Abstract
Main conclusion
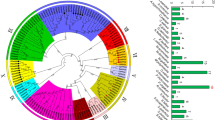
74 phytocyanin genes were identified in the Populus trichocarpa genome. Phylogenetic analysis grouped the PC proteins into four subfamilies (UCs, PLCs, SCs, and ENODLs). Closely related PC proteins share similar motifs, implying similar functions. Expression profiles of PtPC genes were analyzed in response to drought and salt-stress.
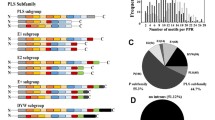
Phytocyanins (PCs) are blue copper proteins associated with electron carrier activity that have a large influence on plant growth and resistance. The majority of PCs are chimeric arabinogalactan proteins (AGPs). In this work, we identified 74 PC genes in Populus trichocarpa and analyzed them comprehensively. Based on the ligands composition of copper-binding sites, glycosylation state, the domain structure and spectral characteristics of PC genes, PCs were divided into four subfamilies [uclacyanins (UCs), plantacyanins (PLCs), stellacyanins (SCs) and early nodulin-like proteins (ENODLs)], and phylogenetic relationship analysis classified them into seven groups. All PtPCs are randomly distributed on 17 of the 19 poplar chromosomes, and they appear to have undergone expansion via segmental duplication. Eight PtPCs do not contain introns, and each group has a similar conserved motif structure. Promoter analysis revealed cis-elements related to growth, development and stress responses, and established orthology relationships of PCs between Arabidopsis and poplar by synteny analysis. Expression profile analysis and qRT-PCR analysis showed that PtPCs were expressed widely in various tissues. Quantitative real-time RT-PCR analysis of PC genes expression in response to salt and drought stress revealed their stress-responses profiles. This work provides a theoretical basis for a further study of stress resistance mechanisms and the function of PC genes in poplar growth and development.









Similar content being viewed by others
Abbreviations
- AG:
-
Arabinogalactan
- AGPs:
-
Arabinogalactan proteins
- ENODLs:
-
Early nodulin-like proteins
- PCs:
-
Phytocyanins
- PCLD:
-
Plastocyanin-like domain
- PLCs:
-
Plantacyanins
- SCs:
-
Stellacyanins
- SP:
-
Signal peptide
- UCs:
-
Uclacyanins
- K s :
-
Number of synonymous substitutions per synonymous site
- K a :
-
Number of non-synonymous substitutions per non-synonymous site
References
Anderson SL, Teakle GR, Martino-Catt SJ, Kay SA (1994) Circadian clock- and phytochrome regulated transcription is conferred by a 78 bp cis-acting domain of the Arabidopsis CAB2 promoter. Plant J 6(4):457–470
Bobb AJ, Chern MS, Bustos MM (1997) Conserved RY-repeats mediate transactivation of seed-specific promoters by the developmental regulator PvALF. Nucleic Acids Res 25(3):641–647
Bowers JE, Chapman BA, Rong J, Paterson AH (2003) Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422(6930):433–438
Cao J, Li X, Lv Y, Ding L (2015) Comparative analysis of the phytocyanin gene family in 10 plant species: a focus on Zea mays. Front Plant Sci 6:515. https://doi.org/10.3389/fpls.2015.00515
Cruz-Garcia F, Nathan HC, Kim D, Mcclure B (2005) Stylar glycoproteins bind to S-RNase in vitro. Plant J 42(3):295–304
Denancé N, Szurek B, Noël LD (2014) Emerging functions of nodulin-like proteins in non-nodulating plant species. Plant Cell Physiol 55(3):469–474
Diab AA, Teulat-Merah B, This D, Ozturk NZ, Benscher D, Sorrells ME (2004) Identification of drought-inducible genes and differentially expressed sequence tags in barley. Theor Appl Genet 109(7):1417–1425
Dong J, Kim ST, Lord EM (2005) Plantacyanin plays a role in reproduction in Arabidopsis. Plant Physiol 138(2):778–789
Eisenhaber B, Wildpaner M, Schultz CJ, Borner GH, Dupree P, Eisenhaber F (2003) Glycosylphosphatidylinositol lipid anchoring of plant proteins. Sensitive prediction from sequence- and genome-wide studies for Arabidopsis and rice. Plant Physiol 133(4):1691–1701. https://doi.org/10.1104/pp.103.023580
