Abstract
Background
Gastric cancer is a common kind of malignancies, with yearly occurrences exceeding one million worldwide in 2017. Typically, ulcerous and cancerous tissues develop abnormal morphologies through courses of progression. Endoscopy is a routinely adopted means for examination of gastrointestinal tract for malignancy. Early and timely detection of malignancy closely correlate with good prognosis. Repeated presentation of similar frames from gastrointestinal tract endoscopy often weakens attention for practitioners to result in true patients missed out to incur higher medical cost and unnecessary morbidity. Highly needed is an automatic means for spotting visual abnormality and prompts for attention for medical staff for more thorough examination.
Methods
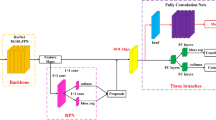
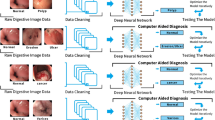
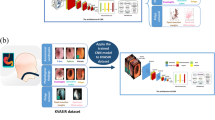
We conduct classification of benign ulcer and cancer for gastrointestinal endoscopic color images using deep neural network and transfer-learning approach. Using clinical data gathered from Gil Hospital, we built a dataset comprised of 200 normal, 367 cancer, and 220 ulcer cases, and applied the inception, ResNet, and VGGNet models pretrained on ImageNet. Three classes were defined—normal, benign ulcer, and cancer, and three separate binary classifiers were built—those for normal vs cancer, normal vs ulcer, and cancer vs ulcer for the corresponding classification tasks. For each task, considering inherent randomness entailed in the deep learning process, we performed data partitioning and model building experiments 100 times and averaged the performance values.
Results
Areas under curves of respective receiver operating characteristics were 0.95, 0.97, and 0.85 for the three classifiers. The ResNet showed the highest level of performance. The cases involving normal, i.e., normal vs ulcer and normal vs cancer resulted in accuracies above 90%. The case of ulcer vs cancer classification resulted in a lower accuracy of 77.1%, possibly due to smaller difference in appearance than those cases involving normal.
Conclusions
The overall level of performance of the proposed method was very promising to encourage applications in clinical environments. Automatic classification using deep learning technique as proposed can be used to complement manual inspection efforts for practitioners to minimize dangers of missed out positives resulting from repetitive sequence of endoscopic frames and weakening attentions.






Similar content being viewed by others
References
World Cancer Report 2014 (2014) World Health Organization
http://globocan.iarc.fr. Accessed 23 Jan 2019
Watanabe K, Nagata N, Shimbo T, Nakashima R, Furuhata E, Sakurai T, Akazawa N, Yokoi C, Kobayakawa M, Akiyama J, Mizokami M, Uemura N (2013) Accuracy of endoscopic diagnosis of Helicobacter pylori infection according to level of endoscopic experience and the effect of training. BMC Gastroenterol 13:128
Menon S, Trudgill N (2014) How commonly is upper gastrointestinal cancer missed at endoscopy? A meta-analysis. Endosc Int Open 2(2):E46–E50
Almadi MA, Sewitch M, Barkun AN (2015) Adenoma detection rates decline with increasing procedural hours in an endoscopist’s workload. Can J Gastroenterol Hepatol 29(6):304–308
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Miyaki R, Yoshida S, Tanaka S, Kominami Y, Sanomura Y, Matsuo T, Oka S, Raytchev B, Tamaki T, Koide T, Kaneda K, Yoshihara M, Chayama K (2015) A computer system to be used with laser-based endoscopy for quantitative diagnosis of early gastric cancer. J Clin Gastroenterol 49(2):108–115
Zhang X, Hu W, Chen F, Liu J, Yang Y, Wang L, Duan H, Si J (2017) Gastric precancerous diseases classification using CNN with a concise model. PLoS ONE 12(9):e0185508
Zhu R, Zhang R, Xue D (2015) Gastric precancerous diseases classification using CNN with a concise model. In: 2015 8th International Congress on Image and Signal Processing (CISP) 14–16 Oct
Billah M, Waheed S, Rahman MM (2017) An automatic gastrointestinal polyp detection system in video endoscopy using fusion of color wavelet and convolutional neural network features. Int J Biomed Imaging. https://doi.org/10.1155/2017/9545920
