Abstract
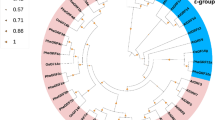
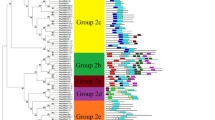
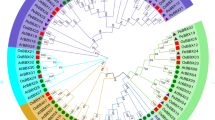
Gibberellic acid-stimulated transcript (GAST) proteins are widely distributed among plant species and are involved in various aspects of plant growth, plant responses to stress, and hormone crosstalk. However, in moso bamboo (Phyllostachys edulis), bioinformatics data and functional information on the GAST family are limited. Here, a total of eight GAST genes were first identified in the whole genome of moso bamboo and then clustered into three groups based on phylogenetic analysis. Each clade is well supported by the conserved intron/exon structures and motifs, suggesting that the members within the same subfamily perform similar functions. A number of development-, hormone-, and stress-related elements are randomly distributed in the promoter sequences of PheGASTs. Tissue expression analysis indicates that most PheGASTs might be associated with moso bamboo flower development and rapid shoot growth. Furthermore, the PheGASTs display phytohormone- and stress-inducible expression patterns in response to various circumstances (abscisic acid, gibberellins, cold, drought, and salinity). Among these, PheGASR1 was assumed to have a distinct regulatory role in rapid shoot growth and abiotic stress responses involving the ABA pathway. Collectively, our results provide extensive insights into the GAST gene family, thereby contributing to the screening of additional candidate genes in moso bamboo.









Similar content being viewed by others
References
Almasia NI, Narhirñak V, Hopp HE, Vazquez-Rovere C (2010) Isolation and characterization of the tissue and development-specific potato snakin-1 promoter inducible by temperature and wounding. Electron J Biotechnol 13:8–9. https://doi.org/10.2225/vol13-issue5-fulltext-12
Alonso-Ramirez A, Rodríguez D, Reyes D, Jiménez JA, Nicolás G, López-Climent M, Gómez-Cadenas A, Nicolás C (2009) Evidence for a role of gibberellins in salicylic acid-modulated early plant responses to abiotic stress in Arabidopsis seeds. Plant Physiol 150:1335–1344. https://doi.org/10.1104/pp.109.139352
Aubert D, Chevillard M, Dorne AM, Arlaud G, Herzog M (1998) Expression patterns of GASA genes in Arabidopsis thaliana: the GASA4 gene is up-regulated by gibberellins in meristematic regions. Plant Mol Biol 36:871–883. https://doi.org/10.1023/A:1005938624418
Bailey TL, Gribskov M (1998) Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14:48–54. https://doi.org/10.1093/bioinformatics/14.1.48
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren JY, Li WW, Noble WS (2009) MEME suite: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. https://doi.org/10.1093/nar/gkp335
Ben-Nissan G, Lee JY, Borohov A, Weiss D (2004) GIP, a Petunia hybrida GA-induced cysteine-rich protein: a possible role in shoot elongation and transition to flowering. Plant J 37:229–238. https://doi.org/10.1046/j.1365-313X.2003.01950.x
Chen D, Chen Z, Wu M, Wang Y, Wang Y, Yan H, Xiang Y (2017) Genome-wide identification and expression analysis of the HD-zip gene family in moso bamboo (Phyllostachys edulis). J Plant Growth Regul 36:323–337. https://doi.org/10.1007/s00344-016-9642-x
de la Fuente JI, Amaya I, Castillejo C, Sanchez-Sevilla JF, Quesada MA, Botella MA, Valpuesta V (2006) The strawberry gene FaGAST affects plant growth through inhibition of cell elongation. J Exp Bot 57:2401–2411. https://doi.org/10.1093/jxb/erj213
Fan C, Ma J, Guo Q, Li X, Wang H, Lu M (2013) Selection of reference genes for quantitative real-time PCR in bamboo (Phyllostachys edulis). PLoS ONE 8:e56573. https://doi.org/10.1371/journal.pone.0056573
Furukawa T, Sakaguchi N, Shimada H (2006) Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet Syst 81:171
Gamuyao R, Nagai K, Ayano M, Mori Y, Minami A, Kojima M, Suzuki T, Sakakibara H, Higashiyama T, Ashikari M, Reuscher S (2017) Hormone distribution and transcriptome profiles in bamboo shoots provide insights on bamboo stem emergence and growth. Plant Cell Physiol. https://doi.org/10.1093/pcp/pcx023
