Abstract
Key message
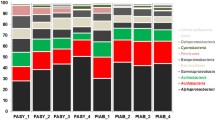
A total of 43 phyla (395 genera) of bacteria and 11 phyla (309 genera) of fungi were identified from normal sapwood (SW), wetwood (WW) and their transition zone wood (TW) in living poplar trees, with bacteria showing significantly higher diversity than fungi. Highly variable community diversities and compositions were found for bacteria but not for fungi among WW, TW and SW. Overall, 22 phyla (95 genera) bacteria and three phyla (11 genera) fungi were significantly different in relative abundance between WW and SW. Nine wood chemical properties were significantly correlated with the relative abundance of top 15 bacterial and fungal genera in all samples.
Abstract
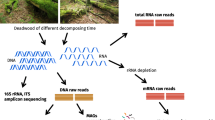
Wetwood represents an abnormal condition of heartwood in living poplar trees. The cause of wetwood formation is still poorly understood, but it is generally considered to involve microbial activities. To facilitate a better understanding of wetwood formation, we sequenced the 16S rRNA and ITS (Internal Transcribed Spacer) genes of distinct regions (16S V4, ITS1) to precisely identify the microbial communities in normal sapwood (SW), wetwood (WW) and their transition zone wood (TW). Nine chemical properties of the wood samples were further determined to elucidate the correlations between the microbial communities and wood characteristics. The total sequences of 16S V4 and ITS1 from WW, TW and SW were assigned to 43 bacterial phyla (395 genera) and 11 fungal phyla (309 genera), indicating a significantly higher diversity of bacteria than fungi. Among SW, TW and WW, significant variations in community diversity and composition were found for bacteria but not for fungi. WW had the highest bacterial and lowest fungal diversities, while SW exhibited the highest fungal diversity. TW generally exhibited intermediate levels between WW and SW. Between WW and SW, 22 bacterial phyla (95 genera) were significantly different in relative abundance. Nine wood chemical properties were significantly correlated with the relative abundance of top 15 bacterial and fungal genera in all wood samples.







Similar content being viewed by others
References
Beckers B, Beeck MOD, Weyens N, Boerjan W, Vangronsveld J (2017) Structural variability and niche differentiation in the rhizosphere and endosphere bacterial microbiome of field-grown poplar trees. Microbiome 5:25. https://doi.org/10.1186/s40168-017-0241-2
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10(1):57–59. https://doi.org/10.1371/journal.pone.0066437
Brill H, Bock E, Bauch J (1981) Über die bedeutung von mikroorganismen im holz von Abies alba Mill. für das tannensterben. Forstwissenschaftliches Centralbltt 100(1):195–206. https://doi.org/10.1007/BF02640636
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peňa AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. PNAS 108(1):4516–4522. https://doi.org/10.1073/pnas.1000080107
Chao L, Zeng D, Sun F, Zhao T, Liu C (1998) Fusarium Proliferatum (Matsushima) Nirenberg—a pathogen causing wetwood in poplar trees. Sci Silvae Sinicae 34(5):61–73
Desai C, Madamwar D (2007) Extraction of inhibitor-free metagenomic DNA from polluted sediments, compatible with molecular diversity analysis using adsorption and ion-exchange treatments. Bioresour Technol 98(4):761–768. https://doi.org/10.1016/j.biortech.2006.04.004
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072. https://doi.org/10.1128/AEM.03006-05
Dische Z (1962) Color reactions of carbohydrates. Methods Carbohyd Chem 1:475–514
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acid Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–998. https://doi.org/10.1038/nmeth.2604
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Etheridge DE, Morin LA (1962) Wetwood formation in balsam fir. Can J Bot 40(10):1335–1345. https://doi.org/10.1139/b62-127
Fry SC (1988) The growing plant cell wall: chemical and metabolic analysis. Longman, London
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methé B, DeSantis TZ, Consortium H-M, Petrosino J-F, Knight R, Birren B-W (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21(3):494–504. https://doi.org/10.1101/gr.112730.110
