Abstract
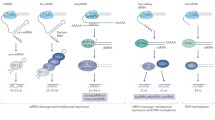
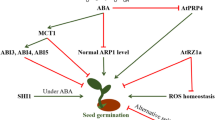
Seeds represent a crucial stage of the seed plants life cycle. It is during seed development that the foundations of the future plant body, and the ability to give rise to a new plant capable of growing under sometimes adverse environmental conditions, are established. Small non-coding RNAs are major regulators of gene expression both at the post-transcriptional and transcriptional levels and, not surprisingly, these elements play major roles in seed development and germination. We review here the current knowledge about small RNA expression and functions in seed development, going from the morphogenesis phase comprehending embryo development and patterning, to the several steps of the maturation phase, ending in the transition to the germination. A special focus is given to the small RNAs for which functional studies have been conducted and their participation in regulatory networks operating in seeds. Many challenges remain ahead for dissecting the complex small RNA landscape in seeds, but this is a highly relevant issue in plant biology and advances in this area will most certainly impact plant breeding.

Similar content being viewed by others
References
Agarwal P, Kapoor S, Tyagi AK (2011) Transcription factors regulating the progression of monocot and dicot seed development. Bioessays 33:189–202. doi:10.1002/bies.201000107
Allen RS, Li J, Stahle MI, Dubroué A, Gubler F, Millar AA (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci USA 104(41):16371–16376. doi:10.1073/pnas.0707653104
Allen RS, Li J, Alonso-Peral MM, White RG, Gubler F, Millar AA (2010) MicroR159 regulation of most conserved targets in Arabidopsis has negligible phenotypic effects. Silence 1(1):18. doi:10.1186/1758-907X-1-18
Alon U (2007) Network motifs: theory and experimental approaches. Nat Rev Genet 8(6):450–461. doi:10.1038/nrg2102
Alonso-Peral MM, Li J, Li Y, Allen RS, Schnippenkoetter W, Ohms S, White RG, Millar AA (2010) The microRNA159-regulated GAMYB-like genes inhibit growth and promote programmed cell death in Arabidopsis. Plant Physiol 154(2):757–771. doi:10.1104/pp.110.160630
Alonso-Peral MM, Sun C, Millar AA (2012) MicroRNA159 can act as a switch or tuning microRNA independently of its abundance in Arabidopsis. PLoS One 7(4):e34751. doi:10.1371/journal.pone.0034751
Arikit S, Xia R, Kakrana A, Huang K, Zhai J, Yan Z, Valdés-López O, Prince S, Musket TA, Nguyen HT, Stacey G, Meyers BC (2014) An atlas of soybean small RNAs identifies phased siRNAs from hundreds of coding genes. Plant Cell 26(12):4584–4601. doi:10.1105/tpc.114.131847
Axtell MJ (2013) Classification and comparison of small RNAs from plants. Annu Rev Plant Biol 64:137–159. doi:10.1146/annurev-arplant-050312-120043
Borges F, Martienssen RA (2015) The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol 16:727–741. doi:10.1038/nrm4085
Breakfield NW, Corcoran DL, Petricka JJ, Shen J, Sae-Seaw J, Rubio-Somoza I, Weigel D, Ohler U, Benfey PN (2012) High-resolution experimental and computational profiling of tissue-specific known and novel miRNAs in Arabidopsis. Genome Res 22(1):163–176. doi:10.1101/gr.123547.111
Cao S, Zhu QH, Shen W, et al. (2013) Comparative profiling of miRNA expression in developing seeds of high linoleic and high oleic safflower (Carthamus tinctorius L.) plants. Front Plant Sci 4. doi:10.3389/fpls.2013.00489