Byrne MF, Chapados N, Soudan F, Oertel C, Linares PM, Kelly R, Iqbal N, Chandelier F, Rex DK (2017) Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut 68:94–100
Tajbakhsh N (2015) Automatic polyp detection in colonoscopy videos using an ensemble of convolutional neural networks. In: IEEE 12th International Symposium on Biomedical Imaging (ISBI)
Li B, Meng MQ (2009) Computer-aided detection of bleeding regions for capsule endoscopy images. IEEE Trans Biomed Eng 56(4):1032–1039
Tamai N, Saito Y, Sakamoto T, Nakajima T, Matsuda T, Sumiyama K, Tajiri H, Koyama R, Kido S (2017) Effectiveness of computer-aided diagnosis of colorectal lesions using novel software for magnifying narrow-band imaging: a pilot study. Endosc Int Open 5(8):E690–E694
Clements LM, Kockelman KM (2017) Economic effects of automated vehicles. J Transp Res Board 2606:2606–2614
Silver D, Huang A, Maddison CJ, Guez A, Sifre L, Driessche G, Schrittwieser J (2016) Mastering the game of Go with deep neural networks and tree search. Nature 529:484–489
Gerhardus D (2003) Robot-assisted surgery: the future is here. J Healthc Manag 48(4):242–251
Maroulis DE, Savelonas MA, Karkanis SA, Iakovidis DK, Dimitropoulos N (2005) Computer-aided thyroid nodule detection in ultrasound images. Proceedings of the 18th IEEE symposium on computer-based medical systems (CBMS’05)
Pizer SM, Amburn EP, Austin JD, Cromartie R, Geselowitz A, Greer T, Romeny B, Zimmerman JB, Zuiderveld K (1987) Adaptive histogram equalization and its variations. Comput Vis Graph Image Process 39(3):355–368
Zuiderveld K (1994) Contrast limited adaptive histogram equalization. Graphics gems IV. Academic Press Professional, Inc., San Diego, pp 474–485
Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L (2009) ImageNet: a large-scale hierarchical image database. In: Proceedings of IEEE conference on computer vision and pattern recognition, pp. 248–255
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. NIPS 1:1097–1105
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Vanhoucke V, Rabinovich A (2015) Going deeper with convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 1–9
He K, Zhang C, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 770–778
Canziani A, Paszke A, Culurciello E (2016) An analysis of deep neural network models for practical applications. ArXiv:1605.07678
Dauphin GMY, Glorot X, Rifai S, Bengio Y, Goodfellow I, Lavoie E, Muller X, Desjardins G, Warde-Farley D, Vincent P, Courville A, Bergstra J (2012) Unsupervised and transfer learning challenge: a deep learning approach. JMLR 27:97–110
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542(7639):115–118
Pogorelov K, Randel KR, Griwodz C, Eskeland SL, de Lange T, Johansen D, Spampinato C, Nguyen DTD, Lux M, Schmidt PT, Riegler M, Halvorsen P (2017) Kvasir: a multi-class image dataset for computer aided gastrointestinal disease detection. In: Proceedings of the 8th ACM on Multimedia Systems Conference, Pages 164–169
Acknowledgements
This work was supported by Institute for Information & communications Technology Promotion (IITP) grant funded by the Korea government (MSIT) (2018-2-00861, Intelligent SW Technology Development for Medical Data Analysis) and the Gachon University Gil Medical Center (Grant No: 2018-5283). Authors Kim KG and Chung JW equally contributed to this work.
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
Authors Jang Hyung Lee, Young Jae Kim, Yoon Woo Kim, Sungjin Park, Yoon Yi Choi, Yoon Jae Kim, Dong Kyun Park, Kwang Gi Kim, and Jun-Won Chung have no financial arrangement or affiliation with any product or services used or discussed in this paper, nor any potential bias against another product or service. The authors declare that there are no conflicts of interest regarding the publication of this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J.H., Kim, Y.J., Kim, Y.W. et al. Spotting malignancies from gastric endoscopic images using deep learning. Surg Endosc 33, 3790–3797 (2019). https://doi.org/10.1007/s00464-019-06677-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00464-019-06677-2