Abstract
Objectives
To investigate the imaging findings and clinical time course of COVID-19 pneumonia.
Methods
A total of 113 baseline and follow-up CT scans from 24 January 2020 to 18 February 2020 were longitudinally collected from 29 confirmed COVID-19 patients in a single center. The changes in the clinical and laboratory characteristics, imaging features, lesion-to-muscle ratio (LMR), and pulmonary inflammation index (PII) at baseline, 1–6 days, 7–13 days, and ≥ 14 days were compared.
Results
Of the 29 COVID-19 patients enrolled, the baseline chest CT scan was obtained 3 ± 2 (0–9) days after the onset of symptoms, and each patient had an average of 4 ± 1 (3–5) CT scans with a mean interval of 5 ± 2 (1–14) days. The percentage of patients with fever, cough, shortness of breath, and myalgia obviously decreased at 7–13 days with regular treatment (p < 0.05). The lymphocyte count, C-reactive protein, interleukin-6, and oxygenation index worsened within 1–6 days but improved sharply at 7–13 days. Compared with those at the other three time points, the LMR, PII, and number of involved lobes at 1–6 days were the highest, and gradually improved after 7–13 days.
Conclusions
Lung lesion development on chest CT reflects the clinical time course of COVID-19 progression over 1–6 days, followed by clinical improvement and the resorption of lesions. CT imaging may be indicated when patients fail to improve within a week of treatment, but repeated chest CT may be unnecessary when the patients show improvements clinically.
Key Points
• Chest CT reflects the development of coronavirus disease 2019 pneumonia (COVID-19).
• COVID-19 usually shows progressive lesions over up to 9 days with subsequent resorption.
• Unusual clinical time course of COVID-19 may indicate repeated chest CT.
Similar content being viewed by others
Introduction
Since December 8, 2019, several cases of unexplained pneumonia have been identified in Wuhan, Hubei Province, China [1,2,3], and a novel coronavirus was found in the throat swab of one patient [4]. This novel coronavirus, also known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was named 2019 novel coronavirus (2019-nCoV) by the World Health Organization (WHO) and is the cause of coronavirus disease 2019 (COVID-19). The genome sequence of SARS-CoV-2 is similar to but distinct from those of two other coronaviruses that have had outbreaks in the past: severe acute respiratory syndrome coronavirus (SARS-CoV; approximately 79% sequence identity) and Middle East respiratory syndrome coronavirus (MERS-CoV; approximately 50% sequence identity) [5].
COVID-19 pneumonia is highly infectious and develops rapidly, and it has led to a serious pandemic. Although chest CT has been pointed out as an important test in the diagnosis of patients with COVID-19 [6, 7], information on the development of CT imaging features along the clinical time course of COVID-19 is still scarce. This retrospective single-center cohort study attempted to disprove the hypothesis that chest CT findings at follow-up do not reflect the clinical time course of COVID-19.
Materials and methods
Patients
Our institutional review board waived the requirement to obtain written informed consent for this retrospective cohort study. Our hospital is one of the four major designated hospitals responsible for the treatment of COVID-19 pneumonia, as assigned by the government in the city. From 24 January 2020 to 18 February 2020, totally 73 patients diagnosed with SARS-CoV-2 infection by real-time reverse transcriptase polymerase chain reaction (RT-PCR) assays for SARS-CoV-2 nucleic acid with nasopharyngeal swab specimens were retrospectively enrolled in our hospital, and 44 patients were excluded. The inclusion criteria were as follows: (1) had an epidemiological history and diagnosed with SARS-CoV-2 infection, (2) at least three chest CT examined during illness. Exclusion criteria included (1) patients with lung tumors or any other causes of pneumonia, (2) chest CT with severe motion artifact, and (3) COVID-19 patients without pneumonia on all the chest CT during illness. Finally, 29 patients with a total of 113 baseline and follow-up CT scans were included in the study. A diagram illustrating the patient selection process is shown in Fig. 1. According to the Diagnosis and Treatment of Novel Coronavirus Pneumonia (trial version six) of China [8], the discharge criteria were as follows: (i) afebrile for more than 3 days, (ii) significant reduction in respiratory symptoms, (iii) acute exudative lesions with substantial improvements on chest radiograph or CT, and (iv) two consecutively negative COVID-19 nucleic acid tests at least 24 h apart. All of these patients were discharged for home isolation for 14 days. The clinical classifications were as follows: ordinary patients all had fever or respiratory symptoms with CT demonstrating pneumonia. Severe patients met any of the following conditions: (i) shortness of breath, respiratory rate ≥ 30 breaths/min; (ii) finger oxygen saturation ≤ 93% at a resting state; or (iii) oxygenation index (arterial oxygen tension/inspiratory oxygen fraction) ≤ 300 mmHg (1 mmHg = 0.133 kPa). Critical patients met any of the following conditions: (i) respiratory failure that required mechanical ventilation, (ii) shock, or (iii) other organ failure requiring intensive care unit monitoring and treatment. Patients with less than 3 CT scans and without pneumonia during the disease course were excluded.
