Abstract
Coronaviruses (CoV) are a family of viral pathogens that infect both birds and mammals, including humans. Seven human coronaviruses (HCoV) have been recognized so far. HCoV-229E, -OC43, -NL63, and -HKU1 account for one-third of common colds with mild symptoms. The other three members are severe acute respiratory syndrome (SARS)-CoV, Middle East respiratory syndrome (MERS)-CoV, and SARS-CoV-2. These viruses are responsible for SARS, MERS, and CoV disease 2019 (COVID-19), respectively. A variety of diagnostic techniques, including chest X-rays, computer tomography (CT) scans, analysis of viral nucleic acids, proteins, or whole virions, and host antibody detection using serological assays have been developed for the detection of these viruses. In this review, we discuss conventional serological tests, such as enzyme-linked immunosorbent assay (ELISA), western blot (WB), immunofluorescence assay (IFA), lateral flow immunoassay (LFIA), and chemiluminescence immunoassay (CLIA), as well as biosensor-based assays that have been developed for diagnosing HCoV-associated diseases since 2003, with an in-depth focus on COVID-19.
Similar content being viewed by others
Introduction
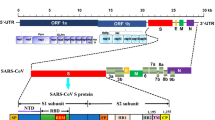
Virus-induced diseases are pervasive worldwide, ranging from mild infections to lethal diseases. Among the diverse families of viruses is the family Coronaviridae, of the order Nidovirales. Orthocoronavirinae is a subfamily of the family Coronaviridae whose members infect birds and mammals, including humans [1, 2]. Human coronaviruses (HCoVs) are responsible for various respiratory diseases, e.g., common colds, bronchitis, and pneumonia [3]. Urban sprawl and farming have contributed to the rapid pace at which these viruses have been evolving [4]. In general, coronaviruses (CoVs) are enveloped viruses with a non-segmented, positive-sense single-stranded RNA genome that is approximately 30 kb in length and encodes four structural proteins; the membrane (M) glycoprotein, the small envelope (E) protein, the spike (S) glycoprotein, and the nucleocapsid (N) phosphoprotein, which encapsulates the viral RNA. The M and E proteins are vital for viral envelope formation and maintenance of virion structure [5, 6]. The S protein, which has a molecular weight of about 180 kDa and a length of 20 nm, consists of two subunits, S1 and S2. The S1 subunit, which contains the receptor-binding domain (RBD), recognizes and binds to its receptor angiotensin-converting enzyme 2 (ACE2), which is found in human lungs. The S2 subunit, by undergoing specific conformational changes, induces membrane fusion, enabling viral entry into the host cell. This protein is the chief target of neutralizing antibodies, which prevent infection and further dissemination by blocking binding to ACE2. The 40- to 60-kDa N protein is the viral protein that is produced and shed in the largest quantities during infection [7,8,9,10,11].
In the case of SARS-CoV-2, the immune system initially generates an uncontrolled response through the hyperactivation of monocytes and macrophages, which leads to an increase in the number of neutrophils and interleukin 6 (IL-6) and a reduction in the lymphocyte population. For the adaptive immune response, T and B lymphocytes are necessary for cell-mediated and humoral immunity, respectively. In cell-mediated immunity, cytotoxic T cells identify and eradicate infected cells, while in humoral immunity, B cells produce neutralizing, immunoglobulin G (IgG), and immunoglobulin M (IgM) antibodies that block viral particles from binding to host cells [12, 13]. IgG and IgM antibodies are generally produced shortly after exposure to the virus, with IgG usually being secreted later than IgM. Zhou et al. demonstrated that the maximum IgM level is reached 9 days after infection and that secretion of IgG antibodies begins during the second week [14, 15]. These antibodies are commonly produced against the viral S, E, or N proteins [16,17,18].
In severe COVID-19 cases, a condition termed “cytokine storm” has been observed. This syndrome results from an excessive uncontrolled inflammatory response that leads to the overproduction of pro-inflammatory cytokines, particularly IL-6. Patients experiencing a cytokine storm may suffer from acute respiratory distress syndrome (ARDS), tissue damage, or organ failure, which can have a fatal outcome [19].
To date, seven CoVs have been identified in humans, including HCoV-229E and -OC43 in the 1960s, SARS-CoV in November 2002, HCoV-NL63 in 2004, HCoV-HKU1 in 2005, MERS-CoV in June 2012, and SARS-CoV-2 in December 2019 [5, 14, 20]. HCoV-229E, -OC43, -NL63, and -HKU1 account for one-third of common colds, often producing mild symptoms, but in some cases, they can cause bronchitis or pneumonia [21]. SARS-CoV, MERS-CoV, and SARS-CoV-2, which are more likely to produce severe symptoms, are responsible for SARS, MERS, and COVID-19, which are life-threatening infectious respiratory tract diseases [22].
More than 8000 SARS-associated cases and 770 deaths were reported during the 2002-2004 SARS epidemic in 29 countries, with a case-fatality rate of roughly 9%. The last SARS-related case was reported in April 2004, and since then, no new cases have been reported [23,24,25].
MERS-CoV, which is closely related to Tylonycteris bat CoV HKU4 and Pipistrellus bat CoV HKU5, may infect humans via direct or indirect contact with dromedary camels. Human- and camel-derived MERS-CoV have shown more than 99.5% nucleic acid sequence identity. Since its outbreak in June 2012 until the end of January 2020, 2519 confirmed MERS-related cases and 866 deaths have been reported. The case-fatality rate was approximately 34%, which is significantly higher than that of SARS [26, 27].
It is widely believed that SARS-CoV-2 was initially transmitted to humans via contact with animals at the Wuhan live animal and seafood market in China in December 2019. Since the genome sequence of this virus is approximately 89% identical to that of the bat SARS-like CoV ZXC21 and 96.2% similar to that of the bat CoV RaTG13, researchers have proposed that SARS-CoV-2 is of bat origin. Moreover, its genomic sequence is 79.6% identical to that of SARS-CoV, resulting in it being renamed from "2019 novel CoV" (2019-nCoV) to "SARS-CoV-2". By the end of August 2020, the ongoing COVID-19 pandemic resulted in more than 25 million confirmed cases and 850,000 deaths in over 213 countries and territories worldwide [15, 28,29,30,31,32,33,34].
In addition to the chest X-rays and CT scans employed to diagnose CoV-related diseases, several other techniques are used to identify the virus directly, by analyzing the viral RNA or proteins or the whole virus. These direct analytical methods include nucleic acid amplification tests (NAATs), such as reverse transcription polymerase chain reaction (RT-PCR) and isothermal nucleic acid amplification assays, ultrasensitive amplification-free RNA biosensors, and whole-virus or viral proteins assays [35,36,37,38,39,40,41,42, 121]. Furthermore, host antibody detection, which is the main focus of this review, is another technique that indirectly identifies the virus, by detecting antibodies that recognize viral particles [43, 44].
The detection of host antibodies produced against viral antigens in serum samples is a well-established method for determining the presence or absence of CoVs. These tests, known as serological assays, are frequently used by clinicians due to their rapidity and low cost when compared to the RT-PCR technique recommended for viral RNA detection. However, the sensitivity of these assays highly depends on the time of viral exposure, which determines the antibody level. These tests are also incapable of determining the precise time of disease onset [45, 46]. Serological assays may be used to determine the portion of the population who have had the disease and recovered from it and are therefore potentially immune to the infection. Although it has been suggested that screening for these antibodies might be used to identify individuals for whom it is safe to return to work [47], on April 24, 2020, the World Health Organization (WHO) stipulated that even people who have recovered from COVID-19 and tested positive for antibodies are not necessarily protected from becoming reinfected. One reason for this uncertainty was the low levels of neutralizing antibodies observed in the sera of several recovered COVID-19 patients, which implied that in addition to humoral immunity, cell-mediated immunity might have been crucial to their recovery [48]. Therefore, more consideration should be given to the potential applications of serological testing.