Chávez-Montes RA, de Fátima Rosas-Cárdenas F, De Paoli E, Accerbi M, Rymarquis LA, Mahalingam G, Marsch-Martínez N, Meyers BC, Green PJ, de Folter S (2014) Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs. Nat Commun 5:3722. doi:10.1038/ncomms4722
Chellappan P, Xia J, Zhou X, Gao S, Zhang X, Coutino G, Vazquez F, Zhang W, Jin H (2010) siRNAs from miRNA sites mediate DNA methylation of target genes. Nucleic Acids Res 38(20):6883–6894. doi:10.1093/nar/gkq590
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 35:21–44. doi:10.1146/annurev.cellbio.042308.113417
Chou CH et al (2016) miRTarBase 2016: updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Res 44(D1):D239–D247. doi:10.1093/nar/gkv1258
Curaba J, Spriggs A, Taylor J, Li Z, Helliwell C (2012) miRNA regulation in the early development of barley seed. BMC Plant Biol 12:120. doi:10.1186/1471-2229-12-120
Devic M, Roscoe T (2016) Seed maturation: simplification of control networks in plants. Plant Sci 252:335–346. doi:10.1016/j.plantsci.2016.08.012
Dolgosheina EV, Morin RD, Aksay G, Sahinalp SC, Magrini V, Mardis ER, Mattsson J, Unrau PJ (2008) Conifers have a unique small RNA silencing signature. RNA 14(8):1508–1515. doi:10.1261/rna.1052008
Dong Z, Han MH, Fedoroff N (2008) The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc Natl Acad Sci USA 105(29):9970–9975. doi:10.1073/pnas.0803356105
Galli V, Guzman F, de Oliveira LF, Loss-Morais G, Körbes AP, Silva SD, Margis-Pinheiro MM, Margis R (2014) Identifying microRNAs and transcript targets in Jatropha seeds. PLoS One 9(2):e83727. doi:10.1371/journal.pone.0083727
Giraudat J, Hauge BM, Valon C, Smalle J, Parcy F, Goodman HM (1992) Isolation of the Arabidopsis ABI3 gene by positional cloning. Plant Cell 4(10):1251–1261. doi:10.1105/tpc.4.10.1251
Goldberg RB, Depaiva G, Yadegari R (1994) Plant embryogenesis—zygote to seed. Science 266:605–614. doi:10.1126/science.266.5185.605
Grigg SP, Galinha C, Kornet N, Canales C, Scheres B, Tsiantis M (2009) Repression of apical homeobox genes is required for embryonic root development in Arabidopsis. Curr Biol 19(17):1485–1490. doi:10.1016/j.cub.2009.06.070
Gutiérrez L, Van Wuytswinkel O, Castelain M, Bellini C (2007) Combined networks regulating seed maturation. Trends Plant Sci 12(7):294–300. doi:10.1016/j.tplants.2007.06.003
Hobert O (2008) Gene regulation by transcription factors and microRNAs. Science 319(5871):1785–1786. doi:10.1126/science.1151651
Hove CA, Lu KJ, Weijers D (2015) Building a plant: cell fate specification in the early Arabidopsis embryo. Development 142(3):420–430. doi:10.1242/dev.111500
Huang D, Koh C, Feurtado JA, Tsang EW, Cutler AJ (2013) MicroRNAs and their putative targets in Brassica napus seed maturation. BMC Genom 14:140. doi:10.1186/1471-2164-14-140
Huo H, Wei S, Bradford KJ (2016) DELAY OF GERMINATION1 (DOG1) regulates both seed dormancy and flowering time through microRNA pathways. Proc Natl Acad Sci USA 113(15):E2199–E2206. doi:10.1073/pnas.1600558113
Junker A, Hartmann A, Schreiber F, Bäumlein H (2010) An engineer’s view on regulation of seed development. Trends Plant Sci 15(6):303–307. doi:10.1016/j.tplants.2010.03.005
Kang M, Zhao Q, Zhu D, Yu J (2012) Characterization of microRNAs expression during maize seed development. BMC Genom 13:360. doi:10.1186/1471-2164-13-360
Knauer S, Holt AL, Rubio-Somoza I, Tucker EJ, Hinze A, Pisch M, Javelle M, Timmermans MC, Tucker MR, Laux T (2013) A protodermal miR394 signal defines a region of stem cell competence in the Arabidopsis shoot meristem. Dev Cell 24(2):125–132. doi:10.1016/j.devcel.2012.12.009
