Abstract
Objectives
To distinguish squamous cell carcinoma (SCC) from lung adenocarcinoma (ADC) based on a radiomic signature
Methods
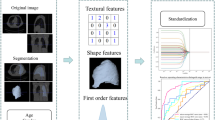
This study involved 129 patients with non-small cell lung cancer (NSCLC) (81 in the training cohort and 48 in the independent validation cohort). Approximately 485 features were extracted from a manually outlined tumor region. The LASSO logistic regression model selected the key features of a radiomic signature. Receiver operating characteristic curve and area under the curve (AUC) were used to evaluate the performance of the radiomic signature in the training and validation cohorts.
Results
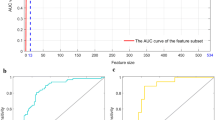
Five features were selected to construct the radiomic signature for histologic subtype classification. The performance of the radiomic signature to distinguish between lung ADC and SCC in both training and validation cohorts was good, with an AUC of 0.905 (95% confidence interval [CI]: 0.838 to 0.971), sensitivity of 0.830, and specificity of 0.929. In the validation cohort, the radiomic signature showed an AUC of 0.893 (95% CI: 0.789 to 0.996), sensitivity of 0.828, and specificity of 0.900.
Conclusions
A unique radiomic signature was constructed for use as a diagnostic factor for discriminating lung ADC from SCC. Patients with NSCLC will benefit from the proposed radiomic signature.
Key points
• Machine learning can be used for auxiliary distinguish in lung cancer.
• Radiomic signature can discriminate lung ADC from SCC.
• Radiomics can help to achieve precision medical treatment.





Similar content being viewed by others
Abbreviations
- ADC:
-
Adenocarcinoma
- AUC:
-
Area Under the Curve
- CI:
-
Confidence Interval
- CT:
-
Computed Tomography
- LASSO:
-
Least Absolute Shrinkage and Selection Operator
- NSCLC:
-
Non-small Cell Lung Cancer
- PET:
-
Positron Emission Tomography
- ROC:
-
Receiver Operating Characteristic
- ROI:
-
Region of Interest
- SCC:
-
Squamous Cell Carcinoma
References
Printz C (2015) Lung cancer new leading cause of death for women in developed countries: Data reflects increased rates of smoking. Cancer 121:1911–1912
Brawley OW (2011) Avoidable cancer deaths globally. CA Cancer J Clin 61:67–68
Ferlay J, Steliarovafoucher E, Lortettieulent J et al (2013) Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. Eur J Cancer 49:1374–1403
Nair VS, Gevaert O, Davidzon G et al (2012) Prognostic PET 18F-FDG uptake imaging features are associated with major oncogenomic alterations in patients with resected non-small cell lung cancer. Cancer Res 72:3725–3734
Siegel RL, Miller KD, Jemal A (2017) Cancer Statistics, 2017. CA Cancer J Clin. 67:7–30
Fukui T, Taniguchi T, Kawaguchi K et al (2015) Comparisons of the clinicopathological features and survival outcomes between lung cancer patients with adenocarcinoma and squamous cell carcinoma. Gen Thorac Cardiovasc Surg 63:507–513
Bodendorf MO, Haas V, Laberke HG (2009) Prognostic value and therapeutic conse-quences of vascular invasion in non-small cell lung carcinoma. Lung Cancer 64:71–78
Usui S, Minami Y, Shiozawa T et al (2013) Differences in the prognostic implications of vascular invasion between lung adenocarcinoma and squamous cell carcinoma. Lung Cancer 82:407–412
Herbst RS, O'Neill VJ, Fehrenbacher L et al (2007) Phase II study of efficacy and safety of bevacizumab in combination with chemotherapy or erlotinib compared with chemotherapy alone for treatment of recurrent or refractory non small-cell lung cancer. J Clin Oncol 25:4743–4750
Di Costanzo F, Mazzoni F, Micol Mela M et al (2008) Bevacizumab in non-small cell lung cancer. Drugs 68:737–746
Nguyenkim TD, Frauenfelder T, Strobel K et al (2015) Assessment of Bronchial and Pulmonary Blood Supply in Non-Small Cell Lung Cancer Subtypes Using Computed Tomography Perfusion. Investig Radiol 50:179–186
Yip C, Landau D, Kozarski R et al (2014) Primary esophageal cancer: heterogeneity as potential prognostic biomarker in patients treated with definitive chemotherapy and radiation therapy. Radiology 270:141–148
Ng F, Ganeshan B, Kozarski R, Miles KA, Goh V (2013) Assessment of primary colorectal cancer heterogeneity by using whole-tumor texture analysis: contrast-enhanced CT texture as a biomarker of 5-year survival. Radiology 266:177–184
Ganeshan B, Goh V, Mandeville HC, Ng QS, Hoskin PJ, Miles KA (2013) Non-small cell lung cancer: histopathologic correlates for texture parameters at CT. Radiology 266:326–336
Ganeshan B, Abaleke S, Young RC, Chatwin CR, Miles KA (2010) Texture analysis of non-small cell lung cancer on unenhanced computed tomography: initial evidence for a relationship with tumour glucose metabolism and stage. Cancer Imaging 10:137–143
Ganeshan B, Skogen K, Pressney I, Coutroubis D and MilesK (2012) Tumour heterogeneity in oesophageal cancer assessed by CT texture analysis: preliminary evidence of an association with tumour metabolism, stage, and survival. Clin Radiol 67:157-164.