Most HCoV-specific antibody detection assays target IgG and IgM antibodies, but there have been tests targeting immunoglobulin A (IgA) antibodies as well [18, 49,50,51,52,53]. The capture antigens in these assays are either whole virus particles from cultured infected cells, the viral membrane S or E proteins, or the N proteins [43].
Apart from the two label-free surface plasmon resonance (SPR)-based optical biosensors, most serological tests have utilized the conventional label-based enzyme-linked immunosorbent assay (ELISA), western blot (WB), immunofluorescence assay (IFA), lateral flow immunoassay (LFIA), and chemiluminescence immunoassay (CLIA) techniques [16, 17, 54,55,56,57]. In this article, we classify the host antibody detection assays into two types: (1) conventional serological tests and (2) biosensors. We then review the studies and commercial assays of each classification. A summary of the reported studies to date for the detection of host antibodies against CoVs is presented in Table 1.
Conventional serological assays
Traditional serological tests regarding CoV-related disease diagnosis mostly rely on the well-known labeled ELISA, WB, IFA, LFIA, and CLIA methods [53, 55, 57,58,59], which are extensively discussed in the following sections. As shown in Table 1, after the SARS outbreak, between the years 2003 and 2013, more than 20 studies were conducted that incorporated these assays to detect host antibodies against SARS-CoV in human serum or plasma samples. Additionally, in 2005 and 2006, seven assays were reported for identifying antibodies against HCoV-OC43 and -229E. After the emergence of MERS in 2012, three serological tests were developed to identify antibodies against MERS-CoV, and since the ongoing COVID-19 pandemic, 11 studies have been reported regarding anti-SARS-CoV-2 detection.
Enzyme-linked immunosorbent assay (ELISA)
The ELISA technique is a quantitative analytical method that identifies the presence or absence of a specific analyte in a given sample by measuring its concentration via antigen-antibody interactions, which lead to a color change in the presence of an enzyme label and its related substrate [60]. The analyte may be either an antigen or, in the case of serological ELISAs, antibodies [43]. ELISA-based serological tests for CoV-associated diseases have employed horseradish peroxidase (HRP) or alkaline phosphatase (AP) as the enzyme labels, using appropriate substrates [61, 62]. The peroxidase substrates exploited in these assays include 3, 3, 5, 5’-tetramethylbenzidine (TMB), o-phenylenediamine dihydrochloride (OPD), and 2, 2’-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS) [53, 63, 64].
ELISA is a heterogeneous enzyme-based immunoassay that can be classified into four distinct types: (1) direct, (2) indirect, (3) sandwich, and (4) competitive [60]. Serological assays reported for CoV detection have used indirect, sandwich, and competitive ELISA principles [44, 64, 65]. As illustrated in Fig. 1A, in an indirect ELISA, a diluted or undiluted serum sample containing the target antibodies is added to antigen-coated plates, where targets form complexes with the capture antigens. After washing to remove unbound molecules, HRP- or AP-labeled detection antibodies are added that recognize the primary or target antibodies. Following further incubation and washing steps, a substrate is added that is converted by the conjugated enzyme to produce a visible color. The intensity of the color of the reaction product is directly correlated with the concentration of the analyte [66].
In a sandwich ELISA-based serological test, target antibodies are recognized by the capture antigens or antibodies in the first step. Subsequently, the enzyme-labeled detection antigens identify the antigen-antibody or capture-target antibody complexes and bind to their target antibody segments. Following the addition of the enzyme-specific substrate, the concentration of the target corresponds directly to the color intensity [60, 66].
As depicted in Fig. 1B, in a competitive ELISA, a serum sample containing the target antibodies and HRP-labeled analogs of the targets are added simultaneously to antigen-coated plates. The targets and analogs compete for the immobilized capture antigens. After the introduction of the peroxidase substrate, the color intensity is measured, which is inversely proportional to the concentration of the target antibodies [67].
As shown in Table 1, 14, one, and 13 distinct ELISA assays have been reported for detection of antibodies against SARS-CoV, MERS-CoV, and SARS-CoV-2, respectively. The MERS diagnostic assay is a competitive ELISA [64]. The IgM-specific test developed by Xiang et al. for SARS-CoV-2 detection and two of the assays proposed by Zhao et al. for COVID-19 diagnosis involving the simultaneous identification of IgG and IgM antibodies and the individual detection of IgM antibodies are sandwich ELISAs [58, 65]. The other assays are based on the indirect ELISA principle. Except for the serological test developed by Zhao et al. for IgM antibody detection, all of the assays detect IgG antibodies [65]. Nine of these assays were designed to identify both IgG and IgM antibodies, and five detected IgG, IgM, and IgA antibodies. The time required for these assays varied from 1 hour to 4 hours and 10 minutes [50, 53].
In 2004, Guan et al. developed two ELISA platforms for the detection of anti-SARS-CoV antibodies. Their assays detected IgG antibodies against the viral N protein within 1 hour and 15 minutes, with a sensitivity of 70.5-100%. Furthermore, they evaluated the tests using a commercial IFA as the gold standard [68, 69]. Liu and colleagues also designed a 1-hour-and-20-minute ELISA procedure to detect IgG antibodies against the SARS-CoV N protein. The assay was able to detect the target antibodies in 1:1000-diluted sera [70].
In 2004, Woo and coworkers reported an ELISA incorporating N proteins as the capture antigens for the detection of SARS-associated IgG, IgM, and IgA antibodies at serum dilutions of 1:40, 1:10, and 1:10, respectively. The assay had a 3-hour-and-15-minute duration. Also, in 2005, they presented two ELISA-based tests. Both assays detected IgG and IgM antibodies in serum samples, one employing the S protein, and the other, the N protein as the immobilized capture molecule. Under optimal conditions, the S-protein-based test took 3 hours and 15 minutes and detected the antibodies at a serum dilution of 1:20. This assay had a sensitivity of 98.6% and 93.9% for IgG and IgM antibodies, respectively. The N-protein-based ELISA had a sensitivity of 94.7% and 55.2% for the respective antibodies [18, 63]. Furthermore, during that year, Che et al. developed an ELISA platform that exploited N proteins as the antigens and detected IgG antibodies against SARS-CoV, HCoV-OC43, and -229E in serum specimens [71].
In the field of MERS diagnosis, a study was performed by Fukushi and coworkers in 2018. They produced a labeled monoclonal IgG antibody against the S protein of MERS-CoV and used it to establish a competitive ELISA. The sensitivity and specificity of the assay for Ethiopian dromedary camel serum samples were 98% and 100%, respectively. The test took about 2 hours and 30 minutes to complete. This competitive ELISA was able to detect the target antibodies in 1:512-diluted camel sera [64].