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42:D68–D73. doi:10.1093/nar/gkt1181
Lan Y, Su N, Shen Yi et al (2012) Identification of novel MiRNAs and MiRNA expression profiling during grain development in indica rice. BMC Genom 13:264
Lee EK, Cibrian-Jaramillo A, Kolokotronis SO, Katari MS, Stamatakis A, Ott M, Chiu JC, Little DP, Stevenson DW, McCombie WR, Martienssen RA, Coruzzi G, Desalle R (2011) A functional phylogenomic view of the seed plants. PLoS Genet 7(12):e1002411. doi:10.1371/journal.pgen.1002411
Li D, Wang L, Liu X, Cui D, Chen T, Zhang H, Jiang C, Xu C, Li P, Li S, Zhao L, Chen H (2013) Deep sequencing of maize small RNAs reveals a diverse set of microRNA in dry and imbibed seeds. PLoS One 8(1):e55107. doi:10.1371/journal.pone.0055107
Li T, Ma L, Geng Y, Hao C, Chen X, Zhang X (2015) Small RNA and degradome sequencing reveal complex roles of miRNAs and their targets in developing wheat grains. PLoS One 10(10):e0139658. doi:10.1371/journal.pone.0139658
Li ZX, Li SG, Zhang L et al (2016) Over-expression of miR166a inhibits cotyledon formation in somatic embryos and promotes lateral root development in seedlings of Larix leptolepis. Plant Cell Tiss Organ Cult 127:461. doi:10.1007/s11240-016-1071-9
Liu PP, Montgomery TA, Fahlgren N, Kasschau KD, Nonogaki H, Carrington JC (2007) Repression of AUXIN RESPONSE FACTOR10 by microRNA160 is critical for seed germination and post-germination stages. Plant J 52(1):133–146. doi:10.1111/j.1365-313X.2007.03218.x
Liu X, Huang J, Wang Y et al (2010) The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in MicroRNA160a, in organogenesis and the mechanism regulating its expression. Plant J 62:416–428. doi:10.1111/j.1365-313X.2010.04164.x
Lobbes D, Rallapalli G, Schmidt DD, Martin C, Clarke J (2006) SERRATE: a new player on the plant microRNA scene. EMBO Rep 7(10):1052–1058. doi:10.1038/sj.embor.7400806
Lotan T, Ohto M, Yee KM, West MA, Lo R, Kwong RW, Yamagishi K, Fischer RL, Goldberg RB, Harada JJ (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93(7):1195–1205. doi:10.1016/S0092-8674(00)81463-4
Lu J, Zhang C, Baulcombe DC, Chen ZJ (2012) Maternal siRNAs as regulators of parental genome imbalance and gene expression in endosperm of Arabidopsis seeds. Proc Natl Acad Sci USA 109(14):5529–5534. doi:10.1073/pnas.1203094109
Luerssen H, Kirik V, Herrmann P, Miséra S (1998) FUSCA3 encodes a protein with a conserved VP1/AB13-like B3 domain which is of functional importance for the regulation of seed maturation in Arabidopsis thaliana. Plant J 15(6):755–764. doi:10.1046/j.1365-313X.1998.00259.x
Luo YC, Zhou H, Li Y, Chen JY, Yang JH, Chen YQ, Qu LH (2006) Rice embryogenic calli express a unique set of microRNAs, suggesting regulatory roles of microRNAs in plant post-embryogenic development. FEBS Lett 580(21):5111–5116. doi:10.1016/j.febslet.2006.08.046
Mahdavi-Darvari F, Noor NM, Ismanizan I (2015) Epigenetic regulation and gene markers as signals of early somatic embryogenesis. Plant Cell Tiss Organ Cult 120:407. doi:10.1007/s11240-014-0615-0
Mallory AC, Vaucheret H (2006) Functions of microRNAs and related small RNAs in plants. Nat Genet 38(Suppl):S31–S36. doi:10.1038/ng1791
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14:1035–1046. doi:10.1016/j.cub.2004.06.022
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17(5):1360–1375. doi:10.1105/tpc.105.031716
