Abstract
Purpose
Breast cancer is the leading cause of cancer death in females. Histone modifications have been shown to have an influence on the gene expression. This study focusses on the histone modifications H3K9ac and H3K4me3 in breast cancer and their impact on survival
Methods
H3K4me3 and H3K9ac expression was immunohistochemically examined in 235 tissue samples.
Results
Positive estrogen receptor status was correlated with a higher IRS of the nuclear (p = 0.033), and of the cytoplasmic H3K4me3 staining (p = 0.009). H3K9ac intensity was associated to the Her2 status (p = 0.045) and to poor prognosis in cells with positive Ki67 status (p = 0.013). A high intensity of nuclear H3K4me3 staining was found to be correlated with a lower 10-year-survival (p = 0.026) and with lower breast cancer-specific survival (p = 0.004). High percentage score (> 190) of H3K9ac expression was correlated to worse breast cancer-specific survival (p = 0.005). Shorter progression-free survival was found in patients with nuclear (p = 0.013) and cytoplasmic H3K4me3expression (p = 0.024) and H3K9ac expression (p = 0.023).
Conclusion
This analysis provides new evidence of histone modifications in breast cancer. High H3K4me3 and H3K9ac expression was correlated with survival rates. Further investigation of histone modifications in breast cancer could lead to a more profound understanding of the molecular mechanisms of cancer development and could result in new therapeutic strategies.
Similar content being viewed by others
Introduction
Breast cancer is the most frequently diagnosed cancer and the leading cause of cancer death in females worldwide, accounting for 25% (1.7 million) of the total new cancer cases and 15% (521.900) of the total cancer-related deaths in 2012 (Torre et al. 2017). Breast cancer remains a significant threat to women all over the world, even though the breast cancer death rates have decreased by 40% between 1975 and 2017 (DeSantis et al. 2019).
Gene expression profiling has had an important impact on the understanding of breast cancer (Bell et al. 2017). For example, the analysis of certain biomarkers such as hormone receptor status, Her2 status and expression of Ki67 has led to the characterization of molecular subtypes of breast cancer that have shown significant differences in terms of their incidence, risk factors, prognosis and sensitivity to treatment (Prat et al. 2015).
Epigenetic alterations, such as DNA methylation and posttranslational modification of histones have been shown to have a considerable influence on the gene expression (Wu et al. 2015). Histones are the central component of the nucleosomes’ subunit. They form an octamer containing the four core histone proteins (H3, H4, H2A, H2B) around which is wrapped a 147-base-pair segment of the DNA (Audia and Campbell 2016). The histones’ N-terminal tails extend from the double-strand DNA and are subject to posttranslational modifications, which include acetylation, methylation, phosphorylation, ADP-ribosylation, glycosylation, sumoylation and ubiquitylation (Zhang et al. 2016). Histone acetylation is primarily associated with gene activation, whereas methylation, depending on its position and state, can either be associated with repression or activation (Wang et al. 2008).
Previous studies have analyzed the impact of specific posttranslational modifications on the gene expression (Lawrence et al. 2016). For example, H3K4 methylation has been intensely studied regarding the enzymes and molecular factors required for methylation (Shilatifard 2008). An association between high H3K4me3 expression and poor prognosis was found in patients with hepatocellular carcinoma and cervical carcinoma (Beyer et al. 2017; He et al. 2012). In cervical cancer, the same observation has been made for H3K9ac (Beyer et al. 2017).
As a thorough investigation regarding the influence of histone modifications on the prognosis of breast cancer patients was lacking, an expression analysis of histone H3 trimethyl K4 (H3K4me3) and histone H3 acetyl K9 (H3K9ac) was performed in this study. 235 tissue samples were examined by immunohistochemical methods and assessed by a semi-quantitative score.
