Abstract
Key message
A set of single-nucleotide polymorphisms (SNPs) was discovered using a de novo transcriptome for radiata pine. The reference transcriptome and SNPs were annotated according to the functional information available in biological databases.
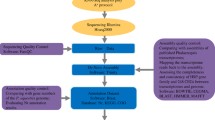
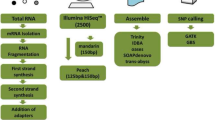
A set of SNPs was discovered using a de novo transcriptome assembly for radiata pine in Chile. De novo transcriptome was generated by Illumina sequences and assembled by Trinity. A set of unique genes and transcription factors were identified. All dataset sequences were mapped against the de novo reference transcriptome and SNPs were called. A total of 300,761 SNPs were identified between samples, where 38,730 SNPs had a functional annotation. Since there is little data available of radiata pine-omics, the aim of this work is to provide to the scientific community a de novo transcriptome assembly and a set of SNPs.
Similar content being viewed by others
References
Ahuja MR, Neale DB (2005) Evolution of genome size in conifers. Silvae Genet 54:126–137. https://doi.org/10.1515/sg-2005-0020
Birol I, Raymond A, Jackman SD, Pleasance S, Coope R et al (2013) Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data. Bioinformatics 29:1492–1497. https://doi.org/10.1093/bioinformatics/btt178
Brown CT, Howe A, Zhang Q, Pyrkosz AB, Brom TH (2012) A reference-free algorithm for computational normalization of shotgun sequencing data. arXiv preprint arXiv:1203.4802
Cairney J, Zheng L, Cowels A, Hsiao J, Zismann V et al (2006) Expressed sequence tags from loblolly pine embryos reveal similarities with angiosperm embryogenesis. Plant Mol Biol 62:485–501. https://doi.org/10.1007/s11103-006-9035-9
Canales J, Bautista R, Label P, Gómez-Maldonado J, Lesur I et al (2014) De novo assembly of maritime pine transcriptome: implications for forest breeding and biotechnology. Plant Biotechnol J 12:286–299. https://doi.org/10.1111/pbi.12136
Cañas RA, Feito I, Fuente-Maqueda JF, Ávila C, Majada J, Cánovas FM (2015) Transcriptome-wide analysis supports environmental adaptations of two Pinus pinaster populations from contrasting habitats. BMC Genom 16:909. https://doi.org/10.1186/s12864-015-2177-x
Cánovas A, Rincon G, Islas-Trejo A, Wickramasinghe S, Medrano JF (2010) SNP discovery in the bovine milk transcriptome using RNA-seq technology. Mamm Genome 21:592–598. https://doi.org/10.1007/s00335-010-9297-z
Carrasco A, Wegrzyn JL, Durán R, Fernández M, Donoso A, Rodriguez V, Neale D, Valenzuela S (2017) Expression profiling in Pinus radiata infected with Fusarium circinatum. Tree Genet Genomes 13:46. https://doi.org/10.1007/s11295-017-1125-0
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6(2):80–92. https://doi.org/10.4161/fly.19695
Clarke K, Yang Y, Marsh R, Xie L, KeK Z (2013) Comparative analysis of de novo transcriptome assembly. Sci China Life Sci 56:156–162. https://doi.org/10.1007/s11427-013-4444-x
Core Team R (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Viena
Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11:485
Dillon SK, Nolan M, Li W, Bell C, Wu HX, Southerton SG (2010) Allelic variation in cell wall candidate genes affecting solid wood properties in natural populations and land races of Pinus radiata. Genetics 185:1477–1487. https://doi.org/10.1534/genetics.110.116582
Dillon SK, Nolan MF, Matter P, Gapare WJ, Bragg JG, Southerton SG (2013) Signatures of adaptation and genetic structure among the mainland populations of Pinus radiata (D. Don) inferred from SNP loci. Tree Genet Genomes 9:1447–1463. https://doi.org/10.1007/s11295-013-0650-8
Eckert AJ, Pande B, Ersoz ES, Wright MH, Rashbrook VK, Nicolet CM, Neale DB (2009) High-throughput genotyping and mapping of single nucleotide polymorphisms in loblolly pine (Pinus taeda L.). Tree Genet Genomes 5:225–234
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Fan F, Cui B, Zhang T, Qiao G, Ding G, Wen X (2014) The temporal transcriptomic response of Pinus massoniana seedlings to phosphorus deficiency. PLoS ONE 9(8):e105068. https://doi.org/10.1371/journal.pone.0105068
Fernández-Pozo N, Canales J, Guerrero-Fernández D, Villalobos DP, Díaz-Moreno SM, Bautista R, Flores-Monterroso A, Guevara MÁ, Perdiguero P, Collada C (2011) EuroPineDB: a high-coverage web database for maritime pine transcriptome. BMC Genom 12:366
Gonzalez-Ibeas D, Martinez-Garcia PJ, Famula RA, Delfino-Mix A, Stevens KA, Loopstra CA, Langley CH, Neale DB, Wegrzyn JL (2016) Assessing the gene content of the megagenome: sugar pine (Pinus lambertiana). G3: genes. Genomes Genetics 6:3787–3802. https://doi.org/10.1534/g3.116.032805