Continuous efforts are still in progress to develop highly sensitive serological ELISA platforms for anti-SARS-CoV-2 detection and COVID-19 diagnosis. Amanat et al. proposed a highly sensitive and specific ELISA for the detection of IgG, IgM, and IgA antibodies in patients’ serum or plasma samples. Recombinant S proteins were employed as the capture molecules. The assay took approximately 3 to 4 hours, depending on whether the samples were activated by heat treatment [53]. Lin et al. and Liu et al. designed two different 1-hour-and-45-minute ELISA-based tests to detect IgG and IgM antibodies in 1:100-diluted sera [44, 55]. Both of these assays utilized N proteins as the immobilized capture antigens. A sensitivity of 81.5% was observed for the assay developed by Liu et al.
In another study, Xiang et al. developed two ELISA-based serological tests for the detection of IgG and IgM antibodies in serum samples. Each assay took 1 hour and 45 minutes. The IgG- and IgM-specific tests employed recombinant SARS-CoV-2 antigens and anti-IgM monoclonal antibodies as the capture molecules, respectively. The sensitivity of the IgG and IgM assays were 87.3%, with a specificity of 100% [58]. Zhang et al. reported a 1-hour-and-15-minute ELISA for the detection of IgG and IgM antibodies in 1:80 and 1:800 serum dilutions, respectively. They investigated the use of three prokaryotically expressed N proteins as well as three eukaryotically expressed S proteins as the capture antigens and concluded that the latter proteins were more appropriate for anti-SARS-CoV-2 serology [56].
Western blot (WB)
The WB method, also known as immunoblotting, is a common laboratory technique applied to separate and identify specific proteins among a pool of protein molecules derived from a particular tissue. This method is also used to examine protein expression [72,73,74]. Furthermore, this technique has been employed in developing serological assays to detect antiviral antibodies in patients’ sera, e.g., WB-based serological tests for human immunodeficiency virus (HIV), influenza A and B viruses, and CoVs [57, 75, 76].
A WB assay typically consists of three steps; (1) protein separation by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), (2) transfer of the separated proteins to a nitrocellulose (NC) or polyvinylidene difluoride (PVDF) membrane, and (3) staining and identification of the proteins using an indirect detection assay. In step one, the negatively charged sodium dodecyl sulfate (SDS) renders the proteins as particles with anionic properties. During electrophoresis, depending on their molecular weight and electric charge, the proteins start moving toward the anode through the polyacrylamide gel, which results in their separation (Fig. 2A). As illustrated in Fig. 2B, in the second step, the separated proteins are transferred to an NC or PVDF membrane by exploiting an electric current that moves the anionic proteins toward the anode; hence, the membrane. In step three, as shown in Fig. 2C, the proteins are detected using an indirect assay, which virtually resembles an indirect ELISA. This assay is performed on the membrane that bears the target molecules [74, 77]. In WB-based serological assays for CoVs, anti-CoV antibodies are first captured by CoV-specific N or S proteins and HRP- or AP-labeled detection antibodies recognizing the antigen-antibody complexes are then added. Finally, an appropriate peroxidase or phosphatase substrate is added to produce a measurable signal, such as a light or color change. The peroxidase substrates employed in CoV-specific WB assays include enhanced chemiluminescence (ECL) and 3, 3’-diaminobenzidine tetrahydrochloride (DAB) [50, 78]. 5-Bromo-4-chloro-3-indolyl-phosphate/nitro blue tetrazolium (BCIP/NBT) is an AP substrate that is used in these assays [61, 79, 80].
From 2004 to 2006, eight WB assays for SARS diagnosis were reported; one detected both IgG and IgM antibodies, two detected IgG, IgM, and IgA antibodies, and the other five were specific to IgG. One of these IgG detection assays, which was proposed by Che et al. in 2005, could also detect anti-HCoV-OC43 and -229E antibodies [71]. The time required for these WB assays ranged from 2 to 5 hours [51, 57].
In 2004, He and coworkers reported a WB assay for the detection of IgG and IgM antibodies against SARS-CoV in 1:100-diluted sera. Immunoblotted N proteins on an NC membrane were used as capture antigens. The assay had a sensitivity of 90.9% and a specificity of 98.3% [78]. Guan et al. developed a 2-hour-and-30-minute WB-based serological test that detected IgG antibodies in serum samples to diagnose SARS. Their assay, which had a sensitivity of 100%, utilized an NC platform with N proteins as the capture molecules [80]. Leung and colleagues developed a WB assay for the detection of IgG, IgM, and IgA antibodies against SARS-CoV in 1:200 serum dilutions. The test took 2 hours and used gel-purified N and S proteins on a PVDF membrane as bioreceptors. It was observed that IgG antibodies secreted by SARS patients reacted more strongly with N proteins than with S proteins [50].
A year later, Wang et al. reported a WB-based test for the detection of SARS-specific IgG antibodies in 1:100-diluted serum specimens. This 2-hour assay made use of an NC membrane with immunoblotted N and three truncated S proteins named S1, S2, and S3. Their results indicated that the N-, S1-, S2-, and S3-based WB assays had a sensitivity of 100%, 50%, 30%, and 70%, respectively [79]. In 2006, Maache and coworkers developed a 5-hour WB assay for the detection of IgG antibodies in 1:1500-diluted sera of SARS patients. They employed the N proteins, as well as the S1 and S2 subunits of the S proteins, as the capture antigens on an NC membrane. Similar to the results obtained by Leung et al. and Wang et al., this study also revealed that the sensitivity of the N-protein-based assay was higher than that of the S-protein-based ones and showed that this rather high sensitivity could have been in part due to the fact that N-protein-specific IgG antibodies are generally secreted profusely during the critical and recovery phases of the disease [57].
Immunofluorescence assay (IFA)
The IFA technique is an approved method that detects target antigens or antibodies in a given sample. These assays employ fluorescent dyes, also known as fluorophores, enabling the visualization of the targets using fluorescence microscopy or imaging, or their quantification through measurement of fluorescent light intensity. There are two different types of IFA: direct and indirect. Immunofluorescence serology uses the latter type. As shown in Fig. 3, in indirect IFA-based serological tests, initially, the immobilized antigens recognize and capture the target antibodies present in the serum samples. Subsequently, fluorophore-labeled detection antibodies are introduced, which tag the targets and allow for their visualization and quantification [81,82,83]. IFA-based serological assays for CoV antibodies have utilized fluorescein isothiocyanate (FITC), rhodamine, cyanine (Cy) 3 and 5, DyLight 649, and quantum dot fluorophores as the fluorescent labels [51, 54, 59, 84, 85].
As shown in Table 1, from 2003 to 2020, there were fifteen studies developing IFA-based serological tests as a means to diagnose SARS, MERS, COVID-19, HCoV-OC43, and HCoV-229E. Four of the assays were applicable for the identification of IgG, IgM, and IgA antibodies, three of them detected the IgG and IgM antibodies, and the others were specific to IgG antibodies only. The time required for these assays varied from 6 minutes to almost 19 hours [54, 59].
In 2004, Chan et al. described an IFA for the detection of SARS-CoV-specific IgG antibodies in 1:40 serum dilutions. The assay, which employed whole virions as the immobilized capture agents, took 1 hour and 30 minutes and had a sensitivity and specificity of 100% [86]. In addition to their WB assay, He and coworkers reported an IFA-based test in which they used whole virus particles as the capture antigens. The assay detected IgG and IgM antibodies in 1:10-diluted sera from SARS patients within 30 and 90 minutes, respectively [78]. Moreover, Wu et al. designed an IFA for the detection of SARS-CoV that identified the IgG, IgM, and IgA antibodies in 1:800 serum dilutions within 1 hour and 10 minutes. This assay utilized virus-infected cells as the source of whole virus antigens and had a sensitivity and specificity of 99.1% and 87.8%, respectively [52].
