Abstract
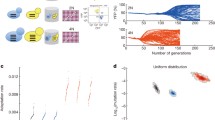
Ploidy is considered a very stable cellular characteristic. Although rare, changes in ploidy play important roles in the acquisition of long-term adaptations. Since these duplications allow the subsequent loss of individual chromosomes and accumulation of mutations, changes in ploidy can also cause genomic instability, and have been found to promote cancer. Despite the importance of the subject, measuring the rate of whole-genome duplications has proven extremely challenging. We have recently measured the rate of diploidization in yeast using long-term, in-lab experiments. We found that spontaneous diploidization occurs frequently, by two different mechanisms: endoreduplication and mating type switching. Despite its common occurrence, spontaneous diploidization is usually selected against, although it can be advantageous under some stressful conditions. Our results have implications for the understanding of evolutionary processes, as well as for the use of yeast cells in biotechnological applications.

Similar content being viewed by others
References
Adamczyk J, Deregowska A, Panek A, Golec E, Lewinska A, Wnuk M (2016) Affected chromosome homeostasis and genomic instability of clonal yeast cultures. Curr Genet 62:405–418. https://doi.org/10.1007/s00294-015-0537-3
Bell G (2010) Experimental genomics of fitness in yeast. Proc Biol Sci/R Soc 277:1459–1467. https://doi.org/10.1098/rspb.2009.2099
Berman J, Hadany L (2012) Does stress induce (para)sex? Implications for Candida albicans evolution. Trends Genet 28:197–203. https://doi.org/10.1016/j.tig.2012.01.004
Chidi BS, Rossouw D, Bauer FF (2016) Identifying and assessing the impact of wine acid-related genes in yeast. Curr Genet 62:149–164. https://doi.org/10.1007/s00294-015-0498-6
Cubillos FA (2016) Exploiting budding yeast natural variation for industrial processes. Curr Genet 62:745–751. https://doi.org/10.1007/s00294-016-0602-6
Cuypers TD, Hogeweg P (2014) A synergism between adaptive effects and evolvability drives whole genome duplication to fixation. PLoS Comput Biol 10:e1003547. https://doi.org/10.1371/journal.pcbi.1003547
Desai MM, Fisher DS, Murray AW (2007) The speed of evolution and maintenance of variation in asexual populations. Curr Biol 17:385–394. https://doi.org/10.1016/j.cub.2007.01.072
Dunham MJ, Badrane H, Ferea T, Adams J, Brown PO, Rosenzweig F, Botstein D (2002) Characteristic genome rearrangements in experimental evolution of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 99:16144–16149. https://doi.org/10.1073/pnas.242624799
Edgar BA, Orr-Weaver TL (2001) Endoreplication cell cycles: more for less. Cell 105:297–306 doi
Fujiwara T, Bandi M, Nitta M, Ivanova EV, Bronson RT, Pellman D (2005) Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 437:1043–1047. https://doi.org/10.1038/nature04217
Gallone B, Steensels J, Prahl T, Soriaga L, Saels V, Herrera-Malaver B, Merlevede A, Roncoroni M, Voordeckers K, Miraglia L, Teiling C, Steffy B, Taylor M, Schwartz A, Richardson T, White C, Baele G, Maere S, Verstrepen KJ (2016) Domestication and divergence of Saccharomyces cerevisiae beer yeasts. Cell 166:1397–1410 e1316. https://doi.org/10.1016/j.cell.2016.08.020
Gerstein AC (2013) Mutational effects depend on ploidy level: all else is not equal. Biol Lett 9:20120614. https://doi.org/10.1098/rsbl.2012.0614
Gerstein AC, Otto SP (2011) Cryptic fitness advantage: diploids invade haploid populations despite lacking any apparent advantage as measured by standard fitness assays. PLoS ONE 6:e26599. https://doi.org/10.1371/journal.pone.0026599
Gerstein AC, Chun HJ, Grant A, Otto SP (2006) Genomic convergence toward diploidy in Saccharomyces cerevisiae. PLoS Genet 2:e145. https://doi.org/10.1371/journal.pgen.0020145
Gerstein AC, Lim H, Berman J, Hickman MA (2017) Ploidy tug-of-war: evolutionary and genetic environments influence the rate of ploidy drive in a human fungal pathogen. Evol Int J Org Evol 71:1025–1038. https://doi.org/10.1111/evo.13205
Gore J, Youk H, van Oudenaarden A (2009) Snowdrift game dynamics and facultative cheating in yeast. Nature 459:253–256. https://doi.org/10.1038/nature07921
Harari Y, Ram Y, Rappoport N, Hadany L, Kupiec M (2018) Spontaneous changes in ploidy are common in yeast. Curr Biol. https://doi.org/10.1016/j.cub.2018.01.062
Hou J, Schacherer J (2016) Negative epistasis: a route to intraspecific reproductive isolation in yeast? Curr Genet 62:25–29. https://doi.org/10.1007/s00294-015-0505-y
Lang GI, Rice DP, Hickman MJ, Sodergren E, Weinstock GM, Botstein D, Desai MM (2013) Pervasive genetic hitchhiking and clonal interference in forty evolving yeast populations. Nature 500:571–574. https://doi.org/10.1038/nature12344
Lee CS, Haber JE (2015) Mating-type gene switching in Saccharomyces cerevisiae. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.MDNA3-0013-2014
