Abstract
Fowl adenoviruses (FAdVs) are the causative agents of multietiological syndromes and diseases in poultry flocks. During a routine diagnostic examination, two FAdVs strains were isolated. Molecular typing of these isolates based on the partial loop L1 HVR1-4 region of the hexon gene sequence revealed the presence of different FAdV isolates: 1/A-61/11z (GenBank accession number KX247012, APP94082), and 8a/E-6/12j (GenBank accession number: KP890032, ALB00550), and comparative genome analysis indicated small differences between these two viruses. The next step of the study was the estimation of the pathogenicity of these isolates in specific-pathogen-free (SPF) chickens. Chickens were divided into three groups, with 20 chickens per group infected intraperitoneally on the first day after hatching. Group I consisted of chickens infected with strain FAdV-1/A-61/11z, group II consisted of chickens infected with strain FAdV-8a/E-6/12j, and group III consisted of uninfected birds. Clinical signs observed in infected chickens included poor growth, apathy, prostration, ruffled feathers, crouching position, and huddling behavior. The mortality rate in chickens infected with FAdV-1/A-61/11z was 10% at 10 days postinfection (dpi), and no mortality was observed in chickens infected with the FAdV-8a-6/12j strain. The mean real-time PCR threshold cycle (Ct) value was 39.70%. The detection limit of these assays was 8 copies, with an efficiency of 91.03% and 95.17% and regression square (R2) values of 0.991 and 0.997, respectively, with a mean pathogen load of 4.8 × 106.0 copies/µl. The assays did not demonstrate cross-reactivity between types 1/A and 8a/E and non-targeted poultry viruses. Adenoviral DNA was detected in the liver, spleen, kidney, gizzard, intestine, bursa of Fabricius, and thymus of every examined dead and euthanized chicken from groups I and II between the third and fourth week postinfection. This is the first study conducted on the pathogenic and apathogenic strains FAdV-1/A and FAdV-8a/E, showing the presence of the virus in multiple tissues in chickens in Poland. This study revealed that it is very likely that the FAdV-1/A-61/11z strain is able to cause clinical inclusion body hepatitis (IBH) in chickens and that it is slightly more virulent than the FAdV-8a/E-6/12j strain, although both are primary pathogens of the disease.
Similar content being viewed by others
Introduction
Fowl adenoviruses (FAdVs) are widely distributed in poultry populations around the world [12, 24, 25] and have been isolated from sick birds as well as from birds without any clinical signs of infection [4, 6, 24, 25]. In recent years, the number of adenovirus infections associated with inclusion body hepatitis (IBH) and gizzard erosion and ulceration (GEU) in poultry flocks in Poland and other European countries has increased. Most cases have been observed in broilers between 3 to 5 weeks of age. As a result of these findings, a study on two FAdVs strains isolated from diseased chickens in Poland, including phylogenetic and pathogenic analysis, was undertaken. FAdVs may cause infection independently, or they may be one of the factors involved in various multi-etiological syndromes [6]. At present, the pathogenic role of adenoviruses is changing, and some FAdVs types (2, 11/D, 8a, and 8b/E) may be responsible for clinical manifestations of IBH [5, 25, 27, 29, 37, 40, 41]. Strains belonging to type/species FAdV-1/A are responsible mainly for adenoviral gizzard erosion (GEU) [8, 10, 15,16,17, 26, 31, 32]. FAdV-4/C strains are responsible for hepatitis-hydropericardium syndrome (HHS), mainly in Asia, some Arab countries and Latin America [7]. Adenoviruses of types 7/E and 1/A are responsible for failure of chicken vaccination and immunodeficiency [28, 35]. The presence of FAdV DNA in infected birds may be detected in many internal organs, including trachea, kidney, liver, spleen, intestines, gizzard, and bursa of Fabricius [1]. Virus tropism depends on the age of the bird and immunological competence of the strain. Different strains have different levels of pathogenicity and virulence, which may determine the outcome of the infection as well as cell tropism [3, 6, 11, 30]. Adenoviruses have an affinity for certain receptors [18, 24, 33].