Multiple CT scans were performed to evaluate the deterioration of the patients’ clinical conditions or discharge indications for patients with negative SARS-CoV-2 nucleic acid results by RT-PCR. Clinical symptoms and laboratory parameters were collected at baseline and at the follow-up time points. The clinical symptoms at baseline were collected no more than 9 days from the onset of the initial symptoms and collected at the same day as the CT scan at the other time points. The laboratory data were collected less than 2 days from the CT scan at each time point. The clinical and laboratory data specifically included symptoms, history of exposure, clinical classifications, white blood cell count, lymphocyte count, C-reactive protein, interleukin-6, and oxygenation index. In case of loss to follow-up, the laboratory data were processed as a defect.
CT protocol
Chest CT examinations were performed without the use of a contrast agent in the following two scanners: CT scanner (Brilliance iCT (128); Philips Healthcare) or CT (SOMATOMgo. Top; Siemens Healthcare). All patients were scanned from the apex to the base of the lung in the supine position during breath-holding using a peak voltage of 120 kVp with an automatic tube current (50~300 mAs). All scans were reconstructed using a high spatial frequency, iterative reconstruction (iDose 4, Philips), or a convolution kernel (BR60, Siemens) with a slice thickness of 1 mm as axial images.
Image interpretation
The images were analyzed independently in a consistent manner by two radiologists with 5 years of experience. In case of a disagreement, a final consensus was reached by a third radiologist with 10 years of experience in diagnosing cardiothoracic conditions. All CT images were analyzed and evaluated on a picture archiving and communication system (PACS), and the three radiologists had access to previous CT scans when evaluating each subsequent CT scan. The data collected from the analysis were as follows: (i) characteristics of the lesions (ground-glass opacity (GGO), mixed ground-glass opacity and consolidation, or consolidation), the number of pulmonary segments that were occupied by these lesions, and the number of involved pulmonary lobes. The following CT features were recorded: interlobular septal thickening, parenchymal bands, air bronchogram, pleural thickening, pleural effusion, and architectural distortion. All CT signs or characteristics were defined and described using the Fleischner Society glossary of terms [9]. The distribution patterns of the lesions were recorded as subpleural, peribronchovascular, random, or diffuse [10]. (ii) The changes in the number and/or size of the lesions detected by two consecutive CT scans were categorized as without lesions, increase in lesions, concurrent increase and decrease, or decrease in lesions. The CT density of the lesions in the same area on the baseline and follow-up CT scans, as well as the CT density of the erector spinae in the same image location as the lesion, was measured (the regions of interest ranged from 1 to 1.5 cm2 and attempted to include different densities of lesions). To ensure consistency, all measurements were performed two times by two radiologists at different image levels, and the average values were calculated. The lesion-to-muscle ratio (LMR) was calculated. In the case of the presence of more than one lung lesion per patient, one index lesion with the largest lesion range on the baseline CT scan was selected in consensus by the two radiologists. If there was no lesion on the baseline CT scan, the index lesion was selected from the follow-up CT scan. (iii) According to the evaluation criterion recommended by the expert group of radiologists of our city, the pneumonia inflammation index (PII) was calculated from each CT scan, and the PII calculation method was as follows: (1) the lesion distribution score had a maximum of 20 points. Per pulmonary anatomy, 1 point was scored for each of the 10 segments of the right lung and of segments 3 to 6, 9, and 10 of the left lung, while segments 1/2 and 8 of the left lung each scored 2 points when affected with pulmonary lesions. (2) The lesion size score had a maximum of 20 points and attributed 1 point to each pulmonary segment with lesions filling more than 50% of its volume. (3) The consolidation score had a maximum of 20 points and attributed 1 point to each pulmonary segment with large patch of consolidation. PII was a ratio that calculated as 100% times one-fortieth (1/40) of the sum of the lesion distribution score, the lesion size score, and the consolidation score. Any sum exceeding 40 was set to 40.
