (2010) The ultrastructure of the development of Tillandsia (Bromeliaceae) trichome. Flora Morphol Distrib Funct Ecol Plants 205(2):94–100. https://doi.org/10.1016/j.flora.2009.02.001
Pécrix Y, Rallo G, Folzer H et al (2011) Polyploidization mechanisms: temperature environment can induce diploid gamete formation in Rosa sp. J Exp Bot 62(10):3587–3597. https://doi.org/10.1093/jxb/err052
Pierre P, Sousa S (2011) Citomixia em plantas: causas, mecanismos e consequências. Brazilian. J Biosci 9:231–240
Polowick PL, Sawhney VK (1992) Ultrastructural changes in the cell wall, nucleus and cytoplasm of pollen mother cells during meiotic prophase I in Lycopersicon esculentum (Mill.) Protoplasma 169(3-4):139–147. https://doi.org/10.1007/BF01323613
Qiu Y, Liao L, Liu S, Mao D, Liu R (2017) Differences on the microsporogenesis and tapetal development of male fertile and cytoplasmic male sterile pepper (Capsicum annuum L.) Grana 56(3):215–227. https://doi.org/10.1080/00173134.2016.1248860
Reis A, Sousa S, Viccini L (2016) High frequency of cytomixis observed at zygotene in tetraploid Lippia alba. Plant Syst Evol 302(1):121–127. https://doi.org/10.1007/s00606-015-1249-3
Ressayre A, Mignot A, Siljak-Yakovlev S, Raquin C (2003) Postmeiotic cytokinesis and pollen aperture number determination in eudicots: effect of the cleavage wall number. Protoplasma 221(3-4):257–268. https://doi.org/10.1007/s00709-002-0075-y
Risso-Pascotto C, Pagliarini MS, do Valle CB (2009) Chromosome number and microsporogenesis of two accessions of Brachiaria dura Stapf (Poaceae). Biota Neotrop 9(2):1–5. https://doi.org/10.1590/S1676-06032009000200024
Rogakou EP, Boon C, Redon C, Bonner WM (1999) Megabase chromatin domains involved in DNA double-strand breaks in vivo. J Cell Biol 146(5):905–915. https://doi.org/10.1083/jcb.146.5.905
Schattat MH, Barton KA, Mathur J (2014) The myth of interconnected plastids and related phenomena. Protoplasma 252(1):359–371. https://doi.org/10.1007/s00709-014-0666-4
Schoenmakers S, Wassenaar E, Laven JSE, Grootegoed JA, Baarends WM (2010) Meiotic silencing and fragmentation of the male germline restricted chromosome in zebra finch. Chromosoma 119(3):311–324. https://doi.org/10.1007/s00412-010-0258-9
Shibata F, Nagaki K, Yokota E, Murata M (2013) Tobacco karyotyping by accurate centromere identification and novel repetitive DNA localization. Chromosom Res 21(4):375–381. https://doi.org/10.1007/s10577-013-9363-y
Sidorchuk YV, Deineko EV, Shumny VK (2007a) Peculiarities of cytomixis in pollen mother cells of transgenic tobacco plants (Nicotiana tabacum L.) with mutant phenotype. Cell Tiss Biol 1(6):570–576. https://doi.org/10.1134/S1990519X07060144
Sidorchuk YV, Deineko EV, Shumny VK (2007b) Role of microtubular cytoskeleton and callose walls in the manifestation of cytomixis in pollen mother cells of tobacco Nicotiana tabacum L. Cell Tiss Biol 1(6):577–581. https://doi.org/10.1134/S1990519X07060156
Sidorchuk YV, Novikovskaya AA, Deineko EV (2016) Cytomixis in the cereal (Gramineae) microsporogenesis. Protoplasma 253(2):291–298. https://doi.org/10.1007/s00709-015-0807-4
Silva N, Machado MFPS, Mangolin CA, Pagliarini MS (2006) Microsporogenesis in somaclones of Cereus peruvianus Mill. (Cactaceae). J Plant Sci 1(1):8–13. https://doi.org/10.3923/jps.2006.8.13
Silva SL, Magalhaes KM, Carvalho R (2017) Karyotype variations in seagrass (Halodule wrightii Ascherson—Cymodoceaceae). Aquat Bot 136:52–55. https://doi.org/10.1016/j.aquabot.2016.09.005
Singhal VK, Kumar P (2008) Impact of cytomixis on meiosis, pollen viability and pollen size in wild populations of Himalayan poppy (Meconopsis aculeata Royle). J Biosci 33(3):371–380. https://doi.org/10.1007/s12038-008-0057-0
Singhal VK, Rana PK, Kumar P, Kaur D (2011) Persistent occurrence of meiotic abnormalities in a new hexaploid cytotype of Thalictrum foetidum from Indian cold deserts. Biologia 66(3):458–464. https://doi.org/10.2478/s11756-011-0033-2
Stegemann S, Bock R (2009) Exchange of genetic material between cells in plant tissue grafts. Science (80- ) 324:649–651. doi: https://doi.org/10.1126/science.1170397