Escobar-Restrepo JM, Huck N, Kessler S, Gagliardini V, Gheyselinck J, Yang WC, Grossniklaus U (2007) The FERONIA receptor-like kinase mediates male–female interactions during pollen tube reception. Science 317(5938):656–660
Ezaki B, Katsuhara M, Kawamura M, Matsumoto H (2001) Different mechanisms of four aluminum (Al)-resistant transgenes for Al toxicity in Arabidopsis. Plant Physiol 127(3):918–927
Ezaki B, Sasaki K, Matsumoto H, Nakashima S (2005) Functions of two genes in aluminium (Al) stress resistance: repression of oxidative damage by the AtBCB gene and promotion of efflux of Al ions by the NtGDI1gene. J Exp Bot 56(420):2661–2671
Fedorova M, Mortel JVD, Matsumoto PA, Cho J, Town CD, VandenBosch KA, Gantt JS, Vance CP (2002) Genome-wide identification of nodule-specific transcripts in the model legume Medicago truncatula. Plant Physiol 130(2):519–537
Giri AV, Anishetty S, Gautam P (2004) Functionally specified protein signatures distinctive for each of the different blue copper proteins. BMC Bioinform 5(1):1–8
Goldsbrough AP, Albrecht H, Stratford R (1993) Salicylic acid-inducible binding of a tobacco nuclear protein to a 10 bp sequence which is highly conserved amongst stress-inducible genes. Plant J 3(4):563–571
Goodstein DM, Shu S, Russell H, Rochak N, Hayes RD, Joni F, Therese M, William D, Uffe H, Nicholas P (2011) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:1178–1186
Gu Z, Steinmetz LM, Gu X, Scharfe C, Davis RW, Li WH (2003) Role of duplicate genes in genetic robustness against null mutations. Nature 421(6918):63–66
Hart PJ, Nersissian AM, Herrmann RG, Nalbandyan RM, Valentine JS, Eisenberg D (1996) A missing link in cupredoxins: crystal structure of cucumber stellacyanin at 1.6 A resolution. Protein Sci 5(11):2175–2183
Hou Y, Guo X, Cyprys P, Zhang Y, Bleckmann A, Cai L, Huang Q, Luo Y, Gu H, Dresselhaus T, Dong J, Qu L (2016) Maternal ENODLs are required for pollen tube reception in Arabidopsis. Curr Biol 26(17):2343–2350
Hu R, Qi G, Kong Y, Kong D, Qian G, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10(1):145. https://doi.org/10.1186/1471-2229-10-145
Hui M, Lin F, Zhu C, Xue C, Hualin Z, Yan X (2014) Genome-wide identification and expression analysis of the IQD gene family in Populus trichocarpa. Plant Sci 229:96–110
Jeßberger N, Krey VM, Rademacher C, Böhm ME, Mohr AK, Ehlingschulz M, Scherer S, Märtlbauer E (2015) From genome to toxicity: a combinatory approach highlights the complexity of enterotoxin production in Bacillus cereus. Front Microbiol 6(6):560
Johnson KL, Cassin AM, Lonsdale A, Bacic A, Doblin MS, Schultz CJ (2017) A motif and amino acid bias bioinformatics pipeline to identify hydroxyproline-rich glycoproteins. Plant Physiol 174(2):886–903
Khan JA, Wang Q, Sjölund RD, Schulz A, Thompson GA (2007) An early nodulin-like protein accumulates in the sieve element plasma membrane of Arabidopsis. Plant Physiol 143(4):1576–1589
Li J, Yu M, Geng LL, Zhao J (2010) The fasciclin-like arabinogalactan protein gene, FLA3, is involved in microspore development of Arabidopsis. Plant J 64(3):482–497
Li J, Gao G, Zhang T, Wu X (2013) The putative phytocyanin genes in Chinese cabbage (Brassica rapa L.): genome-wide identification, classification and expression analysis. Mol Genet Genom 288(1–2):1–20
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290(5494):1151–1155
Ma H, Jie Z (2010) Genome-wide identification, classification, and expression analysis of the arabinogalactan protein gene family in rice (Oryza sativa L.). J Exp Bot 61(10):2647–2668
Ma H, Zhao J (2010) Genome-wide identification, classification, and expression analysis of the arabinogalactan protein gene family in rice (Oryza sativa L.). J Exp Bot 61(10):2647–2668