González-Martinez SC, Wheeler NC, Ersoz E, Nelson CD, Neale DB (2007) Association genetics in Pinus taeda L. I. Wood property traits. Genetics 175:399–409. https://doi.org/10.1534/genetics.106.061127
González-Martínez SC, Huber D, Ersoz E, Davis JM, Neale DB (2008) Association genetics in Pinus taeda L. I. Carbon isotope discrimination. Heredity 101:19
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652. https://doi.org/10.1038/nbt.1883
Granato IS, Galli G, de Oliveira Couto EG, e Souza MB, Mendonça LF, Fritsche-Neto R (2018) snpReady: a tool to assist breeders in genomic analysis. Mol Breed 38:102
Guerrero PC, Bustamante RO (2007) Can native tree species regenerate in Pinus radiata plantations in Chile?: evidence from field and laboratory experiments. For Ecol Manag 253:97–102
Hall DE, Yuen MM, Jancsik S, Quesada AL, Dullat HK, Li M, Henderson H, Arango-Velez A, Liao NY, Docking RT (2013) Transcriptome resources and functional characterization of monoterpene synthases for two host species of the mountain pine beetle, lodgepole pine (Pinus contorta) and jack pine (Pinus banksiana). BMC Plant Biol 13:80
Howe GT, Yu J, Knaus B, Cronn R, Kolpak S, Dolan P, Lorenz WW, Dean JF (2013) A SNP resource for Douglas-fir: de novo transcriptome assembly and SNP detection and validation. BMC Genom 14:137. https://doi.org/10.1186/1471-2164-14-137
Huse SM, Huber JA, Morrison HG, Sogin ML, Welch DM (2007) Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8:R143
Jannink JL, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genom 9:166–177. https://doi.org/10.1093/bfgp/elq001
Jin J, Tian F, Yang DC, Meng YQ, Kong L, Luo J, Gao G (2017) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45:D1040–D1045. https://doi.org/10.1093/nar/gkw982
Kumar R, Qiu J, Joshi T, Valliyodan B, Xu D, Nguyen HT (2007) Single feature polymorphism discovery in rice. PLoS ONE 2(3):e284
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) 1000 Genome project data processing subgroup. the sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li B, Fillmore N, Bai Y, Collins M, Thomson JA, Stewart R, Dewey CN (2014) Evaluation of de novo transcriptome assemblies from RNA-Seq data. Genome Biol 15:553. https://doi.org/10.1186/s13059-014-0553-5
Liu JJ, Sturrock RN, Benton R (2013) Transcriptome analysis of Pinus monticola primary needles by RNA-seq provides novel insight into host resistance to Cronartium ribicola. BMC Genom 14:884
Liu L, Zhang S, Lian C (2015) De Novo transcriptome sequencing analysis of cDNA library and large-scale unigene assembly in japanese red pine (Pinus densiflora). Int J Mol Sci 16:29047–29059. https://doi.org/10.3390/ijms161226139
Liu JJ, Schoettle AW, Sniezko RA, Sturrock RN, Zamany A, Williams H, Ha A, Chan D, Danchok B, Savin DP, Kegley A (2016) Genetic mapping of Pinus flexilis major gene (Cr4) for resistance to white pine blister rust using transcriptome-based SNP genotyping. BMC Genom 17:753. https://doi.org/10.1186/s12864-016-3079-2
López de Heredia U, Vázquez-Poletti JL (2016) RNA-seq analysis in forest tree species: bioinformatic problems and solutions. Tree Genet Genomes 12:30. https://doi.org/10.1007/s11295-016-0995-x
Lorenz WW, Ayyampalayam S, Bordeaux JM, Howe GT, Jermstad KD, Neale DB, Rogers DL, Dean JFD (2012) Conifer DBMagic: a database housing multiple de novo transcriptome assemblies for 12 diverse conifer species. Tree Genet Genomes 8:1477–1485. https://doi.org/10.1007/s11295-012-0547-y
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003) The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4:981–994. https://doi.org/10.1038/nrg1226
Merino I, Abrahamsson M, Sterck L, Craven-Bartle B, Canovas F, von Arnold S (2016) Transcript profiling for early stages during embryo development in Scots pine. BMC Plant Biol 16:255. https://doi.org/10.1186/s12870-016-0939-5
Morse AM, Peterson DG, Islam-Faridi MN, Smith KE, Magbanua Z, Garcia SA, Kubisiak TL, Amerson HV, Carlson JE, Nelson CD, Davis JM (2009) Evolution of genome size and complexity in pinus. PLoS ONE 4(2):e4332. https://doi.org/10.1371/journal.pone.0004332
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628. https://doi.org/10.1038/nmeth.1226
Müller T, Ensminger I, Schmid KJ (2012) A catalogue of putative unique transcripts from Douglas-fir (Pseudotsuga menziesii) based on 454 transcriptome sequencing of genetically diverse, drought stressed seedlings. BMC Genom 13:673