Materials and methods
Patients and specimens
A panel of 235 tissue samples from patients who underwent surgery at the Ludwig-Maximilians-University, Munich between 1998 and 2000 due to malignant breast cancer was used. The mean patients’ age was 58.2 ± 13.3 years. 43.8% of the breast cancer cases were classified as Luminal A, 31.5% as Luminal B and Her2 negative, 6.8% as Luminal B and Her2 positive, 3.0% as Her2 positive and hormone receptor negative; 13.2% were triple negative carcinomas. 65.1% of the patients were diagnosed with a tumor size smaller than 2 cm, 28.1% had a tumor between 2 and 5 cm. At least one regional lymph node was affected in 39.6% of the cases. 7.2% were ranked as low grade, 36.6% as intermediate grade and 22.6% as high-grade tumors. In 33.6%, the grading-classification was missing. DCIS and LCIS fractions were observed in 51.1% of the cases. For more patients’ characteristics’ see Table 1.
The endpoints were defined as following: OS = overall survival, period of time from the date of surgery until the date of death or date of last follow-up; DSS = disease-specific survival, period of time from the date of surgery until the breast cancer-dependent death; PFS = progression-free survival, period of time until local recurrence or metastasis were diagnosed; local PFS = period of time until a local recurrence was diagnosed; DDFS = distant disease-free survival, period of time until metastasis is diagnosed.
Ethics approval
The tissue samples were originally collected for histopathological diagnostics. They were no longer used for clinical tests when being selected for this study. Patient data was anonymized and the authors were blinded for the patients’ information as well as for survival time during the analysis. The study was approved by the local ethics committee of the Ludwig-Maximilians-University of Munich (Reference No. 048-08; 2008) and was performed according to the Declaration of Helsinki.
Immunohistochemistry
The formalin-fixated and paraffin-embedded tissues were first dewaxed in xylol. After rinsing the tissue in 100% ethanol, the endogenous peroxidase was inactivated in 3% H2O2 in methanol and the samples were rehydrated in a descending alcohol series. To unmask the antigen, the samples were heated up to 100 °C in a dilution with citrate buffer for 5 min. After washing the samples in distilled water and PBS-buffer, a blocking solution was applied to prevent unspecific staining due to binding of the antibodies to electrostatic charges in the tissue. The samples were then incubated at 4 °C with the primary antibody for 16 h (see Table 2). A Purified solution of Antibodies bought from the producer was used. The specificity of the antibodies was already tested by CHiP Sequence by other authors and the producer (Abcam; Lima-Fernandes et al. 2019). After intensifying the staining with a Post-Block Solution, the HRP polymer was applied. The excess HRP Polymer was removed and the binding of the antibody was made visible through an enzymatic reaction using diaminobenzidine (DAB). A counterstaining was performed in haemalaun (2 min), followed by dehydration in an ascending alcohol series and covering of the samples. Placenta tissues were used as positive and negative controls for each staining of H3K4me3 and H3K9ac (Fig. 1b and d). The results of the staining were analyzed using two different scores. The immunoreactive score (IRS-Score) multiplies the intensity of the staining (0 = not stained, 1 = low intensity, 2 = moderate intensity, 3 = high intensity) with the percentage of stained tumor cells (0 = 0%, 1 = 1–10%, 2 = 11–50%, 3 = 51–80%, 4 = > 80%). The result is a value between 0 and 12. The %Score multiplies the staining intensity (0 = not stained, 1 = low intensity, 2 = moderate intensity, 3 = high intensity) with the percentage of stained tumor cells, allowing a finer differentiation of the samples regarding the percentage of stained cells (range: 0–300%).
Statistics
IBM SPSS Statistics version 25 (Armonk, NY, USA) was used. Spearman’s-rank correlation coefficient was employed to calculate bivariate correlations. To compare independent groups non-parametric tests (NPAR: Kruskal–Wallis test, Mann–Whitney U test) were used. Survival times were shown by Kaplan–Meier estimates and calculated by log-rank-test, thresholds were defined experimentally. The p value of the tests had to be < 0.05 to be statistically significant.
Results
H3K4me3 staining in breast cancer
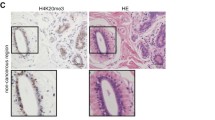
71.4% of the samples showed a nuclear staining with a median immunoreactive score of 2 and a median percentage score of 40 (Fig. 1a). 26.0% displayed no nuclear staining. 20.4% of the samples had low staining (IRS ≤ 2), while 44.3% showed an enhanced staining (IRS > 2) and, therefore, high expression of H3K4me3 in the nucleus. 9.3% of the samples could not be analyzed. An additional staining of the cytoplasm was found in 35.1% of the samples. Placenta tissue was used as positive control (Fig. 1b).