Ma H, Zhao H, Liu Z, Zhao J (2011) The phytocyanin gene family in rice (Oryza sativa L.): genome-wide identification, classification and transcriptional analysis. PLoS One 6(10):e25184
Ma Y, Yan C, Li H, Wu W, Liu Y (2017) Bioinformatics prediction and evolution analysis of arabinogalactan proteins in the plant kingdom. Front Plant Sci 8:1–17
MacMillan CP, Mansfield SD, Stachurski ZH, Evans R, Southerton SG (2010) Fasciclin-like arabinogalactan proteins: specialization for stem biomechanics and cell wall architecture in Arabidopsis and Eucalyptus. Plant J 62(4):689–703
Mashiguchi K, Yamaguchi I, Suzuki Y (2004) Isolation and identification of glycosylphosphatidylinositol-anchored arabinogalactan proteins and novel β-glucosyl Yariv-reactive proteins from seeds of rice (Oryza sativa). Plant Cell Physiol 45(12):1817–1829
Mashiguchi K, Asami T, Suzuki Y (2009) Genome-wide identification, structure and expression studie, and mutant collection of 22 early nodulin-like protein genes in Arabidopsis. Biosci Biotechnol Biochem 73(11):2452–2459
Nejad ES, Askari H, Soltani S (2012) Regulatory TGACG-motif may elicit the secondary metabolite production through inhibition of active cyclin-dependent kinase/cyclin complex. Plant Omics 5(6):553–558
Nersissian AM, Immoos C, Hill MG, Hart PJ, Williams G, Herrmann RG, Valentine JS (1998) Uclacyanins, stellacyanins, and plantacyanins are distinct subfamilies of phytocyanins: plant-specific mononuclear blue copper proteins. Protein Sci 7(9):1915–1929
Ozturk ZN, Talamé V, Deyholos M, Michalowski CB, Galbraith DW, Gozukirmizi N, Tuberosa R, Bohnert HJ (2002) Monitoring large-scale changes in transcript abundance in drought- and salt-stressed barley. Plant Mol Biol 48(5–6):551–573
Petersen TN, Brunak S, Von HG, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8(10):785–786
Poon S, Heath R, Clarke A (2013) A chimeric arabinogalactan protein promotes somatic embryogenesis in cotton cell culture. Plant Physiol 160(2):684–695
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3(6):1101–1108
Schultz CJ, Rumsewicz MP, Johnson KL, Jones BJ, Gaspar YM, Bacic A (2002) Using genomic resources to guide research directions. The arabinogalactan protein gene family as a test case. Plant Physiol 129(4):1448–1463
Seifert GJ, Roberts K (2007) The biology of arabinogalactan proteins. Annu Rev Plant Biol 58(1):137–161
Sessa G, Morelli G, Ruberti I (1993) The Athb-1 and -2 HD-Zip domains homodimerize forming complexes of different DNA binding specificities. EMBO J 12(9):3507–3517
Shen Q, Ho TH (1995) Functional dissection of an abscisic acid (ABA)-inducible gene reveals two independent ABA-responsive complexes each containing a G-box and a novel cis-acting element. Plant Cell 7(3):295–307
Showalter AM, Keppler B, Lichtenberg J, Gu D, Welch LR (2010) A bioinformatics approach to the identification, classification, and analysis of hydroxyproline-rich glycoproteins. Plant Physiol 153(2):485–513
Showalter AM, Keppler B, Liu X, Lichtenberg J, Welch LR (2016) Bioinformatic identification and analysis of hydroxyproline-rich glycoproteins in Populus trichocarpa. BMC Plant Biol 16(1):229
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16(8):2001–2019
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Tan L, Qiu F, Lamport DTA, Kieliszewski MJ (2004) Structure of a hydroxyproline (Hyp)-arabinogalactan polysaccharide from repetitive Ala-Hyp expressed in transgenic Nicotiana tabacum. J Biol Chem 279(13):1315656–1315665
Vandepoele K, Simillion C, Van de Peer Y (2003) Evidence that rice and other cereals are ancient aneuploids. Plant Cell 15(9):2192–2202
Wang Y, Feng L, Zhu Y, Li Y, Yan H, Xiang Y (2015) Comparative genomic analysis of the WRKY III gene family in populus, grape, arabidopsis and rice. Biol Direct 10:48. https://doi.org/10.1186/s13062-015-0076-3