Huang Y, Wei X, Zhou S, Liu M, Tu Y, Li A, Chen P, Wang Y, Zhang X, Tai H, Peng L, Xia T (2015) Steam explosion distinctively enhances biomass enzymatic saccharification of cotton stalks by largely reducing cellulose polymerization degree in G. barbadense and G. hirsutum. Bioresour Technol 181:224–230. https://doi.org/10.1016/j.biortech.2015.01.020
Jeremic D, Cooper P, Srinivasan U (2004) Comparative analysis of balsam fir wetwood, heartwood, and sapwood properties. Can J Forest Res 34:1241–1250. https://doi.org/10.1139/X08-193
Jiang X, Zhang L, Xu B, Zheng S (1993) Studies on the occurrence, distribution and wood properties of wet heartwood in Populus deltoides I-69. For Res 6(5):480–485
Jin W, Chen L, Hu M, Sun D, Li A, Li Y, Hu Z, Zhou S, Tu Y, Xia T, Wang Y, Xie G, Li Y, Bai B, Peng L (2016) Tween-80 is effective for enhancing steam-exploded biomass enzymatic saccharification and ethanol production by specifically lessening cellulase absorption with lignin in common reed. Appl Energ 175:82–90. https://doi.org/10.1016/j.apenergy.2016.04.104
Knutson DM (1973) The bacteria in sapwood, wetwood, and heartwood of trembling aspen (Populus tremuloides). Can J Bot 51:498–500. https://doi.org/10.1139/b73-060
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AF, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl B-D, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Scott JA, Senés C, Smith ME, Suija A, Taylor DL, Telleria MT, Weiss M, Larsson KH (2013) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22(21):5271–5277. https://doi.org/10.1111/mec.12481
Krause C, Gagnon R (2005) Wet heartwood distribution in the stem, stump, and root wood of black spruce in the Quebec boreal forest, Canada. North J Appl For 22(1):12–18
Krause C, Gagnon R (2006) The relationship between site and tree characteristics and the presence of wet heartwood in black spruce in the boreal forest of Quebec, Canada. Can J For Res 36(6):1519–1526. https://doi.org/10.1139/x06-026
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL, Buckler ES, Ley RE (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. PNAS 110(16):6548–6553. https://doi.org/10.1073/pnas.1302837110
Peng L, Hocart CH, Redmond JW, Williamson RE (2000) Fractionation of carbohydrates in Arabidopsis root cell walls shows that three radial swelling loci are specifically involved in cellulose production. Planta 211(3):406–414. https://doi.org/10.1007/s004250000301
Roque RM, Muńoz-Acosta F, Jeremic D, Berrocal A (2009) Visual identification, physical properties, ash composition, and water diffusion of wetwood in Gmelina arborea. Can J For Res 39(3):537–545. https://doi.org/10.1139/X08-193
Sakamoto Y, Kato A (2002) Some properties of the bacterial wetwood (watermark) in Salix sachalinensis caused by Erwinia salicis. IAWA J 23(2):179–190. https://doi.org/10.1163/22941932-90000296
Sano Y, Fujikawa S, Fukazawa K (1995) Detection and features of wetwood in Quercus mongolica var. grosseserrata. Trees 9(5):261–268. https://doi.org/10.1007/BF00202016
Schink B, Ward JC (1984) Microaerobic and anaerobic bacterial activities involved in formation of wetwood and discoloured wood. IAWA J 5(2):105–109. https://doi.org/10.1163/22941932-90000872
Schink B, Ward JC, Zeikus JG (1981a) Microbiology of wetwood: importance of pectin degradation and Clostridium species in living trees. Appl Environ Microbiol 42(3):526–532
Schink B, Ward JC, Zeikus JG (1981b) Microbiology of wetwood: role of anaerobic bacterial populations in living trees. J Gen Microbiol 123: 313–322. https://doi.org/10.1099/00221287-123-2-313
Schmidt O, Liese W (1994) Occurrence and significance of bacteria in wood. Holzforschung 48:271–277. https://doi.org/10.1515/hfsg.1994.48.4.271
Schofield JA, Hagerman AE, Harold A (1998) Loss of tannins and other phenolics from willow leaf litter. J Chem Ecol 24(8):1409–1421. https://doi.org/10.1023/A:1021287018787
Shaw DC, Edmonds RL, Littke WR, Browning JE, Russel KW (1995) Incidence of wetwood and decay in pre-commercially thinned western hemlock stands. Can J For Res 25(8):1269–1277. https://doi.org/10.1139/x95-140
Sluiter A, Hames B, Ruiz R, Scarlata C, Sluiter J, Templeton D, Crocker D (2008) Determination of structural carbohydrates and lignin in biomass. Technical Report, NREL/TP-510-42618, NREL, Golden Co