The correlations between the H3K4me3 expression and several important clinical parameters, such as the estrogen receptor status and the Her2 status, as well as the H3K9ac expression, were analyzed by applying Spearman’s rank correlation coefficient (see Table 3).
An association between H3K4me3 and the tumor cells’ estrogen receptor status was observed: Positive estrogen receptor status was correlated with a higher IRS of the nuclear staining (p = 0.033, Rho = 0.147; Fig. 2a–c): the median of nuclear H3K4me3 expression in estrogen positive cells was 3, compared to 2 in estrogen negative cells. ER expression was also associated to a higher intensity (p = 0.006, Rho = 0.186) and IRS (p = 0.009, Rho = 0.179) of the cytoplasmic staining (Fig. 2d–f; see Table 3). Although the difference is significant, the correlation is weak and cannot be seen in the boxplot.
Correlation of H3K4me3 Nuc and Cyt and ER. a Boxplot: Difference of H3K4me3 Nuc expression in regard to ER status; b H3K4me3Nuc staining (IRS 9) and ER positive; c H3K4me3Nuc expression (IRS 2) and ER negative; d Boxplot: H3K4me3 Cyt expression in regard to ER status; e H3K4me3Cyt staining (IRS 8) and ER positive; f H3K4me3Cyt expression (IRS 3) and ER negative
No significant correlation was found regarding the histological subtype, T-stage, N-stage, grading, Ki67 status, PR, Her2 status and clinical subtype (Luminal A, Luminal B, basal like triple negative Her2 positive typ luminal, Her2 positive typ non-luminal) (see Table 3).
A correlation between the H3K9ac staining and the nuclear H3K4me3 staining (p = 0.000; Rho = 0.633), as well as the cytoplasmic H3K4me3 staining (p = 0.000; Rho = 0.448) was found, showing that intense H3K4me3 staining is associated with increased H3K9ac expression (see Table 4).
H3K9ac staining in breast cancer
A total of 72.4% of the samples showed a nuclear staining with a median IRS of 3 and a median percentage score of 60 (Fig. 1c). Unlike the H3K4me3 staining, no additional staining of the cytoplasm was found. 28.5% of the samples had low staining (IRS ≤ 3), while 40.9% showed an enhanced staining (IRS > 3) and 26.4% did not show any staining. 4.2% of the samples could not be analyzed. Placenta tissue was used as positive control (Fig. 1d).
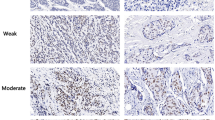
As mentioned above, a correlation between the H3K4me3 staining and the H3K9ac staining was found (see Table 4). Furthermore, a high H3K9ac staining intensity was shown to be correlated with a positive Her2 status of the tumor cells (p = 0.045, ρ = 0.134; Table 3; Fig. 3), but the correlation was weak.
The categorization of the molecular subtype is an important part of the breast cancer diagnosis, as it influences the patient’s treatment and allows to give an approximate prognosis. To further refine the prognosis in specific subcategories, more markers are useful. The expression of H3K9ac was identified as a potential marker. Regarding the Ki67 status, H3K9ac expression was found not to be directly associated to Ki67 expression. But in samples with positive Ki67 status (defined as more than 14% of the tumor cells being positive for Ki67), H3K9ac expression was associated with poor prognosis (p = 0.013; Fig. 4).
No significant correlation was found regarding the T-stage, N-stage, estrogen receptor status, grading, the PR and the clinical subtype.
Role of H3K4me3 and H3K9ac on survival
A high intensity of nuclear H3K4me3 staining (intensity = 3) was found to be correlated with a lower 10-year survival in breast cancer patients (p = 0.026; Fig. 5a). Taking into consideration only the patients that died due to breast cancer, we found out, that patients had to have a %Score > 110 to show a significantly better breast cancer-specific survival (p = 0.004; Fig. 5b). The cytoplasmic expression of H3K4me3 had no visible effect on the survival of the patients.
The examination of the role of H3K9ac showed no significant effect on the overall survival. Regarding the breast cancer-specific survival, patients with a high %Score had a worse prognosis (p = 0.005; Fig. 5c). The threshold needed for significant results was %Score > 190.