Margis R, Fusaro AF, Smith NA, Curtin SJ, Watson JM, Finnegan EJ, Waterhouse PM (2006) The evolution and diversification of Dicers in plants. FEBS Lett 580(10):2442–2450. doi:10.1016/j.febslet.2006.03.072
Meinke D (1995) Molecular genetics of embryogenesis. Annu Rev Plant Physiol Plant Mol Biol 46:369–394
Meng Y, Shao C, Chen M (2011a) Toward microRNA-mediated gene regulatory networks in plants. Brief Bioinform 12:645–659. doi:10.1093/bib/bbq091
Meng Y, Shao C, Gou L, Jin Y, Chen M (2011b) Construction of MicroRNA- and MicroRNA*-mediated regulatory networks in plants. RNA Biol 8(6):1124–1148. doi:10.4161/rna.8.6.17743
Miguel CM, Rupps A, Raschke J, Rodrigues AS, Trontin J-F (2016) Impact of molecular studies on somatic embryogenesis development for implementation in conifer multi-varietal forestry. In: Park YS, Bonga JM, Moon HK (eds) Vegetative Propagation of Forest trees. National Institute of Forest Science, Seoul, pp 373–421
Miyashima S, Honda M, Hashimoto K et al (2013) A comprehensive expression analysis of the Arabidopsis MICRORNA165/6 gene family during embryogenesis reveals a conserved role in meristem specification and a non-cell-autonomous function. Plant Cell Physiol 54(3):375–384. doi:10.1093/pcp/pcs188
Morin RD, Aksay G, Dolgosheina E, Ebhardt HA, Magrini V, Mardis ER, Sahinalp SC, Unrau PJ (2008) Comparative analysis of the small RNA transcriptomes of Pinus contorta and Oryza sativa. Genome Res 18(4):571–584. doi:10.1101/gr.6897308
Mosher RA, Melnyk CW, Kelly KA, et al. (2009) Uniparental expression of PolIV-dependent siRNAs in developing endosperm of Arabidopsis. Nat Lett. doi:10.1038/nature08084
Nambara E, Nonogaki H (2012) Seed biology in the 21st century: perspectives and new directions. Plant Cell Physiol 53(1):1–4. doi:10.1093/pcp/pcr184
Niu SH, Liu C, Yuan HW, Li P, Li Y, Li W (2015) Identification and expression profiles of sRNAs and their biogenesis and action-related genes in male and female cones of Pinus tabuliformis. BMC Genom 16:693. doi:10.1186/s12864-015-1885-6
Nodine MD, Bartel DP (2010) MicroRNAs prevent precocious gene expression and enable pattern formation during plant embryogenesis. Genes Dev 24(23):2678–2692. doi:10.1101/gad.1986710
Nonogaki H (2010) MicroRNA gene regulation cascades during early stages of plant development. Plant Cell Physiol 51(11):1840–1846. doi:10.1093/pcp/pcq154
Nystedt B et al (2013) The Norway spruce genome sequence and conifer genome evolution. Nature 497(7451):579–584. doi:10.1038/nature12211
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263. doi:10.1038/nature01958
Park W, Li J, Song R, Messing J, Chen X (2002) CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 12(17):1484–1495. doi:10.1016/S0960-9822(02)01017-5
Park MY, Wu G, Gonzalez-Sulser A, Vaucheret H, Poethig RS (2005) Nuclear processing and export of microRNAs in Arabidopsis. Proc Natl Acad Sci USA 102:3691–3696. doi:10.1073/pnas.0405570102
Peng T, Sun H, Du Y, Zhang J, Li J et al (2013) Characterization and expression patterns of microRNAs involved in rice grain filling. PLoS One 8(1):e54148. doi:10.1371/journal.pone.0054148
Prigge MJ, Otsuga D, Alonso JM, Ecker JR, Drews GN, Clark SE (2005) Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17(1):61–76. doi:10.1105/tpc.104.026161
Rodrigues JA, Ruan R, Nishimura T et al (2013) Imprinted expression of genes and small RNA is associated with localized hypomethylation of the maternal genome in rice endosperm. Proc Natl Acad Sci USA 110(19):7934–7939. doi:10.1073/pnas.1306164110