A year later, in addition to their ELISA and WB tests, Che et al. developed an IFA-based serological test for the detection of IgG antibodies against SARS-CoV, HCoV-OC43, and HCoV-229E in human sera. The assay employed whole virus molecules as capture antigens [71]. Manopo and coworkers described an S protein-based IFA for the detection of SARS-specific IgG antibodies in 1:100 serum dilutions. The 1-hour-and-20-minute assay had 100% sensitivity and specificity [87]. In 2006, Zhu et al. reported an IFA-based test for the detection of IgG and IgM antibodies in 1:800-diluted sera from SARS and HCoV-229E patients. The assay made use of N proteins as the capture molecules [84].
In 2013, Reusken et al. developed an IFA assay for the detection of IgG and IgM antibodies against both SARS- and MERS-CoV in human serum samples. The 2-hour assay exploited the S1 subunit’s receptor-binding domains (RBDs) to capture the target antibodies, and detected them in 1:160- and 1:20480-diluted sera for SARS- and MERS-CoV, respectively [85]. Additionally, in 2019, Hoy and colleagues designed an IFA-based serological test that used electrospun polystyrene microfibers as three-dimensional membrane filters and FITC as the fluorescent label for the detection of anti-N protein antibodies against MERS-CoV. This 6-minute assay was able to simultaneously detect anti-human serum albumin antibodies (anti-HSA), as well as the anti-MERS-CoV antibodies. The detection limit for the anti-N protein antibodies in the spiked antibody solutions was 200 μg/ml [59].
A year later, in 2020, during the ongoing COVID-19 pandemic, Khan and coworkers described an IFA-based protein microarray composed of 67 antigens on an NC-coated slide to examine the cross-reactivity between the previous CoVs and the latest one, SARS-CoV-2. This assay examined the seroreactivity of IgG antibodies in 1:100 serum dilutions toward the CoVs. The assay time was roughly 15 to 19 hours, since it involved the overnight incubation of sera. The results demonstrated that despite the high reactivity of the IgG antibodies to the previous CoVs, these antibodies showed low seroreactivity to SARS-CoV-2, with some reactivity to the S2 subunit and the N protein [54].
Lateral flow immunoassay (LFIA)
LFIA, also known as the immunochromatographic test (ICT), is a rapid paper-based assay capable of determining the presence of an analyte in a given sample within a time frame of only 5 to 30 minutes. The applications of LFIA-based tests include food and environment quality assessments as well as medical diagnosis. Test strips have been utilized to analyze samples of whole blood, plasma, serum, urine, sweat, or saliva as body fluids specimens. The assay’s platform is a strip consisting of an inert backing on which four overlapping zones have been assembled using a pressure-sensitive adhesive. These zones are (1) the sample, and (2) the conjugate pads, (3) the NC membrane, and (4) the absorbent pad [88, 89].
As illustrated in Fig. 4, in LFIA-based serological tests, the sample is first dropped onto the sample pad and then channeled into the next zone, i.e., the conjugate pad, through capillary flow. This pad commonly contains tagged capture antigens or anti-antibodies specific to the target molecules. The labels can be colloidal gold, monodisperse latex, colloidal carbon, fluorophores, or quantum dots. For instance, the use of colloidal gold enables the visualization of a red product by the naked eye. After the interaction between the targets and the capture molecules, the resulting complexes flow to the NC membrane, comprising the test and control lines, on which detection antigens and control antigens or antibodies have been immobilized, respectively. Lastly, the absorbent or wick pad absorbs the processed sample through the capillary effect, thus preventing it from flowing back into the NC membrane and the conjugate pad. Generally, the test result is positive when both of these lines adopt a color visible by the naked eye or observable using a fluorescence analyzer. If the color is seen only on the control line, it signifies a negative result, and if the color appears only on the test line, it indicates that the test is inaccurate and was not appropriately implemented [88,89,90].
In 2004, in addition to their ELISA-based tests discussed in section 2-1, Guan et al. developed two LFIAs that detected the IgG antibodies against SARS-CoV in undiluted sera within 15 minutes. Both assays employed colloidal gold-tagged anti-IgG antibodies, N proteins, and proteins unique to SARS-CoV as the capture, detection, and control agents, respectively (Fig. 4A). A commercial IFA was used to validate these tests, which were found to have a sensitivity and specificity of 70.5-100% and 97.7-100%, respectively [68, 69]. Wu et al. described an LFIA for the detection of anti-N-protein antibodies in the sera of SARS patients. The assay’s conjugate pad contained colloidal gold-conjugated N proteins and mouse IgG antibodies. N proteins and anti-mouse IgG antibodies were immobilized onto the test and control lines, respectively. The assay had a sensitivity of 33.6% and a specificity of 98.2% [52].
In 2020, as a means to identify COVID-19 patients, Xiang et al. reported an ICT that was employed to detect the IgG and IgM antibodies associated with SARS-CoV-2 in the patients’ plasma specimens. The 10-minute assay had a sensitivity and specificity of 82.4% and 100%, respectively [58]. Zhang et al. designed an LFIA for the detection of IgG and IgM antibodies in 1:160-diluted sera of COVID-19 patients within 5 to 10 minutes. This test made use of colloidal gold-labeled RBDs of recombinant S proteins, recombinant S1 subunits, and secondary polyclonal antibodies as the capture, detection, and control probes, respectively (Fig. 4B). The sensitivity of the assay was 86.89%, and its specificity was 99.39% [56].
Chemiluminescence immunoassay (CLIA)
CLIA is a rapid and ultrasensitive immunoassay that is widely applied for the detection of biological and environmental molecules, such as proteins, nucleic acids, hormones, and pollutants. This label-based technique employs either chemiluminescent tags such as luminol and isoluminol or enzymatic labels such as HRP or AP, followed by the addition of a luminol-based substrate that generates a light signal, the intensity of which can be measured using a chemiluminescent analyzer [91]. Fig. 5 illustrates a CLIA-based serological test utilizing enzymatic tags with their specific chemiluminescent substrates.
In addition to their ELISA-based serological test, Lin and coworkers developed a CLIA using magnetic bead-conjugated recombinant N proteins, AP-labeled anti-immunoglobulin antibodies, and Lumigen APS-5 as the capture agents, detection probes, and the AP-specific chemiluminescent substrate, respectively. This assay detected IgG and IgM antibodies against SARS-CoV-2 in undiluted serum samples within 23 minutes. The test had a sensitivity of 82.28% for both antibody types, and a specificity of 97.5% and 81.25% for IgG and IgM antibodies, respectively [55].
Cai and colleagues designed a magnetic CLIA to detect IgG and IgM antibodies in the sera of COVID-19 patients. This assay, which took approximately 30 minutes, was based on magnetic-bead-bound peptides derived from the S, N, or open reading frame 1a and 1b (ORF1a/b) proteins specific to SARS-CoV-2. Their results indicated that the CLIA employing the S proteins as the capture antigens demonstrated the best performance, with a positive detection rate of 71.4% and 57.2% for IgG and IgM antibodies, respectively [92]. Qu et al. described a CLIA-based serological test that identified IgG and IgM antibodies against SARS-CoV-2 in serum specimens. They employed S and N proteins to heighten the assay’s sensitivity. The assay had a specificity of 95% for both types of antibodies and a sensitivity of 90% and 80% for IgG and IgM antibodies, respectively [93].