The pathogenic properties of FAdVs differ among the various types, which are defined by differences in their hexon proteins. [12, 20]. The latency period may be very short, ranging from to 48 h. Antibodies against one type cannot protect against infection with other types [12]. Neutralizing antibodies against FAdV-9 may neutralize the antigen of FAdV-3, but they are not specific [6]. Cross-neutralisation has also been observed between FAdV-2 and FAdV-11/D. These types are closely related, and cross-neutralisation between them has been reported previously [28].
The aim of this study was to estimate the occurrence and pathogenicity of adenovirus strains FAdV-1-61/11z and FAdV-8a-6/12j in experimentally infected specific-pathogen-free (SPF) chickens.
Materials and methods
Chicken embryo fibroblast (CEF) cultures
CEF cultures were prepared from 9- to 11-day-old SPF chicken embryos (Lohmann, Germany) according to standard procedures. Eagle’s growth medium (MEM) supplemented with 10% foetal bovine serum and 1% antibiotic-antimycotic mixture (Gibco, UK) was used. The maintenance medium consisted of MEM with 1% antibiotic-antimycotic mixture. A monolayer of the CEFs was formed after about 24 h of incubation at 37.5 °C.
Virus strains
Virus strain FAdV-1/A-61/11z was isolated from the gizzard of a 21-day-old broiler Ross 308, in which postmortem examination revealed changes in the liver that were characteristic of IBH. The field strain FAdV-8a/E-6/12j was isolated from the intestine of a 42-week-old MESSA hen, in which Marek’s disease virus had also been detected.
Standards
Tenfold serial dilutions of the standard virus strain ATCC grp1 were used for the quantification of unknown samples.
DNA quantification
For the quantification of DNA, amplicons of 830 bp from the positive samples were purified and ligated into pGEM-T vector (3015 bp) as per the manufacturer’s instructions (Promega USA). Clones were then isolated and quantified.
Calculation of TCID50
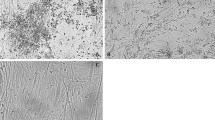
The virus strains were propagated in CEF cultures and were incubated at 37 °C for five days in the presence of 5% of CO2. The cells were examined daily for the appearance of a cytopathic effect (CPE), which is characteristic of an adenovirus infection. Three passages were made. The isolates were kept at -20 °C for the next step of the study. The TCID50 values of the virus strains were determined in 24-well plates (Thermo Scientific, USA) covered with CEF cultures (18-24 h). The CEFs were infected with tenfold dilutions of the virus stocks, from 10−1.0 to 10−7.0: three wells for each dilution and three wells for the negative control. The plates were incubated at 37.5 °C with 85% humidity in an atmosphere of 5% CO2, and the cells were examined daily for the appearance of CPE, using a microscope (Zeiss HXP 120, Germany). After six to seven days of incubation, the TCID50 was determined by the method of Reed & Muench [34]. The DNA of two FAdV strains was extracted from CEFs using a QIAamp Mini-Kit (QIAGEN, Germany) according to the manufacturer’s instructions. The negative DNA controls were extracted from uninfected CEFs. The DNA was stored at -20 °C and later used as the template for sequencing. Extracted FAdVs DNA was tested by PCR and RT-PCR to confirm the absence of other pathogens such as avian reovirus (ARV) [39], chicken anemia virus (CAV) [2], infectious bursal disease virus (IBDV) [38], Marek’s disease (MD) [28], and plaque passage was performed.
Amplification of a target gene by PCR
The primers, FAdVF JSN (5’aatgtcacnaccgaraaggc3’) and FAdVR JSN (5’cbgcbtrcatgtactggta 3’) were used for PCR. They were designed using Primer 3 software (http://bioinfo.ut.ee/primer3-0.4.0/) and were based on a loop L1 HVR1-4 fragment of the hexon gene that is strictly conserved among FAdV types, at nucleotide position 178-1017 (based on FAdV-1, GenBank accession number AC_000014).