Reyes JL, Chua NH (2007) ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J 49(4):592–606. doi:10.1111/j.1365-313X.2006.02980.x
Schallau A, Kakhovskaya I, Tewes A (2008) Phylogenetic footprints in fern spore- and seed-specific gene promoters. Plant J 53:414–424. doi:10.1111/j.1365-313X.2007.03354.x
Schwartz BW, Yeung EC, Meinke DW (1994) Disruption of morphogenesis and transformation of the suspensor in abnormal suspensor mutants of Arabidopsis. Development 120:3235–3245.
Seefried WF, Willmann MR, Clausen RL, Jenik PD (2014) Global regulation of embryonic patterning in arabidopsis by MicroRNAs. Plant Physiol 165(2):670–687. doi:10.1104/pp.114.240846
Song QX, Liu YF, Hu XY, Zhang WK, Ma B, Chen SY, Zhang JS (2011) Identification of miRNAs and their target genes in developing soybean seeds by deep sequencing. BMC Plant Biol 11:5. doi:10.1186/1471-2229-11-5
Sreenivasulu N, Wobus U (2013) Seed-development programs: a systems biology-based comparison between dicots and monocots. Annu Rev Plant Biol 64:189–217. doi:10.1146/annurev-arplant-050312-120215
Stone SL, Kwong LW, Yee KM, Pelletier J, Lepiniec L, Fischer RL, Goldberg RB, Harada JJ (2001) LEAFY COTYLEDON2 encodes a B3 domain transcription factor that induces embryo development. Proc Natl Acad Sci USA 98(20):11806–11811. doi:10.1073/pnas.201413498
Takada S, Jürgens G (2007). Transcriptional regulation of epidermal cell fate in the Arabidopsis embryo. Development 134(6):1141–1150. doi:10.1242/dev.02803
Tang X, Bian S, Tang M, Lu Q, Li S et al (2012) MicroRNA-mediated repression of the seed maturation program during vegetative development in Arabidopsis. PLoS Genet 8(11):e1003091. doi:10.1371/journal.pgen.1003091
Tuteja JH, Zabala G, Varala K, Hudson M, Vodkin LO (2009) Endogenous, tissue-specific short interfering RNAs silence the chalcone synthase gene family in Glycine max seed coats. Plant Cell 21(10):3063–3077. doi:10.1105/tpc.109.069856
Tyson JJ, Novák B (2010) Functional motifs in biochemical reaction networks. Annu Rev Phys Chem 61:219–240. doi:10.1146/annurev.physchem.012809.103457
Vaucheret H (2008) Plant ARGONAUTES. Trends Plant Sci 13(7):350–358. doi:10.1016/j.tplants.2008.04.007
Vazquez F, Blevins T, Ailhas J, Boller T, Meins F (2008) Evolution of Arabidopsis MIR genes generates novel microRNA classes. Nucleic Acids Res 36(20):6429–6438. doi:10.1093/nar/gkn670
Vialette-Giraud ACM, Andres-Robin A, Chambrier P, Tavares R, Scutt CP (2016) The analysis of gene regulatory networks in plant evo-devo. J Exp Bot 67(9):2549–2563. doi:10.1093/jxb/erw119
Wan LC, Wang F, Guo X et al (2012) Identification and characterization of small non-coding RNAs from Chinese fir by high throughput sequencing. BMC Plant Biol 12:146
Wang F, Perry SE (2013) Identification of direct targets of FUSCA3, a key regulator of Arabidopsis seed development. Plant Physiol 161(3):1251–1264. doi:10.1104/pp.112.212282
Wang CY, Zhang S, Yu Y et al (2014) MiR397b regulates both lignin content and seed number in Arabidopsis via modulating a laccase involved in lignin biosynthesis. Plant Biotechnol J 12(8):1132–1142. doi:10.1111/pbi.12222
Wang J, Jian H, Wang T et al (2016) Identification of microRNAs actively involved in fatty acid biosynthesis in developing Brassica napus seeds using high-throughput sequencing. Front Plant Sci 7:1570. doi:10.3389/fpls.2016.01570
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132:3657–3668.