Lambin P, Rios-Velazquez E, Leijenaar R et al (2012) Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer 48:441–446
Kumar V, Gu Y, Basu S et al (2012) Radiomics:the process and the challenges. Magn Reson Imaging 30:1234–1248
Choi ER, Lee HY, Jeong JY et al (2016) Quantitative image variables reflect the intratumoral pathologic heterogeneity of lung adenocarcinoma. Oncotarget 7:67302–67313
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: Images Are More than Pictures,They Are Data. Radiology 278:563–577
Aerts HJ, Velazquez ER, Leijenaar RT et al (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006
Itakura H, Achrol AS, Mitchell LA et al (2015) Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci Transl Med 7:303ra138
Li H, Zhu Y, Burnside ES et al (2016) MR Imaging Radiomics Signatures for Predicting the Risk of Breast Cancer Recurrence as Given by Research Versions of MammaPrint, Oncotype DX, and PAM50 Gene Assays. Radiology 281:382–391
Zinn PO, Singh SK, Kotrotsou A et al (2016) 139 Clinically Applicable and Biologically Validated MRI Radiomic Test Method Predicts Glioblastoma Genomic Landscape and Survival. Neurosurgery 63:156–157
Huang Y, Liu Z, He L et al (2016) Radiomics Signature: A Potential Biomarker for the Prediction of Disease-Free Survival in Early-Stage (I or II) Non-Small Cell Lung Cancer. Radiology 281:152234
Yamamoto S, Korn RL, Oklu R et al (2014) ALK Molecular Phenotype in Non–Small Cell Lung Cancer: CT Radiogenomic Characterization. Radiology 272:568–576
Gevaert O, Xu J, Hoang CD et al (2012) Non-small cell lung cancer: identifying prognostic imaging biomarkers by leveraging public gene expression microarray data--methods and preliminary results. Radiology 264:387–396
Parmar C, Lambin P, Aerts HJ et al (2015) Radiomic feature clusters and prognostic signatures specific for Lung and Head and Neck cancer. Sci Rep 5:11044
Chi MA, Jian H (2016) Asymptotic properties of LASSO in high-dimensional partially linear models. Science China Mathematics 59:1–20
Rao SJ (2003) Regression Modeling Strategies: With Applications to Linear Models, Logistic Regression, and Survival Analysis. J Am Stat Assoc 98:257–258
Lloyd CJ (2015) Theory & Methods: Fitting Roc Curves Using Non-linear Binomial Regression. Aust. N Z J Stat 42:193–204
Ball DL, Fisher RJ, Burmeister BH et al (2013) The complex relationship between lung tumor volume and survival in patients with non-small cell lung cancer treated by definitive radiotherapy: A prospective, observational prognostic factor study of the Trans-Tasman Radiation Oncology Group (TROG 99.05). Radiother Oncol 106:305–311
Khayyata S, Yun S, Pasha T et al (2009) Value of P63 and CK5/6 in distinguishing squamous cell carcinoma from adenocarcinoma in lung fine-needle aspiration specimens. Diagn Cytopathol 37:178–183
Montezuma D, Azevedo R, Lopes P et al (2013) A panel of four immunohistochemical markers (CK7, CK20, TTF-1, and p63) allows accurate diagnosis of primary and metastatic lung carcinoma on biopsy specimens. Virchows Arch 463:749–754
Wang XN, Wang SJ, Pandey V et al (2015) Trefoil factor 3 as a novel biomarker to distinguish between adenocarcinoma and squamous cell carcinoma. Medicine 94:e860
Patnaik S, Mallick R, Kannisto E et al (2015) MiR-205 and MiR-375 MicroRNA Assays to Distinguish Squamous Cell Carcinoma from Adenocarcinoma in Lung Cancer Biopsies. J Thorac Oncol 10:446–453
Ha S, Choi H, Cheon GJ et al (2014) Autoclustering of Non-small Cell Lung Carcinoma Subtypes on 18F-FDG PET Using Texture Analysis: A Preliminary Result. Nucl Med Mol Imaging 48:278–286
Funding
This work was supported by the National Natural Science Foundation of China (81227901, 81771924, 81501616, 61231004, 81671851, and 81527805), National Key R&D Program of China (2017YFA0205200, 2017YFC1308700, 2017YFC1308701, 2017YFC1309100), the Science and Technology Service Network Initiative of the Chinese Academy of Sciences (KFJ-SW-STS-160), the Instrument Developing Project of the Chinese Academy of Sciences (YZ201502), the Beijing Municipal Science and Technology Commission (Z161100002616022), and the Youth Innovation Promotion Association CAS.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Jie Tian.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was obtained from all subjects (patients) in this study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Electronic supplementary material
ESM 1
(DOC 82 kb)
Rights and permissions
About this article
Cite this article
Zhu, X., Dong, D., Chen, Z. et al. Radiomic signature as a diagnostic factor for histologic subtype classification of non-small cell lung cancer. Eur Radiol 28, 2772–2778 (2018). https://doi.org/10.1007/s00330-017-5221-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-017-5221-1