PCR was conducted in a final volume of 25 µL of reaction mixture, which contained 2.5 μL of 10x PCR buffer, 1 μL of dNTPs (10 mM) (Fermentas, USA), 1.5 μL of each of the primers (10 μM), 2 μL of the DNA template, 11.5 μL of sterile water, and 1.0 μL of DNA polymerase. After pre-denaturation at 95 °C for five min, the extract was denatured at 94 °C for 45 s, and the primer was annealed at 55 °C for 1 min, followed by product elongation at 72 °C for 2 min and a final elongation at 72 °C for 10 min. Thirty-five amplification cycles were performed using a basic gradient thermocycler (Biometra, Germany). The results were considered positive if the resulting DNA product had the predicted size of 830 bp. After the amplification, the two PCR products were purified using a NucleoSpin Extract II Kit (Macherey-Nagel, France) and sent to GenoMed (Warsaw) for sequencing.
Amplification of the target gene by real-time PCR
Real-time PCR was conducted in an ABI7500 system (Applied Biosystems, USA) in a final volume of 25 µL containing 12.5 µL of 2x Master Mix SYBR Green (QIAGEN, Germany) 1.0 µL of the primers FAdV-1F and FAdV-1R or FAdV-8aF and FAdV-8aR (40 pM each), 8.5 µL of PCR-grade water, and 2.0 µL of DNA template. For type 8a/E, the primers for real-time PCR were FAdV-8aF (5’-GACAGAGGTCCTTCCTTCAA-3’) and FAdV-8aR (5’-TCAGGCTATCGGTAAAGTCC-3’), and the Taq Man probe was JSNRT/8a/E (5’-AATCCCTACTCGAACACCCC-3’). For type 1/A, the primers were FAdV-1 F (5’-TTCGAGATCAAGAGGCCAGT-3’), and FAdV-1 R (5’-GGTCGAAGTTGCGTAGGAAG-3’), and the Taq Man probe was JSN RT1/A (5’-AATCCCTACTCCAACACCCC-3’). After a pre-denaturation step at 95 °C for 15 min, 40 cycles were done with subsequent signal acquisition, at 94 °C for 30 s and 55 °C for 45 s for the second step. This was followed by melting curve analysis at temperatures ranging from 55 °C to 95 °C. The fluorescence signal at the cycle threshold (Ct) exceeded a defined threshold limit. Samples that reached the threshold before 36 cycles were considered positive. The quantity of adenovirus DNA was determined from a standard curve generated using a set of five nucleic acid standards ranging from 108 to 104 copies/ml for type 1/A and 8a/E, respectively.
Sensitivity and specificity
Sensitivity and specificity were determined based on an amplification curve and a Ct value obtained using tenfold dilutions from 101.0 to 105.0 of DNA extracted from CEFs infected with the reference strains, representing types FAdV-1/A and FAdV-8a/E, which were detected using specific primers. The sensitivity, expressed as copies per µl, was determined for both FAdV types. The specificity of the primers was checked to exclude cross-reactivity with other pathogens.
Sequence alignment and phylogenetic analysis
Sequencing was performed using the Sanger method and a GS FLX/Titanium sequencer (Roche, Switzerland) at Genomed (Warsaw). The sequences obtained for the loop L1 region of the hexon gene of the FAdVs strains 1/A-61/11z (KX247012) and 8a/E-6/12j (KP890032) were compared with sequences of adenovirus strain 1/A (AC_000014) and fowl adenovirus 8a strain (KT862810), respectively, obtained from the GenBank database to determine the level of nucleotide and amino acid sequence identity and to determine the type/species. Sequences of five additional adenovirus species A-E were also obtained from the GenBank database. A phylogenetic tree was constructed using the maximum-likelihood method, and molecular analysis was performed using the computer software MEGA7, Geneious 6.2, and BLAST.
Animals and ethics statement
Sixty one-day-old SPF chickens (Lohmann Tierzucht, Germany) were used to determine the pathogenicity of two FAdV strains. The trial protocol was approved by the local ethics commission. During an experimental period of 4 weeks, the chickens were housed in an animal facility. Every effort was made to minimize their suffering.