Gao J, Zhang Y, Zhang C, Qi F, Li X, Mu S, Peng Z (2014) Characterization of the floral transcriptome of Moso bamboo (Phyllostachys edulis) at different flowering developmental stages by transcriptome sequencing and RNA-seq analysis. PLoS ONE 9:e98910. https://doi.org/10.1371/journal.pone.0098910
Gaut BS (2010) Evolutionary dynamics of grass genomes. New Phytol 154:15–28. https://doi.org/10.1046/j.1469-8137.2002.00352.x
Ge W, Zhang Y, Cheng Z, Hou D, Li X, Gao J (2017) Main regulatory pathways, key genes and microRNAs involved in flower formation and development of moso bamboo (Phyllostachys edulis). Plant Biotechnol J 15:82–96. https://doi.org/10.1111/pbi.12593
Guyot R, Keller B (2004) Ancestral genome duplication in rice. Genome 47:610. https://doi.org/10.1139/g04-016
Jiang X, Lou C, Yuan N, Yue JJ, Gu XP (2016) Effects of exogenous GA3 on culm form and culm fiber quality of Phyllostachys edulis seedlings neonatal tiller. J Nanjing For Univ 40:121–126
Jie Q, Kang SG, Hah C, Jang JC (2016) Molecular and cellular characterization of GA-stimulated transcripts GASA4 and GASA6 in Arabidopsis thaliana. Plant Sci 246:1–10. https://doi.org/10.1016/j.plantsci.2016.01.009
Kellogg EA (1998) Relationships of cereal crops and other grasses. Proc Natl Acad Sci USA 95:2005–2010
Ko CB, Woo YM, Lee DJ, Lee MC, & Kim CS (2007) Enhanced tolerance to heat stress in transgenic plants expressing the GASA4 gene. Plant Physiol Biochem 45(9):722–728. https://doi.org/10.1016/j.plaphy.2007.07.010
Lee SC, Han SK, Kim SR (2015) Salt- and ABA-inducible OsGASR1 is involved in salt tolerance. J Plant Biol 58:96–101. https://doi.org/10.1007/s12374-014-0497-z
Lee SC, Kim SJ, Han SK, An G, Kim SR (2017) A gibberellin-stimulated transcript, OsGASR1, controls seedling growth and α-amylase expression in rice. J Plant Physiol 214:116–122. https://doi.org/10.1016/j.jplph.2017.04.010
Li L, Hu T, Li X, Mu S, Cheng Z, Ge W, Gao J (2016) Genome-wide analysis of shoot growth-associated alternative splicing in moso bamboo. Mol Genet Genom 291:1695–1714. https://doi.org/10.1007/s00438-016-1212-1
Li L, Cheng Z, Ma Y, Bai Q, Li X, Cao Z, Wu Z, Gao J (2017) The association of hormone signalling genes, transcription and changes in shoot anatomy during moso bamboo growth. Plant Biotechnol J 16(1):72–85. https://doi.org/10.1111/pbi.12750
Liu ZH, Zhu L, Shi HY, Chen Y, Zhang JM, Zheng Y, Li XB (2013) Cotton GASL genes encoding putative gibberellin-regulated proteins are involved in response to GA signaling in fiber development. Mol Biol Rep 40:4561. https://doi.org/10.1007/s11033-013-2543-1
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290:1151. https://doi.org/10.1126/science.290.5494.1151
Nahirñak V, Almasia NI, Hopp HE, Vazquezrovere C (2012) Snakin/GASA proteins: involvement in hormone crosstalk and redox homeostasis. Plant Signal Behav 7:1004–1008. https://doi.org/10.4161/psb.20813
Nahirñak V, Rivarola M, Urreta MGD, Paniego N, Hopp HE, Almasia NI, Vazquez-Rovere C (2016) Genome-wide analysis of the snakin/GASA gene family in Solanum tuberosum cv. Kennebec Am J Potato Res 93:172–188. https://doi.org/10.1007/s12230-016-9494-8
Peng Z et al (2013a) The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nat Genet 45:456–461. https://doi.org/10.1038/ng.2569
Peng Z, Zhang C, Zhang Y, Hu T, Mu S, Li X, Gao J (2013b) Transcriptome sequencing and analysis of the fast growing shoots of moso bamboo (Phyllostachys edulis). PLoS ONE 8:e78944. https://doi.org/10.1371/journal.pone.0078944
Qi F, Hu T, Peng Z, Gao J (2013) Screening of reference genes used in qRT-PCR and expression analysis of PheTFL1 gene in moso bamboo. Acta Bot Boreali-Occident Sin 33:48–52
Rozas J (2009) DNA sequence polymorphism analysis using DnaSP. Methods Mol Biol 537:337
Rubinovich L, Weiss D (2010) The Arabidopsis cysteine-rich protein GASA4 promotes GA responses and exhibits redox activity in bacteria and in planta. Plant J 64:1018–1027. https://doi.org/10.1111/j.1365-313X.2010.04390.x
