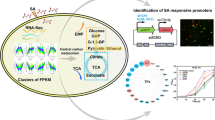
Abstract
In this work, we expanded and updated a genome-scale metabolic model of Streptomyces clavuligerus. The model includes 1021 genes and 1494 biochemical reactions; genome-reaction information was curated and new features related to clavam metabolism and to the biomass synthesis equation were incorporated. The model was validated using experimental data from the literature and simulations were performed to predict cellular growth and clavulanic acid biosynthesis. Flux balance analysis (FBA) showed that limiting concentrations of phosphate and an excess of ammonia accumulation are unfavorable for growth and clavulanic acid biosynthesis. The evaluation of different objective functions for FBA showed that maximization of ATP yields the best predictions for cellular behavior in continuous cultures, while the maximization of growth rate provides better predictions for batch cultures. Through gene essentiality analysis, 130 essential genes were found using a limited in silico media, while 100 essential genes were identified in amino acid-supplemented media. Finally, a strain design was carried out to identify candidate genes to be overexpressed or knocked out so as to maximize antibiotic biosynthesis. Interestingly, potential metabolic engineering targets, identified in this study, have not been tested experimentally.




Similar content being viewed by others
References
Paradkar A (2013) Clavulanic acid production by Streptomyces clavuligerus: biogenesis, regulation and strain improvement. J Antibiot 66:411–420
Jensen SE (2012) Biosynthesis of clavam metabolites. J Ind Microbiol Biotechnol 39:1407–1419
Ozcengiz G, Demain AL (2013) Recent advances in the biosynthesis of penicillins, cephalosporins and clavams and its regulation. Biotechnol Adv 31:287–311
Zelyas NJ, Cai H, Kwong T, Jensen SE (2008) Alanylclavam biosynthetic genes are clustered together with one group of clavulanic acid biosynthetic genes in Streptomyces clavuligerus. J Bacteriol 190:7957–7965
Medema MH, Alam MT, Heijne WH, van den Berg MA, Muller U, Trefzer A, Bovenberg RA, Breitling R, Takano E (2011) Genome-wide gene expression changes in an industrial clavulanic acid overproduction strain of Streptomyces clavuligerus. Microb Biotechnol 4:300–305
Borodina I, Krabben P, Nielsen J (2005) Genome-scale analysis of Streptomyces coelicolor A3(2) metabolism. Genome Res 15:820–829
D’Huys PJ, Lule I, Vercammen D, Anné J, Van Impe JF, Bernaerts K (2012) Genome-scale metabolic flux analysis of Streptomyces lividans growing on a complex medium. J Biotechnol 161:1–13
Medema MH, Trefzer A, Kovalchuk A, van den Berg M, Müller U, Heijne W, Wu L, Alam MT, Ronning CM, Nierman WC, Bovenberg RA (2010) The sequence of a 1.8-mb bacterial linear plasmid reveals a rich evolutionary reservoir of secondary metabolic pathways. Genome Biol Evol 2:212–224
Magrane M, Consortium U (2011) UniProt Knowledgebase: a hub of integrated protein data. Database (Oxford) 2011:bar009
Devoid S, Overbeek R, DeJongh M, Vonstein V, Best AA, Henry C (2013) Automated genome annotation and metabolic model reconstruction in the SEED and Model SEED. In: Systems metabolic engineering: methods and protocols, pp 17–45
Schellenberger J, Que R, Fleming RM, Thiele I, Orth JD, Feist AM, Zielinski DC, Bordbar A, Lewis NE, Rahmanian S, Kang J (2011) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v2.0. Nat Protoc 6:1290–1307
Karp PD, Ouzounis CA, Moore-Kochlacs C, Goldovsky L, Kaipa P, Ahrén D, Tsoka S, Darzentas N, Kunin V, López-Bigas N (2005) Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res 33:6083–6089
Wattam AR, Abraham D, Dalay O, Disz TL, Driscoll T, Gabbard JL, Gillespie JJ, Gough R, Hix D, Kenyon R, Machi D (2014) PATRIC, the bacterial bioinformatics database and analysis resource. Nucl Acids Res 42:D581-D591