Experimental setting
The chickens were divided into three equal groups. The chickens in group I were infected with strain FAdV-1/A-61/11z, which had been obtained from an IBH case, at a dose of 104.0 TCID50/0.2 mL. The chickens in group II were infected with strain FAdV-8a/E-6/12j at a dose of 104.0 TCID50/0.2 mL. The chickens in group III were injected with 0.2 ml of PBS. All chickens were infected and injected intraperitoneally. The doses of the viruses were deemed to be infectious, but not lethal. The birds were kept in isolated rooms, in negatively pressurized BSL3 animal facilities and provided with compound feed and water ad libitum. They were observed daily for clinical signs through a monitoring system throughout the experiment. At 1, 2, 3 and, 4 weeks postinfection, four birds from each group were bled and euthanized according to the required procedure, and necropsy examinations were performed. All birds were euthanized with the approval of the ethics commission. During post-mortem examination, samples of the bursa of Fabricius, liver, and other organs with pathological changes were collected. Samples were also collected from dead birds. DNA was extracted from of 100 mg tissue homogenate in 200 µl PBS, and real-time PCR was conducted.
Virus detection and quantification
The copy number of FAdV-1/A-61/11z and FAdV-8a/E-6/12j in each internal organ was determined by real-time PCR. Each tissue was tested in duplicate. A sample was considered positive when both replicates had similar Ct values (+/-3) and matching curve profiles. Serial dilutions of the standard plasmid from 1 × 101 to 1 × 1010 copies/µl were used to create a standard curve.
Results
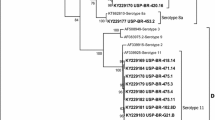
The nucleotide sequence of the loop L1 region of the hexon gene was determined for the isolates FAdV-1/A-61/11z and FAdV-8a/E-6/12j, and each was compared to that of a selected reference strain: AC_000014 and KT862810 (FAdV-1/A and FAdV-8a/E), respectively. A phylogenetic tree is shown in Fig. 1, and sequence comparisons are shown in Fig. 2. A molecular analysis of adenovirus strains showed that they are divided into two main branches representing types A/1 and E/8a. Interestingly, strain FAdV-1/A-61/11z is not closely related to the other GEU-causing strains, but it is closely related to the reference strain AC000014 (Fig. 2). Strain FAdV-1/A-61/11z was found to contain 17 substitutions and 5 deletions (Fig. 2). Three differences were found between the sequences of FAdV-8a/E-6/12j and FAdV-8aTR 59: 436 C→T, 504 A→G, and 507 T→C. The results are in agreement with the ICTV classification of adenoviruses.
Determination of the TCID50 of the propagated strain
In the next step of the study, the TCID50 of the 1/A-61/11z strain was found to be 105.5/ml, and that of the 8a/E-6/12j strain was found to be 104.5/ml, using the Reed-Muench method [34].
Clinical signs and lesions
1/A and 8a/E isolates from chickens with IBH from field outbreaks of the disease were used for experimental infection to test whether they would cause IBH in chickens. The results of the clinical observation of healthy, infected, and dead birds are presented in Table 1. Starting from the second day, depression was observed in up to 14 chickens in group I, which were infected with the FAdV-1/A-61/11z strain. The chickens adopted a crouching position, with ruffled feathers. At the end of the first week postinfection, two of the birds died. A postmortem examination revealed petechial hemorrhage and hepatomegaly in the liver and other internal organs. The liver was pale, fragile, and somewhat swollen. Yellowish mucous membranes were also observed. In the remaining birds from this group, clinical signs decreased on the tenth day postinfection (Table 1), and no clinical signs were observed thereafter. Necropsy at the second and third week postinfection showed splenomegaly in three birds. Hepatomegaly, with a fragile consistency of liver tissue, petechiae, and ecchymoses were also observed in three birds. The bursa of Fabricius was smaller, and involution (weight ratio) was observed in two birds. Kidneys were pale and enlarged, with petechiae. At the fourth week postinfection, the birds were in good condition before euthanasia. Post-mortem examination did not reveal any pathological changes. The mortality rate in this group was 10%. No gizzard erosions were seen in any of the birds.
Clinical manifestations such as depression were observed in up to seven chickens in group II, which were inoculated with the FAdV-8a/E-6/12j strain, starting from 48 h postinfection. The chickens showed a sudden onset of depression characterized by ruffled feathers and sitting in a crouching position. On the third day postinfection, necropsy revealed hepatomegaly and yellowish mucous membranes, and only minimal swelling in the liver was observed in two out of four chickens. The rest of the internal organs did not show any pathological changes. At the end of the second week, the clinical symptoms decreased. On the second day postinfection, the chickens did not show any clinical signs. Post-mortem examination at the first and second weeks postinfection showed hepatomegaly with a change in the physiological colour of the liver and the mucous membrane in three birds. In the rest of the internal organs, no pathological changes were found. The birds did not show any clinical symptoms from the third and fourth weeks postinfection until the end of the experiment. At the time that the chickens were euthanized, they were in good condition. No deaths occurred in this group.