Neale DB, Wegrzyn JL, Stevens KA, Zimin AV, Puiu D et al (2014) Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol 15:1–13
Niu SH, Li ZX, Yuan HW, Chen XY, Li Y, Li W (2013) Transcriptome characterisation of Pinus tabuliformis and evolution of genes in the Pinus phylogeny. BMC Genom 14:263
Nystedt B, Street NR, Wetterbom A, Zuccolo A, Lin YC et al (2013) The Norway spruce genome sequence and conifer genome evolution. Nature 497:579–584. https://doi.org/10.1038/nature12211
Paradis E (2010) pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26:419–420
Parchman TL, Geist KS, Grahnen JA, Benkman CW, Buerkle CA (2010) Transcriptome sequencing in an ecologically important tree species: assembly, annotation, and marker discovery. BMC Genom 11:180
Parent GJ, Raherison E, Sena J, MacKay JJ (2015) Forest tree genomics: review of progress. In: Advances in botanical research, Elsevier, pp 39–92
Pinosio S, González-Martínez SC, Bagnoli F, Cattonaro F, Grivet D, Marroni F, Lorenzo Z, Pausas JG, Verdú M, Vendramin GG (2014) First insights into the transcriptome and development of new genomic tools of a widespread circum-Mediterranean tree species, Pinus halepensis Mill. Mol Ecol Resour 14:846–856. https://doi.org/10.1111/1755-0998.12232
Pop M, Salzberg SL (2008) Bioinformatics challenges of new sequencing technology. Trends Genet 24:142–149. https://doi.org/10.1016/j.tig.2007.12.006
Prager EM, Fowler DP, Wilson AC (1976) Rates of evolution in conifers (Pinaceae). Evolution 30:637. https://doi.org/10.2307/2407806
Rigault P, Boyle B, Lepage P, Cooke JEK, Bousquet J, MacKay JJ (2011) A white spruce gene catalog for conifer genome analyses. Plant Physiol 157:14–28. https://doi.org/10.1104/pp.111.179663
Rogers DL (2004) In situ genetic conservation of a naturally restricted and commercially widespread species, Pinus radiata. For Ecol Manag 197:311–322. https://doi.org/10.1016/j.foreco.2004.05.022
Simao FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. https://doi.org/10.1093/bioinformatics/btv351
Stevens KA, Wegrzyn JL, Zimin A, Puiu D, Crepeau M et al (2016) Sequence of the sugar pine megagenome. Genetics 204:1613–1626. https://doi.org/10.1534/genetics.116.193227
Visser EA, Wegrzyn JL, Steenkmap ET, Myburg AA, Naidoo S (2015) Combined de novo and genome guided assembly and annotation of the Pinus patula juvenile shoot transcriptome. BMC Genom 16:1057. https://doi.org/10.1186/s12864-015-2277-7
Wachowiak W, Trivedi U, Perry A, Cavers S (2015) Comparative transcriptomics of a complex of four European pine species. BMC Genom 16:234. https://doi.org/10.1186/s12864-015-1401-z
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63
Wegrzyn JL, Lee JM, Tearse BR, Neale DB (2008) TreeGenes: a forest tree genome database. Int J Plant Genom, Article ID 412875. https://doi.org/10.1155/2008/412875
Wegrzyn JL, Main D, Figueroa B, Choi M, Yu J et al (2011) Uniform standards for genome databases in forest and fruit trees. Tree Genet Genomes 8(3):549–557. https://doi.org/10.1007/s11295-012-0494-7
Wilhelm BT, Marguerat S, Watt S, Schubert F, Wood V, Goodhead I, Penkett CJ, Rogers J, Bähler J (2008) Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453:1239–1243. https://doi.org/10.1038/nature07002
Yeaman S, Hodgins KA, Suren H, Nurkowski KA, Rieseberg LH, Holliday JA, Aitken SN (2014) Conservation and divergence of gene expression plasticity following c. 140 million years of evolution in lodgepole pine (Pinus contorta) and interior spruce (Picea glauca × Picea engelmannii). New Phytol 203:578–591. https://doi.org/10.1111/nph.12819
Zheng X, Levine D, Shen J, Gogarten S, Laurie C, Weir B (2012) A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28:3326–3328. https://doi.org/10.1093/bioinformatics/bts606
Zimin AV, Stevens KA, Crepeau MW, Puiu D, Wegrzyn JL, Yorke JA, Langley CH, Neale DB, Salzberg SL (2017) An improved assembly of the loblolly pine mega-genome using long-read single-molecule sequencing. GigaScience 6:1–4. https://doi.org/10.1093/gigascience/giw016
Acknowledgements
This project was financed by Genómica Forestal SA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Data archiving statement
The data have been submitted to the NCBI. The bioproject ID related to this paper is PRJNA495832.
Additional information
Communicated by Daniel G. Peterson.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Durán, R., Rodriguez, V., Carrasco, A. et al. SNP discovery in radiata pine using a de novo transcriptome assembly. Trees 33, 1505–1511 (2019). https://doi.org/10.1007/s00468-019-01875-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-019-01875-w