Statistical analysis
Statistical analyses were performed using SPSS Statistics Software (version 26; IBM). Quantitative variables are presented as the mean ± standard deviation (minimum-maximum) for normally distributed variables or interquartile ranges for non-normally distributed variables, and categorical data are reported as counts. The qualitative nominal variables were compared using the chi-square test or Fisher’s exact test. Comparisons of the quantitative data among the four time points were performed using the Kruskal-Wallis H test. Two groups of normally distributed quantitative variables were compared by Fischer’s least significant difference test. A p < 0.05 was defined as significant. Prism (5, GraphPad) was employed to create the bar graph. In cases of missing follow-up data, patients with missing data were excluded.
Results
Clinical and laboratory findings of the patients in four time points
A total of 29 patients (males 21, females 8) with confirmed COVID-19 were retrospectively enrolled in this study, with an average age of 44 ± 14 years (range, 14–73 years old). The clinical classifications were as follows: ordinary for 22 patients, severe for 6 patients, and critical for 1 patient. More demographic data are listed in Table 1. The baseline chest CT was obtained 3 (1–5) days after disease onset. Based on the quartiles of the duration from the baseline CT scan to the follow-up CT, the patients were assigned to four groups: 29 patients were assigned to baseline, 29 to 1–6 days, 27 to 7–13 days, and 28 to ≥ 14 days. The clinical and laboratory findings of the patients at the four time points are listed in Table 2. The most common symptoms of initial illness onset included fever (25/29) and cough (24/29). Many of the patients exhibited progressive shortness of breath on days 1–6 (11/29), and the percentage of patients with fever, cough, shortness of breath, and myalgia obviously decreased at 7–13 days (p < 0.05). The lymphocyte count, C-reactive protein, interleukin-6, and oxygenation index worsened within 1–6 days and improved sharply at 7–13 days (p < 0.05). A few laboratory data were missed, as follows: white blood cell count, lymphocyte count, and C-reactive protein of 4 patients at baseline. Interleukin-6 at the four time points were 6, 2, 3, and 2 patients, respectively; the oxygenation index was 6, 1, 0, and 5, respectively. The reasons for these missing laboratory data were not examined. Outpatients did not receive standardized laboratory tests at the beginning of illness, and some tests were unnecessary in the convalescent stage.
Time course of CT imaging findings
Twenty-nine patients had a total of 113 chest CT scans, and each patient had an average of 4 ± 1 (3–5) CT scans with a mean interval of 5 ± 3 (1–14) days. The most common features on chest CT were GGO at baseline and mixed GGO and consolidation within 1–6 days and 7–13 days, and the less common feature was consolidation (Fig. 2) (Fig. 3). Most lesions were found in subpleural or peribronchovascular locations. The changes in other imaging features are demonstrated in Fig. 4.
Table 3 summarizes the CT characteristics at the four time points. The LMR and PII showed typical time courses with increase from 1 to 6 days and subsequent decreases to lower than baseline levels for LMR and slightly higher than baseline values for PII after 14 days (p < 0.05) (Fig. 5). With an extension of the follow-up time points, the incidence of parenchymal bands and architectural distortion gradually increased (p < 0.05).