After the first day postinfection, neither clinical symptoms nor gross lesions characteristic of inclusion body hepatitis were found in the chickens of group III. Chickens that were euthanized in the first, second, third, and fourth weeks of the experiment did not show any pathological lesions.
Detection and quantification of virus particles
The detection range of the real-time PCR assays used for detection of 1/A and 8a/E regarding specificity was evaluated to exclude cross-reactivity with other pathogens. The sensitivity was found to be 8 copies/µl for both 1/A and 8a/E. Significantly higher copy numbers of the hexon gene in the internal organs of chickens in group I, which were infected with the FAdV-1-61/11z strain, were observed in comparison to group II, and the amount of virus was between 107 and 1015 copies per mg of tissue (R2 = 0.991, Eff% = 91.03). The highest copy numbers of the hexon gene were found in the gizzard, liver, and intestine at the first, third, and fourth weeks postinfection, and these values ranged from 1012 to 1014 copies per mg of tissue. The lowest copy numbers of the hexon gene (about 100.5 copies per mg of tissue) were found in the spleen, kidney, bursa of Fabricius, and thymus at the third week postinfection. The liver, thymus and gizzard had the highest copy numbers in the first, second and fourth weeks postinfection, and the kidney, thymus and spleen had the lowest copy numbers in the third week of the experiment. These results are presented in Fig. 3. There was no correlation between certain lesions and the presence of a particular virus type. The copy number of the hexon gene of the adenovirus strain FAdV-8a-6/12j in internal organs of chickens from group II was determined to be between 100.5 and 108 copies per mg of tissue (R2 = 0.997, Eff% = 95.177), depending on the examined organs and time postinfection. The copy numbers were highest in gizzards in the first week postinfection and lowest in the bursa of Fabricius in the second, third, and fourth week postinfection. However, the copy number of the hexon gene was lower than that obtained for chickens infected with the FAdV-1-61/11z strain, which produced about 1014 copies per mg of tissue at week 1 in the liver and thymus, and at week 4, about 1014 copies per mg of tissue were found in the liver, gizzard, and intestine. The lowest copy number of the hexon gene was found at week 4 postinfection in the gizzard (about 8 copies per mg of tissue). The results are presented in Fig. 4. No evidence of the presence of the hexon gene was obtained by real-time PCR in the tissues of chickens from group III at any time point. The results are presented in Table 1.
Discussion
A sequence alignment revealed that the strains examined in this study represented different types and some differences were observed in the loop L1 HVR1-4 region of the hexon gene.
To date, the mechanisms of adenovirus pathogenicity are not well understood [13]. Strains of the same type or species can have different pathogenicity. These same strains may be found in both GEU and IBH cases. The full genome sequence of the FAdV-1-61/11z virus strain was reported by Matczuk et al. in 2017 [22]. Interestingly, some strains of FAdV-8a may be implicated in the development of GEU [16, 17, 19] and IBH [14, 21, 23, 24, 37]. Adenoviruses have been suggested to play a role in immunodeficiency [24, 28, 35]. They can establish latency for a period of time, after which the pathogenic process can be induced by different pathogens, which stimulate them to replicate. Adenovirus strains are mainly isolated from broilers, and rarely from reproduction flocks [11, 22]. However, there is still no correlation between the disease profile in production flocks and the presence of a particular type. IBH is normally detected in meat-production birds and may be found in birds younger than seven days of age. The literature suggests that many different types, including FAdV-4/C, 2-3-9-11/D are associated with IBH outbreaks in many countries in chickens less than 3 weeks old.