Role of H3K4me3 and H3K9ac on progression-free survival
In addition to the impact on the patient’s general survival, nuclear H3K4me3 expression was also correlated with the progression-0free survival. The distant disease-free survival, as well as the local disease-free survival, was decreased in patients with %Score > 150 (p = 0.005 and p = 0.049; Fig. 6a and b). Combining these two parameters, a significantly shorter general progression-free survival was found in these patients (p = 0.017; Fig. 6c).
Role of H3K4me3 and H3K9ac for progress-free survival. Nuclear H3K4me3-expression for distant (a) and local (b) recurrence-free survival. c recurrence-free survival in patients with nuclear H3K4me3 expression; recurrence (d) and distant disease free (e) survival in association to cytoplasmic H3K4me3 expression; f recurrence-free survival with H3K9ac expression
While the cytoplasmic expression of H3K4me3 seemed to have no impact on the overall survival, it was correlated with a shorter progression-free survival in patients with a %Score ≥ 60 (p = 0.034; Fig. 6d). These patients also showed a shorter distant disease-free survival (p = 0.024; Fig. 6e). Patients with %Score > 110 also showed a shorter local disease-free survival.
A shorter recurrence-free survival was also found in patients with a high nuclear expression of H3K9ac (%Score > 225, p = 0.023; Fig. 6f).
Discussion
This study showed that specific histone modifications are important in breast cancer patients. H3K4me3 expression was correlated with positive estrogen receptor status, while H3K9ac staining was correlated with positive Her2 receptor status. Although results were highly significant, the correlations themselves were weak.
High H3K4me3 and H3K9ac expression were correlated with shorter breast cancer-specific survival as well as shorter progression-free survival. Overall survival was decreased in patients with high nuclear H3K4me3 staining.
Histone modifications have been of great scientific interest in the past years. After synthesis of histones, posttranslational modifications like methylation or acetylation are performed. These can take place in the nucleus or in the cytoplasm (Annunziato and Hansen 2000; Wu et al. 2012). Posttranslational alterations, which can mainly be found at the loose N-termini, but also within the global domain of histones, have been shown to regulate the structure, accessibility and replication of DNA and play an important role in fundamental cellular mechanisms in the cell cycle (Zhang et al. 2016). Aberrant histone modifications have been linked to the pathogenesis of several diseases, including inflammatory diseases and cancer, as they cause a shift in the gene expression and the overall metabolic state of the cell (Shanmugam et al. 2018).
Previous studies have concentrated on the impact of specific histone modifications on the cell: it has been shown that while histone acetylation is mainly associated with gene activation, methylation can be associated with either repression or activation, depending on its position and level (mono/di/trimethylated) (Kimura 2013; Lee et al. 1993; Vakoc et al. 2006). In this study, we analyzed the impact of H3K4me3 and H3K9ac on the survival of breast cancer patients, as these modifications have been described to be associated with active chromatin (Ruthenburg et al. 2007).
H3K4 methylation is a modification occurring at the fourth lysine residue from the N-terminus of Histone H3. It can be mono-, di- and trimethylated, which adds to the complexity of the analysis of its impact on the genome (Takahashi and Shilatifard 2010). H3K4me3 is generally associated with transcriptional activation and has been proposed as a predictive factor of poor prognosis in several types of cancer, such as liver and cervical cancer (Li et al. 2018). Poorer prognosis has also been described for patients with cervical cancer with a high expression of H3K9ac (Beyer et al. 2017).
By its neutralizing acetyl-group, H3K9ac leads to a de-condensation of the DNA structures and to an activation of transcription (Lee et al. 1993). The effect of an acetylated H3 at position 9 depends on the tumor entity: high H3K9ac levels seem to be associated with a poor prognosis in cervical cancer (Beyer et al. 2017), while patients with glioma have better prognosis with high H3K9ac levels (Liu et al. 2010).