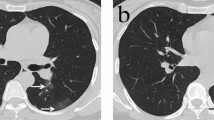
Typical changes on chest CT of COVID-19 pneumonia in a 47-year-old male with a 5-day history of fever. a Baseline CT scan demonstrated subpleural GGO in the bilateral upper lobes, a lesion-to-muscle ratio (LMR) = − 11.0, and pulmonary inflammation index (PII) = 37.5%. b On day 3 after the baseline scan, the lesions increased significantly in size, number, and density compared with those at baseline; the LMR = − 8.3, and PII = 65.0%. c On day 8, compared with that on day 3, the size of the lesion decreased, and the density increased, which was accompanied by the appearance of consolidation and thickening of the right oblique fissure; the LMR = − 7.6, and PII = 52.5%. d On day 12, the density of the lesions increased compared with that on day 8, and parenchymal bands formed in the left lobe; the LMR = − 10.0, and PII = 55.0%. e, f On day 17, compared with day 12, the size and number of the lesions decreased, but there were parenchymal bands and architectural distortion; the LMR = − 10.5, and PII = 38.0%. g On day 25, the lesions remained stable compared with those on day 17; the LMR = − 11.2, and PII = 38.0%
Discussion
The most important findings of our retrospective cohort study of 29 patients with COVID-19 were that the chest CT findings at follow-up reflect the clinical time course of COVID-19. GGO, mixed GGO, and consolidation were the major chest CT findings of COVID-19. Fibrosis changes could be seen during the remission stage.
Our results suggested that the most severe stage of illness was at 1–6 days. During this time point, the clinical symptoms worsened, the lymphocyte count decreased to the lowest level, and C-reactive protein increased to the highest level. A previous study revealed that lymphocytes and C-reactive protein could be used as important indexes in the evaluation of COVID-19 severity [11]. LMR and PII were the highest at 1–6 days compared with those at the other three time points. The increase in LMR was the result of increased lesion density, which indicated that the lesion transitioned from GGO to consolidation. Previous studies suggested that consolidation was related to the severity of COVID-19 pneumonia, and the transition from GGO to consolidation was an indicator of disease progression [11, 12].
All patients were hospitalized and received regular treatments, including oxygen therapy in different ways according to the severity of hypoxemia, supportive treatment, antiviral treatment, traditional Chinese medical therapy, antibiotic therapy if a bacterial infection was present, and corticosteroid therapy if necessary. Although thorough treatment was administered, most patients still exhibited illness progression from 1 to 6 days. This may be caused by the large number of viral replications at this time point; the immune cells overactivate and release a large number of cytokines, resulting in a cytokine storm, which is described as an exuberant immune responses following infection and is associated with excessive levels of proinflammatory cytokines and widespread tissue damage [13]. However, with regular treatment, it still takes time to build an immune response and produce antibodies to suppress virus replication. The time points used for grouping in our study did not include illness onset because the “baseline” examinations took place at different time points of the natural course of COVID-19 pneumonia in each patient, but the baseline CT scan was the time point for treatment initiation. Considering that the baseline chest CT was obtained 3 (1–5) days after the onset of symptoms in our cohort, we believe that the lung lesions might progress to a peak approximately 9 days after the onset of symptoms. However, a recent study from Wuhan, China, suggested that the peak time of COVID-19 pneumonia was approximately 2 weeks after illness onset [12]. It is currently unclear if the difference in timing relates to normal variations in the time course of COVID-19, to differences in patient management, or to other unrevealed reasons. We speculate that the reasons for the timing difference are as follows. First, the patients in our cohort were transferred from Wuhan or had second or higher generation SARS-CoV-2 infections with lower virulence than the initial hosts of pathogens, as pathogens tend to reduce their virulence to maximize their between-host transmission [14]. Second, the patients in our cohort received timely treatment.
At 7–13 days, the LMR decreased but remained higher than that at baseline, and the PII was slightly lower than that at 1–6 days. Compared with the changes in clinical and laboratory data, the improvements in imaging were less significant. We presume that a short plateau stage or a gradual decreasing trend for the lesions on chest CT may exist, and the improvements in imaging may occur slightly later than the changes in clinical and laboratory data. Therefore, imaging examinations may be unnecessary for patients when they show improvements clinically.
After 14 days of regular treatment, the pneumonia improved significantly on chest CT. Some patients had no remaining traces in their pulmonary tissue after lesion absorption. In most patients, there was parenchymal band formation and architectural distortion after lesion absorption. These features indicated the appearance of interstitial changes, suggesting the development of fibrosis that could be seen during the remission stage [12, 15].