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Gevorgyan A, Bushell ME, Avignone-Rossa C, Kierzek AM (2011) SurreyFBA: a command line tool and graphics user interface for constraint-based modeling of genome-scale metabolic reaction networks. Bioinformatics 27:433–434
Yousofshahi M, Ullah E, Stern R, Hassoun S (2013) MC3: a steady-state model and constraint consistency checker for biochemical networks. BMC Syst Biol 7:1–8
Orth JD, Thiele I, Palsson B (2010) What is flux balance analysis? Nat Biotechnol 28:245–248
Durot M, Bourguignon PY, Schachter V (2009) Genome-scale models of bacterial metabolism: reconstruction and applications. FEMS Microbiol Rev 33:164–190
Bushell ME, Kirk S, Zhao HJ, Avignone-Rossa CA (2006) Manipulation of the physiology of clavulanic acid biosynthesis with the aid of metabolic flux analysis. Enzym Microb Technol 39:149–157
Edwards JS, Palsson B (2000) The Escherichia coli MG1655 in silico metabolic genotype: its definition, characteristics, and capabilities. Proc Natl Acad Sci USA 97:5528–5533
Schuetz R, Kuepfer L, Sauer U (2007) Systematic evaluation of objective functions for predicting intracellular fluxes in Escherichia coli. Mol Syst Biol 3:119
Blazeck J, Alper (2010) Systems metabolic engineering: genome-scale models and beyond. Biotechnol J 5:647–659
Stanford NJ, Millard P, Swainston N (2015) RobOKoD: microbial strain design for (over)production of target compounds. Front Cell Dev Biol 3:17
Kim M, Sang Yi J, Kim J, Kim JN, Kim MW, Kim BG (2014) Reconstruction of a high-quality metabolic model enables the identification of gene overexpression targets for enhanced antibiotic production in Streptomyces coelicolor A3(2). Biotechnol J 9:1185–1194
Hucka M, Finney A, Sauro HM, Bolouri H, Doyle JC, Kitano H (2003) The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics 19:524–531
Hutter F, Hoos HH, Leyton-Brown K (2010) Automated configuration of mixed integer programming solvers. In: International conference on integration of artificial intelligence (AI) and operations research (OR) techniques in constraint programming. Springer, Berlin, pp 186–202
Tahlan K, Anders C, Wong A, Mosher RH, Beatty PH, Brumlik MJ, Griffin A, Hughes C, Griffin J, Barton B, Jensen SE (2007) 5S clavam biosynthetic genes are located in both the clavam and paralog gene clusters in Streptomyces clavuligerus. Chem Biol 14:131–142
Mercier C (2006) A Genome-scale investigation of clavulanic acid biosynthesis by Streptomyces clavuligerus in batch and chemostat cultures using transcriptomic and fluxomic analysis. PhD Thesis. University of Surrey, UK
Aharonowitz Y, Demain AL (1978) Carbon catabolite of cephalosphorin production in Streptomyces clavuligerus. Antimicrob Agents Chemother 14:159–164
Müller JC, Toome V, Pruess DL, Blount JF, Weigele M (1983) Ro 22-5417, a new clavam antibiotic from Streptomyces clavuligerus. III. Absolute stereochemistry. J Antibiot(Tokio) 36:208–212
Romero J, Liras P, Martín JF (1986) Utilization of ornithine and arginine as specific precursors of clavulanic acid. Appl Environ Microbiol 52:892–897
Romero J, Liras P, Martín JF (1984) Dissociation of cephamycin and clavulanic acid biosynthesis in Streptomyces clavuligerus. Appl Microbiol Biotechnol 20:318–325
Pérez-Redondo R, Santamarta I, Bovenberg R, Martín JF, Liras P (2010) The enigmatic lack of glucose utilization in Streptomyces clavuligerus is due to inefficient expression of the glucose permease gene. Microbiology 156:1527–1537
Lebrihi A, Germain P, Lefebvre G (1987) Phosphate repression of cephamycin and clavulanic acid production by Streptomyces clavuligerus. Appl Microbiol Biotechnol 26:130–135
Aharonowitz Y, Demain AL (1979) Nitrogen nutrition and regulation of cephalosporin production in Streptomyces clavuligerus. Can J Microbiol 25:61–67