Biosensors
Biosensors offer features such as rapid analysis, high sensitivity and specificity, and cost-effectiveness. Therefore, they are promising candidates for the detection of pathogenic particles. To date, a variety of electrochemical, electrical, optical, piezoelectric, and mechanical biosensors have been reported as media for virus detection [94,95,96].
Regarding SARS-CoV-specific serology, two label-free SPR-based optical biosensors were reported by Park et al. in 2009 and 2011. In the first study, they developed an SPR-based assay for the rapid detection of anti-SARS-CoV E protein antibodies. They fabricated a gold micro-patterned chip and immobilized the E proteins on the gold surface through gold binding polypeptides. After SPR imaging analysis, the biosensor identified as little as 200 ng of the target antibodies per ml within 10 minutes in spiked antibody solutions (Fig. 6A). In their other work, they presented an SPR-like reflective biosensor for anti-E protein antibody detection. They exploited functionalized colloidal silica photonic crystals and the SPR-like characteristics of their reflectance spectra and detected SARS-CoV-associated antibodies within an hour (Fig. 6B) [16, 17].
Schematic illustrations of two optical biosensors for SARS-CoV-specific antibody detection. (A) The first one is an SPR-based biosensor employing E proteins immobilized on a gold surface as the capture probes. (B) The second biosensor is an SPR-like reflective platform utilizing the same proteins attached to a colloidal silica photonic crystal-coated chip
Commercial serological assays for COVID-19 diagnosis
Since the COVID-19 pandemic, a multitude of serology-based tests have been under development, and several have reached the commercialization status [102]. Table 2 lists various commercialized assays that have received United States Food and Drug Association (FDA) Emergency Use Authorization (EUA) or European CE approval. These assays are ELISA-, IFA-, LFIA, or CLIA-based. As shown in Table 2, the ELISA-, LFIA-, and CLIA-based tests have comparatively high sensitivity and specificity. Due to the urgent demand for COVID-19 screening worldwide, LFIA-based serological testing seems to be favored due to its high sensitivity and specificity, rapid response, and easy application. It is less operator-dependent than other assays, which even if automated, still require sophisticated equipment. Moreover, because COVID-19 is often asymptomatic and the level of IgM antibodies tends to decline within 14 days of the disease onset, a rapid diagnostic assay, e.g., an LFIA, detecting IgG and IgM antibodies simultaneously is recommended.
Concluding remarks and future perspectives
The conventional serological assays, including ELISA, WB, IFA, LFIA, and CLIA, as well as biosensors as platforms for the detection of antibodies against HCoVs in human whole blood, serum, or plasma samples, have been discussed in this review. During the SARS, MERS, and COVID-19 outbreaks, these tests have been mostly applied alongside NAATs as rapid and inexpensive alternatives for large-scale screening. Despite the statement of WHO on April 24, 2020, regarding the most recently identified HCoV, SARS-CoV-2, these assays have been applied for analyzing the transmission of CoVs and identifying newly infected and recovered patients who have potentially become immune to the disease. The sensitivity of serological assays varies among patients and is profoundly dependent on the time of the infection; since the amount of anti-CoV antibodies secreted generally depends on the time since viral exposure.
In conclusion, further research is required to understand the effect of the host antibodies on SARS-CoV-2; in order to determine whether these assays can be used to identify individuals who have become immune to the disease. Nonetheless, their rapid response and low cost have made serological tests attractive candidates for screening large numbers of people with suspected infection, in contrast to the time-consuming and costly NAATs, such as RT-PCR assays. Among the conventional ELISA-, IFA-, LFIA-, and CLIA-based serological tests, an LFIA identifying both IgG and IgM antibodies seems to be a desirable choice due to its relatively high sensitivity and specificity and rapid response, which satisfies the urgent need for COVID-19 screening and a simple testing procedure.
References
Naji H (2020) The emerging of the 2019 novel coronavirus 2019-nCoV. Eur J Med Health Sci. https://doi.org/10.24018/ejmed.2020.2.1.169
Chan JFW, Li KSM, To KKW et al (2012) Is the discovery of the novel human betacoronavirus 2c EMC/2012 (HCoV-EMC) the beginning of another SARS-like pandemic? J Infect 65:477–489. https://doi.org/10.1016/j.jinf.2012.10.002
Lim YX, Ng YL, Tam JP, Liu DX (2016) Human coronaviruses: a review of virus–host interactions. Diseases. https://doi.org/10.3390/diseases4030026
Jones BA, Grace D, Kock R et al (2013) Zoonosis emergence linked to agricultural intensification and environmental change. Proc Natl Acad Sci 110:8399–8404. https://doi.org/10.1073/pnas.1208059110
Kahn JS, McIntosh K (2005) History and recent advances in coronavirus discovery. Pediatr Infect Dis J 24:S223–S227. https://doi.org/10.1097/01.inf.0000188166.17324.60
Fehr AR, Perlman S (2015). Coronaviruses: an overview of their replication and pathogenesis. In Coronaviruses. Methods in molecular biology. Humana Press, New York, pp 1–23
Sun C, Chen L, Yang J et al (2020) SARS-CoV-2 and SARS-CoV spike-RBD structure and receptor binding comparison and potential implications on neutralizing antibody and vaccine development. bioRxrv. https://doi.org/10.1101/2020.02.16.951723
Schmaljohn AL (2013) Protective antiviral antibodies that lack neutralizing activity: precedents and evolution of concepts. Curr HIV Res 11:345–353. https://doi.org/10.2174/1570162x113116660057
Bosch BJ, De HCAM, Rottier PJM (2004) Coronavirus spike glycoprotein, extended at the carboxy terminus with green fluorescent protein, is assembly competent. J Virol 78:7369–7378. https://doi.org/10.1128/JVI.78.14.7369
Fan H, Ooi A, Tan YW et al (2005) The nucleocapsid protein of coronavirus infectious bronchitis virus: crystal structure of its N-terminal domain and multimerization properties. Structure 13:1859–1868. https://doi.org/10.1016/j.str.2005.08.021
Jiang S, Hillyer C, Du L (2020) Neutralizing antibodies against SARS-CoV-2 and other human coronaviruses. Trends Immunol 41:355–359. https://doi.org/10.1016/j.it.2020.03.007
di Gabriella M, Cristina S, Concetta R et al (2020) SARS-Cov-2 infection: response of human immune system and possible implications for the rapid test and treatment. Int Immunopharmacol 84:106519. https://doi.org/10.1016/j.intimp.2020.106519