Rubinovich L, Ruthstein S, Weiss D (2014) The Arabidopsis cysteine-rich GASA5 is a redox-active metalloprotein that suppresses gibberellin responses. Mol Plant 7:244–247. https://doi.org/10.1093/mp/sst141
Sun S, Wang H, Yu H, Zhong C, Zhang X, Peng J, Wang X (2013) GASA14 regulates leaf expansion and abiotic stress resistance by modulating reactive oxygen species accumulation. J Exp Bot 64:1637–1647. https://doi.org/10.1093/jxb/ert021
Sun H, Li L, Lou Y, Zhao H, Gao Z (2016) Genome-wide identification and characterization of aquaporin gene family in moso bamboo (Phyllostachys edulis). Mol Biol Rep 43:437–450. https://doi.org/10.1007/s11033-016-3973-3
Székely G (2009) Oxidative stress in plants. Stud Univ Babes-Bolyai Biol 10:19–19
Trapalis M, Li SF, Parish RW, Li SF (2017) The Arabidopsis GASA10 gene encodes a cell wall protein strongly expressed in developing anthers and seeds. Plant Sci 260:71. https://doi.org/10.1016/j.plantsci.2017.04.003
Wang L, Wang Z, Xu Y, Joo SH, Kim SK, Xue Z, Xu Z, Wang Z, Chong K (2009) OsGSR1 is involved in crosstalk between gibberellins and brassinosteroids in rice. Plant J 57:498–510. https://doi.org/10.1111/j.1365-313X.2008.03707.x
Wang T, Wang H, Cai D, Gao Y, Zhang H, Wang Y, Lin C, Ma L, Gu L (2017) Comprehensive profiling of rhizome-associated alternative splicing and alternative polyadenylation in moso bamboo (Phyllostachys edulis). Plant J 91(4):684–699. https://doi.org/10.1111/tpj.13597
Wigoda N, Ben-Nissan G, Granot D, Schwartz A, Weiss D (2006) The gibberellin-induced, cysteine-rich protein GIP2 from Petunia hybrida exhibits in planta antioxidant activity. Plant J 48:796–805. https://doi.org/10.1111/j.1365-313X.2006.02917.x
Wu H, Lv H, Li L, Liu J, Mu S, Li X, Gao J (2015) Genome-wide analysis of the AP2/ERF transcription factors family and the expression patterns of DREB Genes in Moso Bamboo (Phyllostachys edulis). PLoS ONE 10:e0126657. https://doi.org/10.1371/journal.pone.0126657
Wu M, Li Y, Chen D, Liu H, Zhu D, Xiang Y (2016) Genome-wide identification and expression analysis of the IQD gene family in moso bamboo (Phyllostachys edulis). Sci Rep 6:24520. https://doi.org/10.1038/srep24520
Yu J, Hu S, Wang J et al (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296:79–92. https://doi.org/10.1126/science.1068037
Zhang S, Wang X (2008) Expression analysis on GASA genes downstream of DELLA in Arabidopsis. Sci Bull 53:2760–2767
Zhang J, Jia H, Li J, Li Y, Lu M, Hu J (2016) Molecular evolution and expression divergence of the Populus euphratica Hsf genes provide insight into the stress acclimation of desert poplar. Sci Rep 6:30050. https://doi.org/10.1038/srep30050
Zhang LY, Geng XL, Zhang HY, Zhou CL, Zhao AJ, Wang F, Zhao Y, Tian XJ, Hu ZR, Xin MM, Yao YY, Ni ZF, Sun QX, Peng HR (2017) Isolation and characterization of heat-responsive gene TaGASR1 from wheat (Triticum aestivum L.). J Plant Biol 60:57–65. https://doi.org/10.1007/s12374-016-0484-7
Zhao H, Peng Z, Fei B, Li L, Hu T, Gao Z, Jiang Z (2014) BambooGDB: a bamboo genome database with functional annotation and an analysis platform Database. Database 2014:bau006. https://doi.org/10.1093/database/bau006
Zimmermann R, Sakai H, Hochholdinger F (2010) The gibberellic acid stimulated-like gene family in maize and its role in lateral root development. Plant Physiol 152:356–365. https://doi.org/10.1104/pp.109.149054
Funding
Fundamental Research Funds of ICBR (Grant Number 1632018006) and the National Natural Science Foundation of China (Grant Number 31570673) supported this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hou, D., Bai, Q., Li, J. et al. The Gibberellic Acid-Stimulated Transcript Gene Family in Moso Bamboo: A Genome-Wide Survey and Expression Profiling During Development and Abiotic Stresses. J Plant Growth Regul 37, 1135–1147 (2018). https://doi.org/10.1007/s00344-018-9805-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-018-9805-z