A study was undertaken to investigate whether adenovirus strains of different types (1/A and 8a/E) have differences in pathogenicity related to their genome structure and sequence variations. A previous study conducted by Niczyporuk in 2014 in which strains belonging to type FAdV-7/E were examined in vitro and in vivo, proved to be crucial for the inhibition of the replication of Marek’s disease virus vaccine strains Rispens/CVI988 and FC126(HVT), and the inhibition of the immunoprophylactic effect after vaccination against Marek’s disease. However, clinical manifestations and post-mortem examinations gave a completely different clinical picture for type 8a/E and type 1/A, and the mortality rates differed. For both types, adenovirus genetic material was located mainly in the large intestine and liver, rarely in the gizzard, and occasionally in the spleen, kidney, and bursa of Fabricius.
The highest replication rates of the FAdV-7/E strain were found in the bursa of Fabricius and liver, and the amount of virus was determined to be between 102.5 and 105 copies per mg in these tissues. A relationship between the adenovirus type and replication in internal organs was also observed by Cook [3]. Steer et al. [36] observed a peak of disease at 5 and 6 d.p.i with type 8b/E and 11/D, respectively. That study suggested that type 8b/E may have a shorter incubation period and be more virulent than 11/D. The hexon gene copy number was also higher than that of type 7/E [28] and similar to that of type 8a/E in our study. However, at week 3, the copy number of the hexon gene in gizzards was 101.0 copies per mg of tissue for type 8a/E and about 1014 copies per mg of tissue for type 1/A. Steer et al. [36] also suggested that types 8b/E and 11/D may play a role as immunosuppressive agents and indicated that types 8b/E, 11/D and other IBH-associated FAdVs have a predilection for the gizzard, based on an in vivo study that suggested that the epithelial lining of the gizzard and alimentary tract is a reservoir of FAdVs and plays an important role in virus transmission. Cook et al. [3] indicated that the replication rate of adenovirus strains reached a peak in the first three weeks postinfection and that those strains had a higher degree of tropism for the digestive tract than for other internal organs. Similar results were obtained by other authors [9, 10, 30], also showing that the rate of replication of strain FAdV-1/A is highest in the gizzard, liver, spleen, and intestine.
To conclude, we found higher copy numbers of the hexon gene in the organs of chickens infected with strain FAdV-1/A-61/11z (group I), than in those infected with strain FAdV-8a/E-6/12j (group II), and the difference in the copy number was about 7 log10. The FAdV-1/A-61/11j strain was also more virulent than the other examined strain. The mutations in the examined region of the hexon gene and the resulting amino acid changes could be important for pathogenicity. This is the first report of an experimental study of FAdV-8a/E in chickens in Poland.
References
Ballman M, Harrach B (2016) Detection and partial genetic characterisation of novel avi- and siadenoviruses in racing and fancy pigeons (Columba livia domestica). Acta Veta Hung 64:514–528
Carrie J, Markowski G, Miller MM, Schat KA (2002) Development of strain- specific real-time PCR and RT-PCR assays for quantitation of chicken anemia virus. J Virol Methods 101:135–147
Cook JK (1983) Fowl adenoviruses: studies on aspects of the pathogenicity of six strains for 1-day-old chicks. Avian Pathol 12:35–43
Choi KS, Kye SJ, Kim JY, Jeon WJ, Lee EK, Park KY (2012) Epidemiological investigation of outbreaks of fowl adenovirus infection in commercial chickens in Korea. Poult Sci 91:2502–2506
Dar A, Gomis S, Shirle I, Mutwiri G, Brownlie R, Potter A, Gerdts V, Tikoo SK (2012) Pathotypic and molecular characterization of fowl adenovirus associated with inclusion body hepatitis in Saskatchewan chickens. Avian Dis 56:73–81