Lin Q, Zhu L, Ni Z et al (2020) Duration of serum neutralizing antibodies for SARS-CoV-2: lessons from SARS-CoV infection. J Microbiol Immunol Infect. https://doi.org/10.1016/j.jmii.2020.03.015
Vashist SK (2020) In vitro diagnostic assays for COVID-19: recent advances and emerging trends. Diagnostics 10:202. https://doi.org/10.3390/diagnostics10040202
Zhou P, Lou YX, Wang XG et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Park TJ, Hyun MS, Lee HJ et al (2009) A self-assembled fusion protein-based surface plasmon resonance biosensor for rapid diagnosis of severe acute respiratory syndrome. Talanta 79:295–301. https://doi.org/10.1016/j.talanta.2009.03.051
Park TJ, Lee S-K, Yoo SM et al (2011) Development of reflective biosensor using fabrication of functionalized photonic nanocrystals. J Nanosci Nanotechnol 11:632–637. https://doi.org/10.1166/jnn.2011.3269
Woo PCY, Lau SKP, Wong BHL et al (2004) Longitudinal profile of immunoglobulin G (IgG), IgM, and IgA antibodies against the severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein in patients with pneumonia due to the SARS coronavirus. Clin Diagn Lab Immunol 11:665–668. https://doi.org/10.1128/CDLI.11.4.665
Siracusano G, Pastori C, Lopalco L (2020) Humoral immune responses in COVID-19 patients: a window on the state of the art. Front Immunol 11:1049. https://doi.org/10.3389/fimmu.2020.01049
Zhang N, Wang L, Deng X et al (2020) Recent advances in the detection of respiratory virus infection in humans. J Med Virol 92:408–417. https://doi.org/10.1002/jmv.25674
van der Hoek L (2007) Human coronaviruses: what do they cause? Antivir Ther 12:651–658
Carlos WG, Dela Cruz CS, Cao B et al (2020) Novel Wuhan (2019-nCoV) coronavirus. Am J Respir Crit Care Med 201:P7–P8. https://doi.org/10.1164/rccm.2014P7
Lam CWK, Chan MHM, Wong CK (2004) Severe acute respiratory syndrome: clinical and laboratory manifestations. Clin Biochem Rev 25:121–132
Wong TW (2006) ‘“Will the SARS epidemic recur?”’ A retrospective analysis of the experts’ opinions. J Epidemiol Community Heal 60:87
Rabaan AA (2017) Middle East respiratory syndrome coronavirus: five years later. Expert Rev Respir Med 11:901–912. https://doi.org/10.1080/17476348.2017.1367288
Hu B, Ge X, Wang L-F, Shi Z (2015) Bat origin of human coronaviruses. Virol J 12:221. https://doi.org/10.1186/s12985-015-0422-1
World Health Organization (2020) MERS situation update, January 2020. https://www.emro.who.int/pandemic-epidemic-diseases/mers-cov/mers-situation-update-january-2020.html
Chan JF-W, Kok K-H, Zhu Z et al (2020) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect 9:221–236. https://doi.org/10.1080/22221751.2020.1719902
Lu H, Stratton CW, Wei Y (2020) The Wuhan SARS-CoV-2—What’s next for China. J Med Virol 92:546–547. https://doi.org/10.1002/jmv.25738
Kwong KCNK, Mehta PR, Shukla G, Mehta AR (2020) COVID-19, SARS and MERS: a neurological perspective. J Clin Neurosci. https://doi.org/10.1016/j.jocn.2020.04.124
Worldometers (2020) COVID-19 Coronavirus Pandemic. https://www.worldometers.info/coronavirus/
World Health Organization (2020) WHO Coronavirus Disease (COVID-19) Dashboard. https://covid19.who.int/
Zheng J (2020) SARS-CoV-2: an emerging coronavirus that causes a global threat. Int J Biol Sci 16:1678–1685. https://doi.org/10.7150/ijbs.45053
Lu R, Zhao X, Li J et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395:565–574. https://doi.org/10.1016/S0140-6736(20)30251-8
Wong HYF, Lam HYS, Fong AH-T et al (2020) Frequency and distribution of chest radiographic findings in COVID-19 positive patients. Radiology. https://doi.org/10.1148/radiol.2020201160
Kim S-H, Ko J-H, Park GE et al (2017) Atypical presentations of MERS-CoV infection in immunocompromised hosts. J Infect Chemother 23:769–773. https://doi.org/10.1016/j.jiac.2017.04.004
Zeng Q, Chen L, Cai X et al (2003) Chest X-ray and CT in the diagnosis of SARS. Chin J Radiol 37:600–603
Zhu X, Wang X, Han L et al (2020) Reverse transcription loop-mediated isothermal amplification combined with nanoparticles-based biosensor for diagnosis of COVID-19. medRxiv. https://doi.org/10.1101/2020.03.17.20037796
Hirotsu Y, Mochizuki H, Omata M (2020) Double-Quencher Probes improved the detection sensitivity of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) by one-step RT-PCR. medRxiv. https://doi.org/10.1101/2020.03.17.20037903
Qiu G, Gai Z, Tao Y et al (2020) Dual-functional plasmonic photothermal biosensors for highly accurate severe acute respiratory syndrome coronavirus 2 detection. ACS Nano. https://doi.org/10.1021/acsnano.0c02439
Diao B, Wen K, Chen J et al (2020) Diagnosis of acute respiratory syndrome coronavirus 2 infection by detection of nucleocapsid protein. medRxiv. https://doi.org/10.1101/2020.03.07.20032524
Layqah LA, Eissa S (2019) An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Microchim Acta 186:224. https://doi.org/10.1007/s00604-019-3345-5
Meyer B, Drosten C, Müller MA (2014) Serological assays for emerging coronaviruses: challenges and pitfalls. Virus Res 194:175–183. https://doi.org/10.1016/j.virusres.2014.03.018
Liu L, Liu W, Zheng Y et al (2020) A preliminary study on serological assay for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in 238 admitted hospital patients. medRxiv. https://doi.org/10.1101/2020.03.06.20031856
Cheng MS, Toh C-S (2013) Novel biosensing methodologies for ultrasensitive detection of viruses. Analyst 138:6219–6229. https://doi.org/10.1039/C3AN01394D
Huang X, Wei F, Hu L et al (2020) Epidemiology and clinical characteristics of COVID-19. Arch Iran Med 23:268–271. https://doi.org/10.34172/aim.2020.09
Abbasi J (2020) The promise and peril of antibody testing for COVID-19. JAMA. https://doi.org/10.1001/jama.2020.6170
World Health Organization (2020) “Immunity passports” in the context of COVID-19. https://www.who.int/news-room/commentaries/detail/immunity-passports-in-the-context-of-covid-19
Ksiazek TG, Erdman D, Goldsmith CS et al (2003) A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med 348:1953–1966. https://doi.org/10.1056/NEJMoa030781
Leung DTM, Tam FCH, Ma CH et al (2004) Antibody response of patients with severe acute respiratory syndrome (SARS) targets the viral nucleocapsid. J Infect Dis 190:379–386. https://doi.org/10.1086/422040
Tan Y-J, Goh P-Y, Fielding BC et al (2004) Profiles of antibody responses against severe acute respiratory syndrome coronavirus recombinant proteins and their potential use as diagnostic markers. Clin Diagn Lab Immunol 11:362–371. https://doi.org/10.1128/CDLI.11.2.362
Wu H-S, Chiu S-C, Tseng T-C et al (2004) Serologic and molecular biologic methods for SARS-associated. Emerg Infect Dis 10:304–310. https://doi.org/10.3201/eid1002.030731