Willmann MR, Mehalick AJ, Packer RL, Jenik PD (2011) MicroRNAs regulate the timing of embryo maturation in Arabidopsis. Plant Physiol 155(4):1871–1884. doi:10.1104/pp.110.171355
Wobus U, Weber H (1999) Seed maturation: genetic programmes and control signals. Curr Opin Plant Biol 2(1):33–38
Wu L, Zhou H, Zhang Q, Zhang J, Ni F, Liu C, Qi Y (2010) DNA methylation mediated by a microRNA pathway. Mol Cell 38(3):465–475. doi:10.1016/j.molcel.2010.03.008
Wu XM, Liu MY, Ge XX, Xu Q, Guo WW (2011) Stage and tissue-specific modulation of ten conserved miRNAs and their targets during somatic embryogenesis of Valencia sweet orange. Planta 233(3):495–505. doi:10.1007/s00425-010-1312-9
Xin M, Yang R, Yao Y, Ma C, Peng H, Sun Q, Wang X, Ni Z (2014) Dynamic parent-of-origin effects on small interfering RNA expression in the developing maize endosperm. BMC Plant Biol 14:192. doi:10.1186/s12870-014-0192-8
Xin M, Yang G, Yao Y, Peng H, Hu Z, Sun Q, Wang X, Ni Z (2015) Temporal small RNA transcriptome profiling unraveled partitioned miRNA expression in developing maize endosperms between reciprocal crosses. Front Plant Sci 6:744. doi:10.3389/fpls.2015.00744
Yang L, Liu Z, Lu F, Dong A, Huang H (2006) SERRATE is a novel nuclear regulator in primary microRNA processing in Arabidopsis. Plant J 47(6):841–850. doi:10.1111/j.1365-313X.2006.02835.x
Yang SW, Chen HY, Yang J, Machida S, Chua NH, Yuan YA (2010) Structure of Arabidopsis HYPONASTIC LEAVES1 and its molecular implications for miRNA processing. Structure 18(5):594–605. doi:10.1016/j.str.2010.02.006
Yang X, Wang L, Yuan D, Lindsey K, Zhang X (2013) Small RNA and degradome sequencing reveal complex miRNA regulation during cotton somatic embryogenesis. J Exp Bot 64(6):1521–1536. doi:10.1093/jxb/ert013
Yi R, Zhu Z, Hu J, Qian Q, Dai J et al (2013) Identification and expression analysis of microRNAs at the grain filling stage in rice (Oryza sativa L.) via deep sequencing. PLoS One 8(3):e57863. doi:10.1371/journal.pone.0057863
Yoshida S, Barbier de Reuille P, Lane B, Bassel GW, Prusinkiewicz P, Smith RS, Weijers D (2014) Genetic control of plant development by overriding a geometric division rule. Dev Cell 29(1):75–87. doi:10.1016/j.devcel.2014.02.002
Zabala G, Campos E, Varala KK, Bloomfield S, Jones SI, Win H, Tuteja JH, Calla B, Clough SJ, Hudson M, Vodkin LO (2012) Divergent patterns of endogenous small RNA populations from seed and vegetative tissues of Glycine max. BMC Plant Biol 12:177. doi:10.1186/1471-2229-12-177
Zhang J, Zhang S, Han S, Wu T, Li X, Li W, Qi L (2012a) Genome-wide identification of microRNAs in larch and stage-specific modulation of 11 conserved microRNAs and their targets during somatic embryogenesis. Planta 236(2):647–657. doi:10.1007/s00425-012-1643-9
Zhang J, Wu T, Li L, Han S, Li X, Zhang S, Qi L (2012b) Dynamic expression of small RNA populations in larch (Larix leptolepis). Planta 237(1):89–101. doi:10.1007/s00425-012-1753-4