Khannapho C, Zhao H, Bonde BK, Kierzek AM, Avignone-Rossa CA, Bushell ME (2008) Selection of objective function in genome scale flux balance analysis for process feed development in antibiotic production. Metab Eng 10:227–233
Sánchez C, Quintero JC, Ochoa S (2015) Flux balance analysis in the production of clavulanic acid by Streptomyces clavuligerus. Biotechnol Prog 31:1226–1236
García Sánchez CE, Torres Sáez RG (2014) Comparison and analysis of objective functions in flux balance analysis. Biotechnol Prog 30:985–991
Persson BC, Gustafsson C, Berg DE, Björk GR (1992) The gene for a tRNA modifying enzyme, m5U54-methyltransferase, is essential for viability in Escherichia coli. Proc Natl Acad Sci USA 89:3995–3998
Redshaw PA, McCann PA, Pentella MA, Pogell BM (1979) Simultaneous loss of multiple differentiated functions in aerial mycelium-negative isolates of Streptomycetes. J Bacteriol 137:891–899
Wu G, Culley DE, Zhang W (2005) Predicted highly expressed genes in the genomes of Streptomyces coelicolor and Streptomyces avermitilis and the implications for their metabolism. Microbiology 151:2175–2187
Song JY, Jensen SE, Lee KJ (2010) Clavulanic acid biosynthesis and genetic manipulation for its overproduction. Appl Microbiol Biotechnol 88:659–669
Jnawali HN, Yoo JC, Sohng JK (2011) Improvement of clavulanic acid production in Streptomyces clavuligerus by genetic manipulation of structural biosynthesis genes. Biotechnol Lett 33:1221–1226
Panichkin VB, Livshits VA, Biryukova IV, Mashko SV (2016) Metabolic engineering of Escherichia coli for l-tryptophan production. Appl Biochem Microbiol 52:783–809
Acknowledgements
The authors thank Professor Marnix H. Medema and Mohammad Tauqeer Alam for providing the template for the Streptomyces clavuligerus model, and Professor Andrzej Kierzek for advice on the SurreyFBA platform [15]. This work was supported by Departamento Administrativo de Ciencia, Tecnología e Innovación—COLCIENCIAS-Colombia (Grant no. 111566945929). L. Toro and L. Pinilla thank COLCIENCIAS-Colombia for scholarships. C. Avignone-Rossa was supported by Grant BB/L02683X/1 from the Biotechnology and Biological Sciences Research Council (BBSRC, United Kingdom).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
. Model content and Simulation Results. Spreadsheet 1: Reactions included in the S. clavuligerus Sclav_iLT1021 model. Spreadsheet 2: Metabolites included the in S. clavuligerus Sclav_iLT1021 model. Spreadsheet 3: Biomass composition. Spreadsheet 4: MC3 output (gapfilling). Spreadsheet 5: Gene and reaction essentiality for complex media (CPX) and defined media (GAPI). Spreadsheet 6: RoBoKoD analysis, OE: Overexpression ranking, KO: Knockout ranking. Spreadsheet 7: Published experimental data used for testing the different objective functions. (XLSX 382 KB)
Supplementary material 2
: Metabolic model in SBML format (XML) (XML 2336 KB)
Supplementary figure S1
: Validation of the Sclav_iLT1021 model using the experimental data from the literature a. Chemostat data from the literature [19]. Color Code: blue and green: specific glycerol and oxygen uptake rate, respectively; black and yellow: specific CO2 and clavulanic acid secretion rate, respectively. a. Model predictions of the specific growth rate and experimental data from the literature [19]. (TIFF 8790 KB)
Supplementary figure S2
: Scatter plot for the correlation of the different objective functions evaluated. a Chemostat, P-limited media [19]; b Chemostat, P-limited media (additional constraints) [28]; c Batch, medium supplemented media with amino acids [31]. (TIFF 2164 KB)
Rights and permissions
About this article
Cite this article
Toro, L., Pinilla, L., Avignone-Rossa, C. et al. An enhanced genome-scale metabolic reconstruction of Streptomyces clavuligerus identifies novel strain improvement strategies. Bioprocess Biosyst Eng 41, 657–669 (2018). https://doi.org/10.1007/s00449-018-1900-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-018-1900-9