Amanat F, Stadlbauer D, Strohmeier S et al (2020) A serological assay to detect SARS-CoV-2 seroconversion in humans. medRxiv. https://doi.org/10.1101/2020.03.17.20037713
Khan S, Nakajima R, Jain A et al (2020) Analysis of serologic cross-reactivity between common human coronaviruses and SARS-CoV-2 using coronavirus antigen microarray. bioRxiv. https://doi.org/10.1101/2020.03.24.006544
Lin D, Liu L, Zhang M et al (2020) Evaluations of serological test in the diagnosis of 2019 novel coronavirus (SARS-CoV-2) infections during the COVID-19 outbreak. medRxiv. https://doi.org/10.1101/2020.03.27.20045153
Zhang P, Gao Q, Wang T et al (2020) Evaluation of recombinant nucleocapsid and spike proteins for serological diagnosis of novel coronavirus disease 2019 (COVID-19). medRxiv. https://doi.org/10.1101/2020.03.17.20036954
Maache M, Komurian-Pradel F, Rajoharison A et al (2006) False-positive results in a recombinant severe acute respiratory western blot assay were rectified by the use of two subunits (S1 and S2) of spike for detection of antibody to SARS-CoV. Clin Diagn Lab Immunol 13:409–414. https://doi.org/10.1128/CVI.13.3.409
Xiang J, Yan M, Li H et al (2020) Evaluation of enzyme-linked immunoassay and colloidal gold-immunochromatographic assay kit for detection of novel coronavirus (SARS-Cov-2) causing an outbreak of pneumonia (COVID-19). medRxiv. https://doi.org/10.1101/2020.02.27.20028787
Hoy CFO, Kushiro K, Yamaoka Y et al (2019) Rapid multiplex microfiber-based immunoassay for anti-MERS-CoV antibody detection. Sens Bio Sens Res. https://doi.org/10.1016/j.sbsr.2019.100304
Aydin S (2015) A short history, principles, and types of ELISA, and our laboratory experience with peptide/protein analyses using ELISA. Peptides 72:4–15. https://doi.org/10.1016/j.peptides.2015.04.012
Wang Y-D, Li Y, Xu G-B et al (2004) Detection of antibodies against SARS-CoV in serum from SARS-infected donors with ELISA and Western blot. Clin Immunol 113:145–150. https://doi.org/10.1016/j.clim.2004.07.003
Carattoli A, Di BP, Grasso F et al (2005) Recombinant protein-based ELISA and immuno-cytochemical assay for the diagnosis of SARS. J Med Virol 76:137–142. https://doi.org/10.1002/jmv.20338
Woo PCY, Lau SKP, Wong BHL et al (2005) Differential sensitivities of severe acute respiratory syndrome (SARS) coronavirus spike polypeptide enzyme-linked immunosorbent assay (ELISA) and SARS coronavirus nucleocapsid protein ELISA for serodiagnosis of SARS coronavirus pneumonia. J Clin Microbiol 43:3054–3058. https://doi.org/10.1128/JCM.43.7.3054
Fukushi S, Fukuma A, Kurosu T et al (2018) Characterization of novel monoclonal antibodies against the MERS-coronavirus spike protein and their application in species-independent antibody detection by competitive ELISA. J Virol Methods 251:22–29. https://doi.org/10.1016/j.jviromet.2017.10.008
Zhao J, Yuan Q, Wang H et al (2020) Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa344
González-Martínez MÁ, Puchades R, Maquieira Á (2018) Immunoanalytical technique: enzyme-linked immunosorbent assay (ELISA). Modern techniques for food authentication. Academic Press, New York, pp 617–657
García-González E, Aramendía M, Álvarez-Ballano D et al (2016) Serum sample containing endogenous antibodies interfering with multiple hormone immunoassays. Laboratory strategies to detect interference. Pract Lab Med 4:1–10. https://doi.org/10.1016/j.plabm.2015.11.001
Guan M, Chan KH, Peiris JSM et al (2004) Evaluation and validation of an enzyme-linked immunosorbent assay and an immunochromatographic test for serological diagnosis of severe acute respiratory syndrome. Clin Diagn Lab Immunol 11:699–703. https://doi.org/10.1128/CDLI.11.4.699
Guan M, Chen HY, Foo SY et al (2004) Recombinant protein-based enzyme-linked immunosorbent assay and immunochromatographic tests for detection of immunoglobulin G antibodies to Severe Acute Respiratory Syndrome (SARS) coronavirus in SARS patients. Clin Diagn Lab Immunol 11:287–291. https://doi.org/10.1128/CDLI.11.2.287
Liu X, Shi Y, Li P et al (2004) Profile of antibodies to the nucleocapsid protein of the severe acute respiratory syndrome (SARS)-associated coronavirus in probable SARS patients. Clin Diagn Lab Immunol 11:227–228. https://doi.org/10.1128/CDLI.11.1.227
Che X-Y, Qiu L-W, Liao Z-Y et al (2005) Antigenic cross-reactivity between severe acute respiratory syndrome-associated coronavirus and human coronaviruses 229E and OC43. J Infect Dis 191:2033–2037. https://doi.org/10.1086/430355
Smith-Norowitz TA, Kusonruksa M, Wong D et al (2012) Long-term persistence of IgE anti-influenza A HIN1 virus antibodies in serum of children and adults following influenza A vaccination with subsequent H1N1 infection: a case study. J Inflamm Res 5:111–116. https://doi.org/10.2147/JIR.S34152
Martins-Gomes C, Silva AM (2018). Western blot methodologies for analysis of in vitro protein expression induced by teratogenic agents. Teratogenicity testing. Methods in molecular biology. Humana Press, New York, pp 191–203
Mahmood T, Yang P-C (2012) Western blot: technique, theory, and trouble shooting. N Am J Med Sci 4:429–434. https://doi.org/10.4103/1947-2714.100998
Qiu D, Tannock GA, Barry RD, Jackson DC (1992) Western blot analysis of antibody responses to influenza virion proteins. Immunol Cell Biol 70:181–191. https://doi.org/10.1038/icb.1992.23
Castejon MJ, Yamashiro R, Oliveira CAF, Veras MASM (2017) Performance validation of western blot for anti-HIV antibody detection in blood samples collected on filter paper (DBS). J Bras Patol e Med Lab 53:5–12. https://doi.org/10.5935/1676-2444.20170002
Kurien BT, Scofi RH (2015) Western blotting: an introduction. Methods in molecular biology. Humana Press, New York, pp 17–30
He Q, Chong KH, Chng HH et al (2004) Development of a western blot assay for detection of antibodies against coronavirus causing severe acute respiratory syndrome. Clin Diagn Lab Immunol 11:417–422. https://doi.org/10.1128/CDLI.11.2.417
Wang Y, Chang Z, Ouyang J et al (2005) Profiles of IgG antibodies to nucleocapsid and spike proteins of the SARS-associated coronavirus in SARS patients. DNA Cell Biol 24:521–527. https://doi.org/10.1089/dna.2005.24.521
Guan M, Chen HY, Tan PH et al (2004) Use of viral lysate antigen combined with recombinant protein in western immunoblot assay as confirmatory test for serodiagnosis of severe acute respiratory syndrome. Clin Diagn Lab Immunol 11:1148–1153. https://doi.org/10.1128/CDLI.11.6.1148
Odell ID, Cook D (2013) Immunofluorescence techniques. J Investig Dermatol 133:e4. https://doi.org/10.1038/jid.2012.455
Dilnessa T, Zeleke H (2017) Cell culture, cytopathic effect and immunofluorescence diagnosis of viral infection. J Microbiol Mod Technol 2:102