Fitzgerald SD (2008) Adenovirus infections. In: Saif YM, Fadly AM, Glisson JR, McDougald LR, Nolan LK, Swayne DE (eds) Diseases of poultry, 12th edn. Blackwell, Ames, pp 251–252
Ganesh K, Suryanarayana V, Raghavan R, Gowda S (2001) Nucleotide sequence of L1 and part of P1 of hexon gene of fowl adenovirus associated with hydropericardium hepatitis syndrome differs with the corresponding region of other fowl adenoviruses. Vet Microbiol 7:1–11
Gjevre AG, Kaldhusdal M, Eriksen GS (2013) Gizzard erosion and ulceration syndrome in chickens and turkeys: a review of causal or predisposing factors. Avian Pathol 42:297–303
Günes A, Marek A, Grafl B, Berger E, Hess M (2012) Real-time PCR assay for universal detection and quantitation of all five species of fowl adenoviruses (FAdV-A to FAdV-E). J Virol Methods 183:147–153
Grafl B, Liebhart D, Gunes A, Wernsdorf P, Aigner F, Bachmeier J, Hess M (2013) Quantity of virulent fowl adenovirus serotype 1 correlates with clinical signs, macroscopical and pathohistological lesions in gizzards following experimental induction of gizzard erosion in broilers. Vet Res 38:1–8
Grgic H, Yang DH, Nagy E (2011) Pathogenicity and complete genome sequence of a fowl adenovirus serotype 8 isolate. Virus Res 156:91–97
Harrach B (2008) Adenoviruses: General Features. In: Mahy BWJ, Van Regenmortel RMH (eds) Encyclopedia of virology (third edition). Academic Press of Elsevier, Oxford, pp 1–9
Harrach B, Kaján GL (2011) Aviadenovirus. Adenoviridae. In: Tidona CA, Darai G (eds) Springer index of viruses. Springer, New York, pp 13–28
Kajan GL, Kecskemeti S, Harrach B, Benko M (2013) Molecular typing of fowl adenoviruses, isolated in Hungary recently, reveals high diversity. Vet Microbiol 167:357–363
Kecskemeti S, Bistyak A, Matiz K, Rt Glavits, Kajan GL, Benko M (2012) Observations on gizzard ulcers caused by adenovirus in chickens. Magyar Allatorvosk Lapia 134:145–149
Lim TH, Kim BY, Kim MS, Jang JH, Lee DH, Kwon YK, Lee JB, Park SY, Choi IS, Song CS (2012) Outbreak of gizzard erosion associated with fowl adenovirus infection in Korea. Poultry Science Association Inc, Fayetteville
Manarolla G, Pisoni G, Moroni P, Gallazzi D, Sironi G, Rampin T (2009) Adenoviral gizzard erosions in Italian chicken flocks. Vet Res 164:754–756
Marek A, Nolte V, Schachner A, Berger E, Schlötterer C, Hess M (2012) Two fiber genes of nearly equal lengths are a common and distinctive feature of fowl adenovirus C members. Vet Microbiol 156:411–417
Marek A, Schulz E, Hess C, Hess M (2010) Comparison of the fibers of Fowl adenovirus A serotype 1 isolates from chickens with gizzard erosions in Europe and apathogenic reference strains. J Vet Diagn Investig 22:937–941
Marek A, Kaján GL, Kosiol C, Harrach B, Schlötterer C, Hess M (2014) Complete genome sequences of pigeon adenovirus 1 and duck adenovirus 2 extend the number of species within the genus Aviadenovirus. Virology 463:107–114
Mase M, Nakamura K, Minami F (2012) Fowl adenoviruses isolated from chickens with inclusion body hepatitis in Japan, 2009–2010. J Vet Med Sci 74:1087–1089
Matczuk AK, Niczyporuk JS, Kuczkowski M, Woźniakowski G, Nowak M, Wieliczko A (2017) Whole genome sequencing of fowl aviadenovirus A—a causative agent of gizzard erosion and ulceration, in adult laying hens. Infect Genet Evol 48:47
McConnell BA, Fitzgerald AS (2008) Group I adenovirus infections. In: Saif YM (ed) Diseases of poultry, 12th edn. Blackwell/Iowa State Press, Ames, pp 252–266
McFerran JB, Adair BM (2008) Group I Adenovirus Infections. In: Saif YM, Barnes HJ, Glisson JR (eds) Diseases of poultry, 12th edn. Blackwell, Ames, pp 214–227
McFerran JB, Adair BM (1977) Avian adenoviruses: a review. Avian Pathol 6:189–217
Meulemans G, Couvreur B, Decaesstecker M, Boschmans M, Van den Berg TP (2004) Phylogenetic analysis of fowl adenoviruses. Avian Pathol 33:164–170
Mittal D, Jindal N, Tiwari AK, Khokhar RS (2014) Characterization of fowl adenoviruses associated with hydropericardium syndrome and inclusion body hepatitis in broiler chickens. Virus Dis 25:114–119