Stegemann S, Keuthe M, Greiner S, Bock R (2012) Horizontal transfer of chloroplast genomes between plant species. PNAS 109:2434–2438. doi: DOI https://doi.org/10.1073/pnas.1114076109
Tarkowska J (1965) Experimental analysis of the mechanism of cytomixis. I. Cytomixis in vegetative tissues. Acta Soc Bot Pol 34:27–44
Tarkowska J (1966) Experimental analysis of the mechanism of cytomixis. II Cytomixis in the pollen mother cells of the lily—Lilium candidum L. Acta Soc Bot Pol 35:25–40
Tarkowska J (1973) The nature of cytomixis. Caryologia 25(sup1):151–157. https://doi.org/10.1080/00087114.1973.10797120
Thyssen G, Svab Z, Maliga P (2012) Cell-to-cell movement of plastids in plants. PNAS 109(7):2439–2443. https://doi.org/10.1073/pnas.1114297109
Tsvetova M, Elkonin L (2013) Cytological investigation of pollen development in Sorghum line with male sterility induced by sodium ascorbate in tissue culture. Am J Plant Sci 4(07):11–18. https://doi.org/10.4236/ajps.2013.47A1002
Wang CY, Li X, QF W, Wang X (2006) Cytoplasmic channels and their association with plastids in male meiocytes of tobacco, onion and lily. Cell Biol Int 30(5):406–411. https://doi.org/10.1016/j.cellbi.2006.01.003
Wang XY, Nie XW, Guo GQ, Pan YF, Zheng GC (Kuo Chang (2002) Ultrastructural characterization of the cytoplasmic channel formation between pollen mother cells of David lily. Caryologia 55(2):161–169. https://doi.org/10.1080/00087114.2002.10589272
Wang XY, Yu CH, Li X, et al (2004) Ultrastructural aspects and possible origin of cytoplasmic channels providing intercellular connection in vegetative tissues of anthers Russ J Plant Physiol 51:110–120. doi: 1021–4437/04/5101–0097
Whelan EDP (1974) Discontinuities in the callose wall, intermeiocyte connections, and cytomixis in angiosperm meiocytes. Can J Bot 52(6):1219–1224. https://doi.org/10.1139/b74-157
Wozny M, Schattat M, Mathur N et al (2012) Color recovery after photoconversion of H2B::mEosFP allows detection of increased nuclear DNA content in developing plant cells. Plant Physiol 158(1):95–106. https://doi.org/10.1104/pp.111.187062
Xiao A, Li H, Shechter D, Ahn SH, Fabrizio l, Erdjument-Bromage H, Ishibe-Murakami S, Wang B, Tempst P, Hofmann K, Patel DJ, Elledge SJ, Allis CD (2009) WSTF regulates the H2A.X DNA damage response via a novel tyrosine kinase activity. Nature 457(7225):57–62. https://doi.org/10.1038/nature07668
Xue Z, Li Y, Zhang L, Shi W, Zhang C, Feng M, Zhang F, Tang D, Yu H, Gu M, Cheng Z (2016) OsMTOPVIB promotes meiotic DNA double-strand break formation in rice. Mol Plant 9(11):1535–1538. https://doi.org/10.1016/j.molp.2016.07.005
Yu CH, Guo GQ, Nie XW, Zheng GC (2004) Cytochemical localization of pectinase activity in pollen mother cells of tobacco during meiotic prophase and its relation to the formation of secondary plasmodesmata and cytoplasmic channels. Acta Bot Sin 46:1443–1453
Zani BG, Edelman ER (2010) Cellular bridges: routes for intercellular communication and cell migration. Comm Integr Biol 3(3):215–220. https://doi.org/10.4161/cib.3.3.11659
Zhang WC, Yan WM, Lou CH (1990) Intercellular movement of protoplasm in vivo in developing endosperm of wheat caryopses. Protoplasma 153(3):193–203. https://doi.org/10.1007/BF01354004
Zhou SQ (2003) Viewing the difference between the diploid and the polyploid in the light of the upland cotton aneuploid. Hereditas 138(1):65–72. https://doi.org/10.1034/j.1601-5223.2003.01689.x
Acknowledgments
The work was supported by the Russian Foundation for Basic Research [16-34-60007 mol_a_dk] and Siberian Branch of the Russian Academy of Science under the program “Molecular genetic bases of regulation of genes expression, morphology, differentiation and cell reprogramming” [0324-2016-0003].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Jaideep Mathur
Electronic supplementary material
ESM 1
(DOCX 26 kb)
Rights and permissions
About this article
Cite this article
Mursalimov, S., Deineko, E. Cytomixis in plants: facts and doubts. Protoplasma 255, 719–731 (2018). https://doi.org/10.1007/s00709-017-1188-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-017-1188-7