Zhang YC, Yu Y, Wang CY, Li ZY, Liu Q, Xu J, Liao JY, Wang XJ, Qu LH, Chen F, Xin P, Yan C, Chu J, Li HQ, Chen YQ (2013a) Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nat Biotechnol 31(9):848–852. doi:10.1038/nbt.2646
Zhang J, Zhang S, Han S, Li X, Tong Z et al (2013b) Deciphering small noncoding RNAs during the transition from dormant embryo to germinated embryo in larches (Larix leptolepis). PLoS One 8(12):e81452. doi:10.1371/journal.pone.0081452
Zhang JH, Zhang SG, Li SG, Han SY, Li WF, Li XM, Qi LW (2014) Regulation of synchronism by abscisic-acid-responsive small noncoding RNAs during somatic embryogenesis in larch (Larix leptolepis). Plant Cell Tiss Organ Cult 116(3):361–370. doi:10.1007/s11240-013-0412-1
Zhang F, Dong W, Huang L, Song A, Wang H, Fang W et al (2015a) Identification of MicroRNAs and their targets associated with embryo abortion during chrysanthemum cross breeding via high-throughput sequencing. PLoS One 10(4):e0124371. doi:10.1371/journal.pone.0124371
Zhang HM, Kuang S, Xiong X, Gao T, Liu C, Guo AY (2015b) Transcription factor and microRNA co-regulatory loops: important regulatory motifs in biological processes and diseases. Brief Bioinform 16(1):45–58. doi:10.1093/bib/bbt085
Zhao YT, Wang M, Fu SX et al (2012) Small RNA profiling in two Brassica napus cultivars identifies microRNAs with oil production- and development-correlated expression and new small RNA classes. Plant Physiol 158:813–823. doi:10.1104/pp.111.187666
Zhu H, Hu F, Wang R, Zhou X, Sze SH, Liou LW, Barefoot A, Dickman M, Zhang X (2011) Arabidopsis Argonaute10 specifically sequesters miR166/165 to regulate shoot apical meristem development. Cell 145(2):242–256. doi:10.1016/j.cell.2011.03.024
Zong J, Yao X, Yin J, Zhang D, Ma H (2009) Evolution of the RNA-dependent RNA polymerase (RdRP) genes: duplications and possible losses before and after the divergence of major eukaryotic groups. Gene 447(1):29–39. doi:10.1016/j.gene.2009.07.004
Author contribution statement
Both authors contributed equally for the structuring and writing of this review.
Acknowledgements
This work was supported through projects funded by (1) the European Commission Seventh Framework Programme (FP7, Grant Agreement No 289841-PROCOGEN), and (2) Fundação para a Ciência e a Tecnologia (FCT), through Grants GREEN-it (UID/Multi/04551/2013), IF/01168/2013 and the doctoral fellowship SFRH/BD/79779/2011 (to ASR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Nese Sreenivasulu.
Rights and permissions
About this article
Cite this article
Rodrigues, A.S., Miguel, C.M. The pivotal role of small non-coding RNAs in the regulation of seed development. Plant Cell Rep 36, 653–667 (2017). https://doi.org/10.1007/s00299-017-2120-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-017-2120-5