Smalla K, Oros-Sichler M, Milling A, Heuer H, Baumgarte S, Becker R, Neuber G, Kropf S, Ulrich A, Tebbe C-C (2007) Bacterial diversity of soil assessed by DGGE, T-RFLP and SSCP fingerprints of PCR-amplified 16S rRNA gene fragment: Do the different methods provide similar results? J Microbiol Meth 69(3):470–479. https://doi.org/10.1016/j.mimet.2007.02.014
Sun J, Zhang Q, Zhou J, Wei Q (2014) Illumina amplicon sequencing of 16S rRNA tag reveals bacterial community development in the rhizosphere of apple nurseries at a replant disease site and a new planting site. PLOS One 9(10):e111744. https://doi.org/10.1371/journal.pone.0111744
Sun M, Xiao T, Ning Z, Xiao E, Sun W (2015) Microbial community analysis in rice paddy soil irrigated by acid mine drainage contaminated water. Appl Microbiol Biotechnol 99(6):2911–2922. https://doi.org/10.1007/s00253-014-6194-5
Tiedemann G, Bauch J, Bock E (1977) Occurrence and significance of bacteria in living trees of Populus nigra L. Eur J For Pathol 7(6):364–374. https://doi.org/10.1111/j.1439-0329.1977.tb00606.x
Verma D, Satyanarayana T (2011) An improved protocol for DNA extraction from alkaline soil and sediment samples for constructing metagenomic libraries. Appl Biochem Biotech 165(2):454–464. https://doi.org/10.1007/s12010-011-9264-5
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267. https://doi.org/10.1128/AEM.00062-07
Wang X, Jiang Z, Ren H (2008) Distribution of wet heartwood in stems of Populus xiaohei from a spacing trial. Scand J For Res 23:38–45. https://doi.org/10.1080/02827580701763706
Wang X, Wang Z, Jiang P, He Y, Mu Y, Lv X, Zhuang L (2018) Bacterial diversity and community structure in the rhizosphere of four Ferula species. Sci Rep 8:5345. https://doi.org/10.1038/s41598-018-22802-y
Ward JC, Pong WY (1980) Wetwood in trees—overview of a timber resource problem. (Gen. Tech. Rep. PNW-112). Pacific Northwest Forest and Range Experiment Station, USDA Forest Service
Ward JC, Zeikus JG (1980) Bacteriological, chemical and physical properties of wetwood in living trees. Mitt Bundesforsch Forst Holzwirtsch 131:133–168
Warshaw JE, Leschine SB, Canale-parola E (1985) Anaerobic cellulolytic bacteria from wetwood of living trees. Appl Environ Microbiol 50(4):807–811
Wasserfallen A, Nölling J, Pfister P, Reeve J, Conway de Macario E (2000) Phylogenetic analysis of 18 thermophilic Methanobacterium isolates supports the proposals to create a new genus, Methanothermobacter gen. nov., and to reclassify several isolates in three species, Methanothermobacter thermautotrophicus comb. nov. Methanothermobacter wolfeii comb. nov., and Methanothermobacter marburgensis sp. nov. Int J Syst Evol Microbiol 50:43–53. https://doi.org/10.1099/00207713-50-1-43
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogentics. PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Wu Z, Zhang M, Wang L, Tu Y, Zhang J, Xie G, Zou W, Li F, Guo K, Li Q, Gao C, Peng L (2013) Biomass digestibility is predominantly affected by three factors of wall polymer features distinctive in wheat accessions and rice mutants. Biotechnol Biofuels 6:183. https://doi.org/10.1186/1754-6834-6-183
Xiang YM (1983) Study on method for determining the pH of wood. Sci Silvae Sinicae 19(4):430–433
Zeikus JG, Henning DL (1975) Methanobacterium arbophilicum sp.nov. An obligate anaerobe isolated from wetwood of living trees. Antonie Van Leeuwenhoek 41(4):543–552. https://doi.org/10.1007/BF02565096
Zeikus JG, Ward JC (1974) Methane formation in living trees: a microbial origin. Science 184(4142):1181–1183. https://doi.org/10.1126/science.184.4142.1181
Zheng HY, Dai YS, Xie CX, Guo TB, Wang ZY (2006) Studies on the cause of formation of poplar red core wood. J Nanjing For Univ 30(2):82–84
Acknowledgements
This study was funded by the National Natural Science Foundation of China (31570665), Hubei Provincial Major Technological Innovation Foundation of China (2016ABA111), Hubei Provincial Natural Science Foundation of China (2014CFB933 and 2009CDA096), and Fundamental Research Funds for the Central Universities of China (201610504037).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Feau.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, X., Hu, X., Peng, Y. et al. Amplicon sequencing reveals different microbial communities in living poplar wetwood and sapwood. Trees 33, 851–865 (2019). https://doi.org/10.1007/s00468-019-01822-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-019-01822-9