Bossuyt X, Cooreman S, De Baere H et al (2013) Detection of antinuclear antibodies by automated indirect immunofluorescence analysis. Clin Chim Acta 415:101–106. https://doi.org/10.1016/j.cca.2012.09.021
Zhu H, Hu S, Jona G et al (2006) Severe acute respiratory syndrome diagnostics using a coronavirus protein microarray. Proc Natl Acad Sci 103:4011–4016. https://doi.org/10.1073/pnas.0510921103
Reusken C, Mou H, Godeke GJ et al (2013) Specific serology for emerging human coronaviruses by protein microarray. Eurosurveillance 18:20441. https://doi.org/10.2807/1560-7917.ES2013.18.14.20441
Chan PKS, Ng K-C, Chan RCW et al (2004) Immunofluorescence assay for serologic diagnosis of SARS. Emerg Infect Dis 10:530–532. https://doi.org/10.3201/eid1003.030493
Manopo I, Lu L, He Q et al (2005) Evaluation of a safe and sensitive Spike protein-based immunofluorescence assay for the detection of antibody responses to SARS-CoV. J Immunol Methods 296:37–44. https://doi.org/10.1016/j.jim.2004.10.012
Koczula KM, Gallotta A (2016) Lateral flow assays. Essays Biochem 60:111–120. https://doi.org/10.1042/EBC20150012
Wong RC, Tse HY (2008) Lateral flow immunoassay. Humana Press, New York
Bahadır EB, Sezgintürk MK (2016) Lateral flow assays: principles, designs and labels. Trends Anal Chem 82:286–306. https://doi.org/10.1016/j.trac.2016.06.006
Zhao L, Sun L, Chu X (2009) Chemiluminescence immunoassay. Trends Anal Chem 28:404–415. https://doi.org/10.1016/j.trac.2008.12.006
Cai X-F, Chen J, Hu J-L et al (2020) A peptide-based magnetic chemiluminescence enzyme immunoassay for serological diagnosis of corona virus disease 2019 (COVID-19). J Infect Dis. https://doi.org/10.1101/2020.02.22.20026617
Qu J, Wu C, Li X et al (2020) Profile of IgG and IgM antibodies against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Clin Infect Dis. https://doi.org/10.1093/cid/ciaa489
Turner APF (2013) Biosensors: sense and sensibility. Chem Soc Rev 42:3184–3196. https://doi.org/10.1039/c3cs35528d
Caygill RL, Blair GE, Millner PA (2010) A review on viral biosensors to detect human pathogens. Anal Chim Acta 681:8–15. https://doi.org/10.1016/j.aca.2010.09.038
Lakshmipriya T, Gopinath SCB (2019) An introduction to biosensors and biomolecules. In Nanobiosensors for biomolecular targeting. Elsevier, New York, pp 1–21
Hsueh P-R, Hsiao C-H, Yeh S-H et al (2003) Microbiologic characteristics, serologic responses, and clinical manifestations in severe acute respiratory syndrome, Taiwan. Emerg Infect Dis 9:1163–1167. https://doi.org/10.3201/eid0909.030367
Peiris JSM, Lai ST, Poon LLM et al (2003) Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 361:1319–1325. https://doi.org/10.1016/S0140-6736(03)13077-2
Shi Y, Yi Y, Li P et al (2003) Diagnosis of severe acute respiratory syndrome (SARS) by detection of SARS coronavirus nucleocapsid antibodies in an antigen-capturing enzyme-linked immunosorbent assay. J Clin Microbiol 41:5781–5782. https://doi.org/10.1128/JCM.41.12.5781
Okba NMA, Müller MA, Li W et al (2020) SARS-CoV-2 specific antibody responses in COVID-19 patients. Emerg Infect Dis. https://doi.org/10.1101/2020.03.18.20038059
Perera RAPM, Mok CKP, Tsang OTY et al (2020) Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Eurosurveillance 25:2000421. https://doi.org/10.2807/1560-7917
Find; Because Diagnosis Matters (2020) SARS-CoV-2 Diagnostic Pipeline. https://www.finddx.org/covid-19/pipeline/?section=immunoassays#diag_tab
EUROIMMUN AG (2020) Anti-SARS-CoV-2 ELISA (IgG) Instruction for use
InBios International Inc. (2020) InBios SCoV-2 DetectTM IgG ELISA Instructions for Use
Beijing Wantai Biological Pharmacy Enterprise Co. Ltd. (2020) WANTAI SARS-CoV-2 Ab ELISA
Bio-Rad Laboratories Inc. (2020) Platelia SARS-CoV-2 Total Ab
Shanghai Fosun Long March Medical Science Co. (2020) Serology Test Evaluation Report for “Fosun COVID-19 IgG/IgM Rapid Antibody Detection Kit” from Shanghai Fosun Long March Medical Science Co., Ltd.
Shanghai Fosun Long March Medical Science Co. (2020) Fosun Novel Coronavirus (2019-nCoV) IgM/IgG Test Kit. https://en.lm-diagnostics.com.cn/prod_view.aspx?nid=3&typeid=89&id=576
Nuccetelli M, Pieri M, Grelli S et al (2020) SARS-CoV-2 infection serology: a useful tool to overcome lockdown? Cell Death Discov 6:38. https://doi.org/10.1038/s41420-020-0275-2
Cellex Inc. (2020) Cellex qSARS-CoV-2 IgG/IgM Rapid Test. https://www.fda.gov/media/136625/download
Healgen Scientiic LLC. (2020) COVID-19 IgG/IgM Rapid Test Cassette (Whole Blood/Serum/Plasma) instruction for use. https://www.fda.gov/media/136625/download
Biohit Healthcare (Hefei) Co. (2020) Biohit SARS-CoV-2 IgM/IgG antibody test kit instruction for use. https://www.fda.gov/media/139283/download
Hangzhou Laihe Biotech Co. (2020) LYHER novel coronavirus (2019-nCoV) IgM/IgG antibody combo test kit (Colloidal Gold) instruction for use. https://www.fda.gov/media/139410/download
Hangzhou Biotest Biotech Co. (2020) RightSignTM COVID-19 IgG/IgM Rapid Test Cassette. https://www.fda.gov/media/138660/download
DiaSorin Inc. (2020) LIAISON® SARS-CoV-2 S1/S2 IgG. https://www.fda.gov/media/138660/download
Abbott Laboratories (2020) SARS-CoV-2 IgG for use with ARCHITECT. https://www.fda.gov/media/137383/download
SIEMENS Healthineers (2020) SARS-CoV-2 total (COV2T) assay for the detection of total antibodies to SARS-CoV-2. https://www.fda.gov/media/138442/download
Ortho Clinical Diagnostics Inc. (2020) Instructions for use VITROS immunodiagnostic products anti-SARS-CoV-2 total reagent pack and calibrator. https://www.fda.gov/media/136967/download
Roche Diagnostics (2020) Elecsys Anti-SARS-CoV-2. https://www.fda.gov/media/137605/download
Egger M, Bundschuh C, Wiesinger K et al (2020) Comparison of the Elecsys® anti-SARS-CoV-2 immunoassay with the EDITM enzyme linked immunosorbent assays for the detection of SARS-CoV-2 antibodies in human plasma. Clin Chim Acta 509:18–21. https://doi.org/10.1016/j.cca.2020.05.049
Shabani E, Dowlatshahi S, Abdekhodaie MJ (2020) Laboratory detection methods for the human coronaviruses. Eur J Clin Microbiol Infect Dis. https://doi.org/10.1007/s10096-020-04001-8
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing interests or personal relationships that could have influenced the work reported in this article.
Additional information
Handling Editor: Sheela Ramamoorthy.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dowlatshahi, S., Shabani, E. & Abdekhodaie, M.J. Serological assays and host antibody detection in coronavirus-related disease diagnosis. Arch Virol 166, 715–731 (2021). https://doi.org/10.1007/s00705-020-04874-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04874-2