Abstract
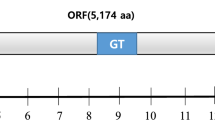
An RNA virus tentatively named Lagenaria siceraria endornavirus-Hubei (LsEV-HuB) was isolated from Lagenaria siceraria var. hispida in Hubei, China. The LsEV-HuB genome consists of 15,098 bp and contains a single open reading frame (ORF) encoding a large protein with several conserved domains, including one helicase domain, one glycosyltransferase domain, two capsular polysaccharide synthesis protein (CPS) domains, and one RNA-dependent RNA polymerase (RdRp) domain. LsEV-HuB has nucleotide and amino acid sequence identities of 72.96% and 77.95%, respectively, to Lagenaria siceraria endornavirus-California (LsEV-CA), the closest relative of LsEV-HuB.


Similar content being viewed by others
References
Adams MJ, Lefkowitz EJ, King AMQ, Harrach B, Harrison RL, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Mushegian AR, Nibert M, Sabanadzovic S, Sanfaçon H, Siddell SG, Simmonds P, Varsani A, Zerbini FM, Gorbalenya AE, Davison AJ (2017) Changes to taxonomy and the International Code of virus classification and nomenclature ratified by the International Committee on Taxonomy of Viruses. Arch Virol 162:2505–2538
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Du ZG, Lin WW, Qiu P, Liu XJ, Guo LF, Wu KC, Zhang SB, Wu ZJ (2016) Complete sequence of a double-stranded RNA from the phytopathogenic fungus Erysiphe cichoracearum that might represent a novel endornavirus. Arch Virol 161:2343–2346
Espach Y, Maree HJ, Burger JT (2012) Complete genome of a novel endornavirus assembled from next-generation sequence data. Virol J 86:13142
Fukuhara T, Moriyama H, Pak JY, Hyakutake H, Nitta T (1993) Enigmatic double-stranded RNA in Japonica rice. Plant Mol Biol 21:1121–1130
Hacker C, Brasier C, Buck K (2005) A double-stranded RNA from a Phytophthora species is related to the plant endornaviruses and contains a putative UDP glycosyltransferase gene. J Gen Virol 86:1561–1570
Kwon SJ, Tan SH, Vidalakis G (2014) Complete nucleotide sequence and genome organization of an endornavirus from bottle gourd (Lagenaria siceraria) in California, USA. Virus Genes 49:163–168
Lim S, Kim KH, Zhao FM, Yoo RH, Igori D, Lee SH, Moon JS (2015) Complete genome sequence of a novel endornavirus isolated from hot pepper. Arch Virol 160:3153–3156
Li W, Zhang T, Sun HY, Deng YY, Zhang AX, Chen HG, Wang KR (2014) Complete genome sequence of a novel endornavirus in the wheat sharp eyespot pathogen Rhizoctonia cerealis. Arch Virol 159:1213–1216
Morris TJ, Dodds JA (1979) Isolation and analysis of double stranded RNA from virus infected plant and fungal tissue. Phytopathology 69:854–858
Moriyama H, Horiuchi H, Nitta T, Fukuhara T (1999) Unusual inheritance of evolutionarily related double-stranded RNAs in interspecific hybrid between rice plants Oryza sativa and Oryza rufipogon. Plant Mol Biol 39:1127–1136
Moriyama H, Nitta T, Fukuhara T (1995) Double-stranded RNA in rice: a novel RNA replicon in plants. Mol Gen Genet 248:364–369
Okada R, Kiyota E, Sabanadzovic S, Moriyama H, Fukuhara T, Saha P, Roossinch MJ, Severin A, Valverde RA (2011) Bell pepper endornavirus: molecular and biological properties, and occurrence in the genus Capsicum. J Gen Virol 92:2664–2673
Okada R, Yong CK, Valverde RA, Sabanadzovic S, Aoki N, Hotate S, Kiyota E, Moriyama H, Fukuhara T (2013) Molecular characterization of two evolutionarily distinct endornaviruses coinfecting common bean (Phaseolus vulgaris). J Gen Virol 94:220–229
Ong JWL, Li H, Sivasithamparam K, Dixon KW, Jones MGK, Wylie SJ (2016) Novel endorna-like viruses, including three with two open reading frames, challenge the membership criteria and taxonomy of the Endornaviridae. Virology 499:203–211
Osaki H, Nakamura H, Sasaki A, Matsumoto N, Yoshida K (2006) An endornavirus from a hypovirulent strain of the violet root rot fungus, Helicobasidium mompa. Virus Res 118:143–149
Park Y, Chen X, Punja ZK (2006) Diversity, complexity and transmission of double-stranded RNA elements in Chalara elegans (synanam. Thielaviopsis basicola). Mycol Res 110:697–704
Pfeiffer P (1998) Nucleotide sequence, genetic organization and expression strategy of the double-stranded RNA associated with the ‘447’ cytoplasmic male sterility trait in Vicia faba. J Gen Virol 79:2349–2358
Roossinck MJ, Sabanadzovic S, Okada R, Valverde RA (2011) The remarkable evolutionary history of endornaviruses. J Gen Virol 92:2674–2678
Sabanadzovic S, Wintermantel WM, Valverde RA, McCreight JD, Aboughanem-Sabanadzovic N (2016) Cucumis melo endornavirus: genome organization, host range and co-divergence with the host. Virus Res 214:49–58
Sela N, Luria N, Dombrovsky A (2012) Genome assembly of Bell pepper endornavirus from small RNA. J Virol 86:7721
Stielow B, Klenk HP, Menzel W (2011) Complete genome sequence of the first endornavirus from the ascocarp of the ectomycorrhizal fungus Tuber aestivum Vittad. Arch Virol 156:343–345
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Villanueva F, Sabanadzovic S, Valverde RA, Navas-Castillo J (2012) Complete genome sequence of a double-stranded RNA virus from avocado. J Virol 86:1282–1283
Yeku O, Frohman MA (2011) Rapid amplification of cDNA ends (RACE). Methods Mol Biol 67:233–238
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This work was supported by the Special Fund for Agro-Scientific Research in the Public Interest (No. 201303028), the National Natural Science Foundation of China (No. 31301638) and the Opening Fund of Engineering Research Center of Ecology and Agricultural Use of Wetland, Ministry of Education (KF201501). We are grateful to Dr. Qingchao Deng at the College of Agriculture, Yangtze University, and Dr. Zhenguo Du at the College of Plant Protection, Fujian Agriculture and Forestry University, for their help in preparing the manuscript.
Conflict of interest
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Sead Sabanadzovic.
Rights and permissions
About this article
Cite this article
Peng, X., Pan, H., Muhammad, A. et al. Complete genome sequence of a new strain of Lagenaria siceraria endornavirus from China. Arch Virol 163, 805–808 (2018). https://doi.org/10.1007/s00705-017-3664-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3664-y