There are some limitations in our study. First, the patients in our study were mainly considered ordinary or severe clinically, with only one critical case. However, some clinically critical patients may have different time courses from those seen in this study. Further studies with larger cohorts of more clinically critical patients are required. Second, the time from illness onset to the baseline CT scan was variable, and there were some patients with missing laboratory data, which are sources of bias and ambiguity in the assessment of the disease process. Despite these shortcomings, it appeared that the time course of the chest CT findings followed the time course of the serum markers of inflammation.
In conclusion, our findings indicate that the time course of CT imaging changes is consistent with the clinical time course of COVID-19. Our work described in detail the clinical and imaging changes of clinically ordinary and severe COVID-19 patients after 2 weeks and revealed that the lung lesions progressed within 1–6 days, followed by clinical improvement and resorption of the lesions after treatment. This work may help to provide evidence that CT imaging may be indicated when patients fail to improve within a week of treatment, but repeated chest CT may be unnecessary when patients show clinical improvements.
Abbreviations
- COVID-19:
-
Coronavirus disease 2019
- CT:
-
Computed tomography
- GGO:
-
Ground-glass opacity
- LMR:
-
Lesion-to-muscle ratio
- PII:
-
Pneumonia inflammation index
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
References
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395:497–506
Hui DS, Azhar EI, Madani TA et al (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health — the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 91:264–266
Lu H, Stratton CW, Tang YW (2020) Outbreak of pneumonia of unknown etiology in Wuhan, China: the mystery and the miracle. J Med Virol 92:401–402
Wu P, Hao X, Lau EHY et al (2020) Real-time tentative assessment of the epidemiological characteristics of novel coronavirus infections in Wuhan, China, as at 22 January 2020. Euro Surveill 25:2000044
Lu R, Zhao X, Li J et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395:565–574
Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J (2020) Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. https://doi.org/10.1148/radiol.2020200343
Zu ZY, Jiang MD, Xu PP et al (2020) Coronavirus disease 2019 (COVID-19): a perspective from China. Radiology. https://doi.org/10.1148/radiol.2020200490
China National Health Commission (2020) Diagnosis and treatment of pneumonitis caused by new coronavirus (trial version 6). In: Beijing China Natl. Heal. Comm http://www.nhc.gov.cn/yzygj/s7653p/202002/8334a8326dd94d329df351d7da8aefc2.shtml. Accessed 19 Feb 2020
Hansell DM, Bankier AA, MacMahon H, McLoud TC, Müller NL, Remy J (2008) Fleischner society: glossary of terms for thoracic imaging. Radiology 246:697–722
Pan F, Ye T, Sun P et al (2020) Time course of lung changes on chest CT during recovery from 2019 novel coronavirus (COVID-19) pneumonia. Radiology 295:715–721. https://doi.org/10.1148/radiol.2020200370
Li K, Wu J, Wu F et al (2020) The clinical and chest CT features associated with severe and critical COVID-19 pneumonia. Invest Radiol 55:327–331
Shi H, Han X, Jiang N et al (2020) Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: a descriptive study. Lancet Infect Dis 20:425–434
Fung SY, Yuen KS, Ye ZW, Chan CP, Jin DY (2020) A tug-of-war between severe acute respiratory syndrome coronavirus 2 and host antiviral defence: lessons from other pathogenic viruses. Emerg Microbes Infect 9:558–570
Yang W, Cao Q, Qin L et al (2020) Clinical characteristics and imaging manifestations of the 2019 novel coronavirus disease (COVID-19):a multi-center study in Wenzhou city, Zhejiang, China. J Infect 80:388–393
Xiong Y, Sun D, Liu Y et al (2020) Clinical and high-resolution CT features of the COVID-19 infection. Invest Radiol 55:332–339
Acknowledgements
The authors would like to acknowledge the Chongqing Radiology Association of China for provide the calculation method of pulmonary inflammation index.
Funding
This work is supported by the Chongqing Health Commission of China (No. 2016MSXM053) and Chongqing science and Technology Commission (cstc2016jcyjAo217).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Yongxia Zhou.
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study/cross sectional study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, Y., Zheng, Y., Yang, Q. et al. Cohort study of chest CT and clinical changes in 29 patients with coronavirus disease 2019 (COVID-19). Eur Radiol 30, 6213–6220 (2020). https://doi.org/10.1007/s00330-020-07007-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-020-07007-0