McDonald MJ, Hsieh YY, Yu YH, Chang SL, Leu JY (2012) The evolution of low mutation rates in experimental mutator populations of Saccharomyces cerevisiae. Curr Biol 22:1235–1240. https://doi.org/10.1016/j.cub.2012.04.056
Meiron H, Nahon E, Raveh D (1995) Identification of the heterothallic mutation in HO-endonuclease of S. cerevisiae using HO/ho chimeric genes. Curr Genet 28:367–373
Ram Y, Hadany L (2016) Condition-dependent sex: who does it, when and why? Philos Trans R Soc Lond Ser B. https://doi.org/10.1098/rstb.2015.0539
Ratcliff WC, Denison RF, Borrello M, Travisano M (2012) Experimental evolution of multicellularity. Proc Natl Acad Sci USA 109:1595–1600. https://doi.org/10.1073/pnas.1115323109
Romano GH, Harari Y, Yehuda T, Podhorzer A, Rubinstein L, Shamir R, Gottlieb A, Silberberg Y, Pe’er D, Ruppin E, Sharan R, Kupiec M (2013) Environmental stresses disrupt telomere length homeostasis. PLoS Genet 9:e1003721. https://doi.org/10.1371/journal.pgen.1003721
Sellis D, Kvitek DJ, Dunn B, Sherlock G, Petrov DA (2016) Heterozygote advantage is a common outcome of adaptation in Saccharomyces cerevisiae. Genetics 203:1401–1413. https://doi.org/10.1534/genetics.115.185165
Selmecki AM, Maruvka YE, Richmond PA, Guillet M, Shoresh N, Sorenson AL, De S, Kishony R, Michor F, Dowell R, Pellman D (2015) Polyploidy can drive rapid adaptation in yeast. Nature 519:349–352. https://doi.org/10.1038/nature14187
Snoek T, Verstrepen KJ, Voordeckers K (2016) How do yeast cells become tolerant to high ethanol concentrations? Curr Genet 62:475–480. https://doi.org/10.1007/s00294-015-0561-3
Steensels J, Verstrepen KJ (2014) Taming wild yeast: potential of conventional and nonconventional yeasts in industrial fermentations. Annu Rev Microbiol 68:61–80. https://doi.org/10.1146/annurev-micro-091213-113025
Storchova Z, Kuffer C (2008) The consequences of tetraploidy and aneuploidy. J Cell Sci 121:3859–3866. https://doi.org/10.1242/jcs.039537
Thompson DA, Desai MM, Murray AW (2006) Ploidy controls the success of mutators and nature of mutations during budding yeast evolution. Curr Biol 16:1581–1590. https://doi.org/10.1016/j.cub.2006.06.070
Toprak E, Veres A, Michel JB, Chait R, Hartl DL, Kishony R (2012) Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat Genet 44:101–105. https://doi.org/10.1038/ng.1034
Tosato V, Sims J, West N, Colombin M, Bruschi CV (2017) Post-translocational adaptation drives evolution through genetic selection and transcriptional shift in Saccharomyces cerevisiae. Curr Genet 63:281–292. https://doi.org/10.1007/s00294-016-0635-x
Ungar L, Harari Y, Toren A, Kupiec M (2011) Tor complex 1 controls telomere length by affecting the level of Ku. Curr Biol 21:2115–2120. https://doi.org/10.1016/j.cub.2011.11.024
Venkataram S, Dunn B, Li Y, Agarwala A, Chang J, Ebel ER, Geiler-Samerotte K, Herissant L, Blundell JR, Levy SF, Fisher DS, Sherlock G, Petrov DA (2016) Development of a comprehensive genotype-to-fitness map of adaptation-driving mutations in yeast. Cell 166:1585–1596, e1522. https://doi.org/10.1016/j.cell.2016.08.002
Voordeckers K, Kominek J, Das A, Espinosa-Cantu A, De Maeyer D, Arslan A, Van Pee M, van der Zande E, Meert W, Yang Y, Zhu B, Marchal K, DeLuna A, Van Noort V, Jelier R, Verstrepen KJ (2015) Adaptation to high ethanol reveals complex evolutionary pathways. PLoS Genet 11:e1005635. https://doi.org/10.1371/journal.pgen.1005635
Yona AH, Manor YS, Herbst RH, Romano GH, Mitchell A, Kupiec M, Pilpel Y, Dahan O (2012) Chromosomal duplication is a transient evolutionary solution to stress. Proc Natl Acad Sci USA 109:21010–21015. https://doi.org/10.1073/pnas.1211150109
Zhang N, Cao L (2017) Starvation signals in yeast are integrated to coordinate metabolic reprogramming and stress response to ensure longevity. Curr Genet 63:839–843. https://doi.org/10.1007/s00294-017-0697-4
Zhu YO, Siegal ML, Hall DW, Petrov DA (2014) Precise estimates of mutation rate and spectrum in yeast. Proc Natl Acad Sci USA 111:E2310-2318. https://doi.org/10.1073/pnas.1323011111
Zhu YO, Sherlock G, Petrov DA (2016) Whole genome analysis of 132 clinical Saccharomyces cerevisiae strains reveals extensive ploidy variation. G3 6:2421–2434. https://doi.org/10.1534/g3.116.029397
Acknowledgements
We thank present and past members of the Kupiec lab for support, ideas and technical help. This work was supported by Grants from the Minerva Stiftung, the Volkswagen Foundation and the Israel Science Foundation to MK and the Stanford Center for Computational, Evolutionary and Human Genomics to YR.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Harari, Y., Ram, Y. & Kupiec, M. Frequent ploidy changes in growing yeast cultures. Curr Genet 64, 1001–1004 (2018). https://doi.org/10.1007/s00294-018-0823-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0823-y