There are much more histone modifications beside H3K4me3 and H3K9ac, which are associated to activation of transcription. For repressive modifications like H3K20me3 and H3K9me3, it was shown, that their levels are elevated in breast cancer cells (Leszinski et al. 2012). Another modification with repressive effect on gene transcription, H3K27me3, Healy et al. could show that it was associated to low grading and inversely to Her-2-neu status (Healey et al. 2014). These results fit to our observance that activating modifications have a positive correlation to the Her2neu status.
In the present study, we showed that H3K4 tri-methylation and H3K4 acetylation are negative prognosticators for breast cancer patients. Even though the mechanisms of histone modifications are not fully understood, several epigenetic therapies have shown great results in cancer treatment. One promising substance class are HDAC inhibitors, which prevent histone deacetylases from detaching the acetyl group from a histone, inducing cell cycle arrest, differentiation and cell death (Eckschlager et al. 2017). The HDAC inhibitors Vorinostat, Romidepsin and Belinostat have been approved by the FDA for the treatment of T cell lymphoma (Zhang et al. 2019). Many studies have concentrated on the combination of epigenetic therapy with well-established therapies and have demonstrated synergistic effects (Cao et al. 2015; Gao et al. 2016; Mann et al. 2007; Marks and Breslow 2007; Rettig et al. 2015). The addition of Vorinostat to Tamoxifen in breast cancer treatment resulted in tumor regression or prolonged disease stabilization in patients who had progressed on prior hormonal therapy (Thomas et al. 2011). In preclinical trials, HDAC inhibitors showed the ability to re-sensitize tamoxifen-resistant cells and prevent hormone therapy resistance (Munster et al. 2011), as well as a potentiation of the immune checkpoint inhibitor blockade in triple negative breast cancer in mice (Terranova-Barberio et al. 2017).
Our results show significant correlations of H3K9ac, H3K4me3 to Her2neu and ER as well as to survival data in breast cancer. As we examined the level of histone modifications at a fixed time by immunohistochemistry, we cannot say, if these results are cause or consequence of the cancer phenotype. Further experiments are, therefore, needed.
Further investigation of histone modifications in breast cancer could lead to a deeper understanding of the molecular mechanisms of cancer development. It could result in reliable screening methods, as well as the identification of new therapeutic targets for breast cancer treatment.
Conclusions
Histone modifications play an important role in the prognosis of breast cancer. H3K4me3 expression was correlated with positive estrogen receptor status, while H3K9ac staining was correlated with positive Her2 receptor status. High H3K4me3 and H3K9ac expression were correlated with shorter breast cancer-specific survival as well as shorter progression-free survival. Overall survival was decreased in patients with high nuclear H3K4me3 staining.
Several epigenetic therapies have already shown great results in cancer treatment. Further investigation of histone modifications in breast cancer could lead to a deeper understanding of the molecular mechanisms of cancer development. It could result in the identification of new therapeutic targets for breast cancer treatment.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Abbreviations
- H3K4me3:
-
Histone H3 trimethyl K4
- H3K4ac:
-
Histone H3 acetyl K9
- DCIS:
-
Ductal carcinoma in situ
- HDAC:
-
Histone deacetylase inhibitor
- LCIS:
-
Lobular carcinoma in situ
- OS:
-
Overall survival
- DSS:
-
Disease-specific survival
- PFS:
-
Recurrence-free survival
- Local PFS:
-
Period of time until a local recurrence was diagnosed
- DDFS:
-
Period of time until metastasis is diagnosed
References
Annunziato AT, Hansen JC (2000) Role of histone acetylation in the assembly and modulation of chromatin structures. Gene Expr 9:37–61. https://doi.org/10.3727/000000001783992687