Washida H, Wu CY, Suzuki A, Yamanouchi U, Akihama T, Harada K, Takaiwa F (1999) Identification of cis-regulatory elements required for endosperm expression of the rice storage protein glutelin gene GluB-1. Plant Mol Biol 40(1):1–12
Wu H, Shen Y, Hu Y, Tan S, Lin Z (2011) A phytocyanin-related early nodulin-like gene, BcBCP1, cloned from Boea crassifolia enhances osmotic tolerance in transgenic tobacco. J Plant Physiol 168(9):935–943
Wu M, Li Y, Chen D, Liu H, Zhu D, Xiang Y (2016) Genome-wide identification and expression analysis of the IQD gene family in moso bamboo (Phyllostachys edulis). Sci Rep UK 6:24520
Xu L, Wang XJ, Wang T, Li LB (2017) Genome-wide identification, classification, and expression analysis of the phytocyanin gene family in Phalaenopsis equestris. Biol Plant 61(3):1–8
Ye H, Du H, Tang N, Li X, Xiong L (2009) Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol Biol 71(3):291–305
Yoshizaki M, Furumoto T, Hata S, Shinozaki M, Izui K (2000) Characterization of a novel gene encoding a phytocyanin-related protein in morning glory (Pharbitis nil). Biochem Biophys Res Commun 268(2):466–470
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18(6):292–298
Zhang R, Tucker MR, Burton RA, Shirley NJ, Little A, Morris J, Milne L, Houston K, Hedley PE, Waugh R, Fincher GB (2016) The dynamics of transcript abundance during cellularisation of developing barley endosperm. Plant Physiol 170(3):1549–1565
Acknowledgements
We thank the members of the Laboratory of Modern Biotechnology for their assistance in this study.
Funding
National Natural Science Foundation of China (31370561) and National Science and Technology Support Plan Corpus (2015BAD07B070104).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Protein backbones of PCs in P. trichocarpa. The coloured sequences at the N and C terminus indicate predicted signal peptides (green) and GPI anchor addition sequences (purple) if present in the sequences. Putative AG sites (blue) are also indicated. Red font indicates N-glycosylation site. Extensins SP3, SP4 and SP5 repeats are indicated (light blue) if present. Note that the putative AG glycomodules are only checked in the proteins containing N-secretion signals (TIFF 32125 kb)
Fig. S2
Exon–intron structures and conserved domains of the predicted PtPC proteins. Exons, introns and untranslated regions (UTRs) are represented by yellow rectangles, grey lines and blue rectangles, respectively (TIFF 437 kb)
Fig. S3
Sliding window plots of Ka/Ks ratios of representative duplicated PtPC gene paralogs. The x-axis denotes the nucleotide position, and the y-axis denotes the Ka/Ks ratio (TIFF 68 kb)
Fig. S4
Extensive microsynteny of PC regions between Arabidopsis and poplar. Arabidopsis and Populus chromosomes are shown in different colours. Numbers along each chromosome box indicate sequence length in megabases. Whole chromosomes of the two species harbouring PC regions are encircled. Black lines represent syntenic relationships between PC regions (TIFF 3195 kb)
Fig. S5
Synteny analysis of PC genes in Populus. Black lines represent syntenic relationships (TIFF 2627 kb)
Fig. S6
Hierarchical clustering of poplar PC gene expression. The heatmap shows hierarchical clustering of PtPC genes across various tissues/organs. Affymetrix microarray data under accession number GSE13990 encompassing results from six organ/tissue types including young leaves (YL), roots (R), xylem (XY), female catkin (FC), male catkin (MC) and mature leaves (ML) were re-analysed (TIFF 621 kb)
Rights and permissions
About this article
Cite this article
Luo, S., Hu, W., Wang, Y. et al. Genome-wide identification, classification, and expression of phytocyanins in Populus trichocarpa. Planta 247, 1133–1148 (2018). https://doi.org/10.1007/s00425-018-2849-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-018-2849-2