Niczyporuk JS (2014) Molecular characteristic on occurrence of fowl adenovirus field strains and effect of efficiency on prophylactic vaccinations against Marek’s disease. Doctoral dissertation, National Veterinary Research Institute, Pulawy, pp 1–148
Niczyporuk JS (2016) Phylogenetic and geographic analysis of fowl adenovirus field strains isolated from poultry in Poland. Arch Virol 1:32–42
Okuda Y, Ono M, Shibata I, Sato S, Akashi H (2006) Comparison of the polymerase chain reaction restriction fragment length polymorphism pattern of the fiber gene and pathogenicity of serotype-1 fowl adenovirus isolates from gizzard erosions and room feces of clinically healthy chickens in Japan. J Vet Diagn Investig 18:162–167
Okuda Y, Ono M, Shibata I, Sato S (2004) Pathogenicity of serotype 8 fowl adenovirus isolated from gizzard erosions of slaughtered broiler chickens. J Vet Med Sci 66:1561–1566
Ono M, Okuda Y, Yazawa S, Shibata I, Tanimura N, Kimura K, Haritani M, Mase M, Sato S (2001) Epizotic outbreaks of gizzard erosion associated with adenovirus infection in chickens. Avian Dis 45:268–275
Pallister J, Wright PJ, Sheppard M (1996) A single gene encoding the fiber is responsible for variations in virulence in the fowl adenoviruses. J Virol 70:5115–5122
Reed L, Muench JH (1938) A simple method of estimating fifty per cent endpoinds. Am J Epidemiol 27:493–497
Schonewille E, Singh A, Gobel TW, Gerner W, Saalmuller A, Hess M (2008) Fowl adenovirus (FAdV) serotype 4 causes depletion of B and T cells in lymphoid organs in specific pathogen-free chickens following experimental infection. Vet Immunol Immunopathol 121:130–139
Steer PA, Sandy JR, O’Rourke D, Scott PC, Browning GF, Noormohammadi AH (2015) Chronological analysis of gross and histological lesions induced by field strains of fowl adenovirus serotypes 1, 8b and 11 in one-day-old chickens. Avian Pathol 44:106–113
Steer PA, Rourke DO, Ghorashi SA, Noormohammadi AH (2011) Application of high- resolution melting curve analysis for typing of fowl adenoviruses in field cases of inclusion body hepatitis. Aust Vet J 89:184–192
Tomás G, Hernández M, Marandino A, Techera C, Grecco S, Hernández D, Banda A, Panzera Y, Pérez R (2017) Development of an RT-qPCR assay for the specific detection of a distinct genetic lineage of the infectious bursal disease virus. Avian Pathol 2:150–156
Woźniakowski G, Niczyporuk JS, Samorek-Salamonowicz E, Gaweł A (2015) The development and evaluation of cross-priming amplification for the detection of avian reovirus. J Appl Microbiol 118:528–536
Zadravec M, Slavec B, Krapež U, Kaján GL, Račnik J, Juntes P, Rojs OZ (2011) Inclusion body hepatitis associated with fowl adenovirus type 8b in broiler flock in Slovenia—a case report. Slov Vet Res 48:107–113
Zadravec M, Brigita Slavec, Krapež U, Gl Kaján, Račnik J, Polona Juntes, Rahela Juršič-Cizerl, Mária Benkõ, Olga Zorman-Rojs (2013) Inclusion body hepatitis (IBH) outbreak associated with fowl adenovirus type 8b in broilers. Acta Vet 63:101–110
Acknowledgements
We acknowledge Mrs. Wieslawa Deska for technical support.
Funding
This study was partially funded by National Veterinary Research Institute (program number S-192 ‘Epidemiology and pathogenicity of adenovirus strains’).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare that they have no competing financial, professional, or non-financial interests that might be perceived to influence the interpretation of data or presentation of information described in this article.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable national and institutional guidelines for the care and use of animals were followed. This article does not contain any studies with human participants.
Additional information
Handling Editor: Roman Pogranichniy.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Niczyporuk, J.S., Czekaj, H. A comparative pathogenicity analysis of two adenovirus strains, 1/A and 8a/E, isolated from poultry in Poland. Arch Virol 163, 3005–3013 (2018). https://doi.org/10.1007/s00705-018-3965-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-018-3965-9