Audia JE, Campbell RM (2016) Histone modifications and cancer. Cold Spring Harbor Perspect Biol 8:a019521. https://doi.org/10.1101/cshperspect.a019521
Bell R, Barraclough R, Vasieva O (2017) Gene expression meta-analysis of potential metastatic breast cancer markers. Curr Mol Med 17:200–210. https://doi.org/10.2174/1566524017666170807144946
Beyer S et al (2017) Histone H3 acetyl K9 and histone H3 Tri methyl K4 as prognostic markers for patients with cervical cancer. Int J Mol Sci. https://doi.org/10.3390/ijms18030477
Cao Q-f, Qian S-b, Wang N, Zhang L, Wang W-m, Shen H-b (2015) TRPM2 mediates histone deacetylase inhibition-induced apoptosis in bladder cancer cells. Cancer Biother Radiopharm 30:87–93. https://doi.org/10.1089/cbr.2014.1697
DeSantis CE et al (2019) Breast cancer statistics, 2019. CA Cancer J Clin 69:438–451. https://doi.org/10.3322/caac.21583
Eckschlager T, Plch J, Stiborova M, Hrabeta J (2017) Histone deacetylase inhibitors as anticancer drugs. Int J Mol Sci. https://doi.org/10.3390/ijms18071414
Gao M et al (2016) Therapeutic potential and functional interaction of carfilzomib and vorinostat in T cell leukemia/lymphoma. Oncotarget 7:29102–29115. https://doi.org/10.18632/oncotarget.8667
He C et al (2012) High expression of trimethylated histone H3 lysine 4 is associated with poor prognosis in hepatocellular carcinoma. Hum Pathol 43:1425–1435. https://doi.org/10.1016/j.humpath.2011.11.003
Healey MA, Hu R, Beck AH, Collins LC, Schnitt SJ, Tamimi RM, Hazra A (2014) Association of H3K9me3 and H3K27me3 repressive histone marks with breast cancer subtypes in the Nurses' Health Study. Breast Cancer Res Treat 147:639–651. https://doi.org/10.1007/s10549-014-3089-1
Kimura H (2013) Histone modifications for human epigenome analysis. J Hum Genet 58:439. https://doi.org/10.1038/jhg.2013.66
Lawrence M, Daujat S, Schneider R (2016) Lateral thinking: how histone modifications regulate gene expression trends in genetics. TIG 32:42–56. https://doi.org/10.1016/j.tig.2015.10.007
Lee DY, Hayes JJ, Pruss D, Wolffe AP (1993) A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell 72:73–84. https://doi.org/10.1016/0092-8674(93)90051-q
Leszinski G, Gezer U, Siegele B, Stoetzer O, Holdenrieder S (2012) Relevance of histone marks H3K9me3 and H4K20me3 in cancer. Anticancer Res 32:2199–2205
Li S, Shen L, Chen KN (2018) Association between H3K4 methylation and cancer prognosis: a meta-analysis. Thorac Cancer 9:794–799. https://doi.org/10.1111/1759-7714.12647
Lima-Fernandes E, Murison A, da Silva Medina T et al (2019) Targeting bivalency de-represses Indian Hedgehog and inhibits self-renewal of colorectal cancer-initiating cells. Nat Commun 10(1):1436. https://doi.org/10.1038/s41467-019-09309-4
Liu B-l et al (2010) Global Histone Modification Patterns as Prognostic Markers to Classify Glioma Patients Cancer. Epidemiol Biomark Prev 19:2888–2896. https://doi.org/10.1158/1055-9965.epi-10-0454
Mann BS, Johnson JR, Cohen MH, Justice R, Pazdur R (2007) FDA approval summary: vorinostat for treatment of advanced primary cutaneous T cell lymphoma. Oncologist 12:1247–1252. https://doi.org/10.1634/theoncologist.12-10-1247
Marks PA, Breslow R (2007) Dimethyl sulfoxide to vorinostat: development of this histone deacetylase inhibitor as an anticancer drug. Nat Biotechnol 25:84. https://doi.org/10.1038/nbt1272
Munster PN et al (2011) A phase II study of the histone deacetylase inhibitor vorinostat combined with tamoxifen for the treatment of patients with hormone therapy-resistant breast cancer. Br J Cancer 104:1828–1835. https://doi.org/10.1038/bjc.2011.156
Prat A et al (2015) Clinical implications of the intrinsic molecular subtypes of breast cancer. Breast (Edinburgh, Scotland) 24(Suppl 2):S26–35. https://doi.org/10.1016/j.breast.2015.07.008
Rettig I et al (2015) Selective inhibition of HDAC8 decreases neuroblastoma growth in vitro and in vivo and enhances retinoic acid-mediated differentiation. Cell Death Dis 6:e1657. https://doi.org/10.1038/cddis.2015.24
Ruthenburg AJ, Li H, Patel DJ, Allis CD (2007) Multivalent engagement of chromatin modifications by linked binding modules. Nat Rev Mol Cell Biol 8:983–994. https://doi.org/10.1038/nrm2298
Shanmugam MK et al (2018) Role of novel histone modifications in cancer. Oncotarget 9:11414–11426. https://doi.org/10.18632/oncotarget.23356
Shilatifard A (2008) Molecular implementation and physiological roles for histone H3 lysine 4 (H3K4) methylation. Curr Opin Cell Biol 20:341–348. https://doi.org/10.1016/j.ceb.2008.03.019
Takahashi YH, Shilatifard A (2010) Structural basis for H3K4 trimethylation by yeast Set1/COMPASS. Adv Enzyme Regul 50:104–110. https://doi.org/10.1016/j.advenzreg.2009.12.005
Terranova-Barberio M et al (2017) HDAC inhibition potentiates immunotherapy in triple negative breast cancer. Oncotarget 8:114156–114172. https://doi.org/10.18632/oncotarget.23169
Thomas S, Thurn KT, Bicaku E, Marchion DC, Munster PN (2011) Addition of a histone deacetylase inhibitor redirects tamoxifen-treated breast cancer cells into apoptosis, which is opposed by the induction of autophagy. Breast Cancer Res Treat 130:437–447. https://doi.org/10.1007/s10549-011-1364-y
Torre LA, Islami F, Siegel RL, Ward EM, Jemal A (2017) Global cancer in women: burden and trends cancer. Epidemiol Biomark Prev 26:444–457. https://doi.org/10.1158/1055-9965.epi-16-0858
Vakoc CR, Sachdeva MM, Wang H, Blobel GA (2006) Profile of histone lysine methylation across transcribed mammalian chromatin. Mol Cell Biol 26:9185–9195. https://doi.org/10.1128/mcb.01529-06
Wang Z et al (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40:897–903. https://doi.org/10.1038/ng.154
Wu FR et al (2012) Differences in H3K4 trimethylation in in vivo and in vitro fertilization mouse preimplantation embryos. Genet Mol Res GMR 11:1099–1108. https://doi.org/10.4238/2012.April.27.9
Wu Y, Sarkissyan M, Vadgama JV (2015) Epigenetics in breast and prostate cancer. Methods Mol Biol (Clifton, NJ) 1238:425–466. https://doi.org/10.1007/978-1-4939-1804-1_23
Zhang P, Torres K, Liu X, Liu CG, Pollock RE (2016) An overview of chromatin-regulating proteins in cells. Curr Protein Peptide Sci 17:401–410. https://doi.org/10.2174/1389203717666160122120310
Zhang Q, Wang S, Chen J, Yu Z (2019) Histone deacetylases (HDACs) guided novel therapies for T cell lymphomas. Int J Med Sci 16:424–442. https://doi.org/10.7150/ijms.30154
Acknowledgements
Open Access funding provided by Projekt DEAL.
Funding
No funding.
Author information
Authors and Affiliations
Contributions
LB did the staining, analyzed the data and wrote the paper. TK Assistance in writing the paper and approvement of the manuscript. ES design of experiments, supervision. DM design of experiments. UJ and ND design of the experiment and supervision. All authors interpreted the data and read the paper and approved the manuscript. This work is part of the dissertation of Luisa Berger.
Corresponding author
Ethics declarations
Conflict of interest
T. Kolben has a relative employed at Roche and holds stock of Roche. T. M. Kolben is employed at Roche and holds stock of Roche. S. Mahner has received Research support, advisory board, honoraria and travel expenses from AstraZeneca, Clovis, Eisai, GlaxoSmithKline, Medac, MSD, Novartis, Olympus, PharmaMar, Roche, Sensor Kinesis, Teva, Tesaro¸ All other authors declare that they have no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Ethics approval
The study was conducted in consent to the Declaration of Helsinki and was approved by the local ethics committee of the Ludwig-Maximilians University of Munich (Reference No. 048-08; 2008).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Berger, L., Kolben, T., Meister, S. et al. Expression of H3K4me3 and H3K9ac in breast cancer. J Cancer Res Clin Oncol 146, 2017–2027 (2020). https://doi.org/10.1007/s00432-020-03265-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-020-03265-z