Abstract
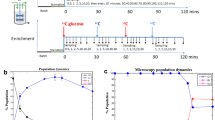
Quiescent cells exploit an array of transcription factors to activate stress response machinery and maintain survival under nutrient-limited conditions. Our recent findings reveal that these transcription factors also play an important role in the exit of quiescence and regrowth. By studying Saccharomyces cerevisiae under a continuous, nutrient-limited condition, we found that Msn2 and Msn4 function as master regulators of glycolytic genes in the quiescent-like phase. They control the timing of transition from quiescence to growth by regulating the accumulation rate of acetyl-CoA, a key metabolite that is downstream of glycolysis and drives growth. These findings suggest a model that Msn2/4 not only protect the cells from starvation but also facilitate their regrowth from quiescence. Thus, understanding the functions of stress response transcription factors in metabolic regulation will provide deeper insight into how quiescent cells manage the capacity of regrowth.


Similar content being viewed by others
References
Broach JR (2012) Nutritional control of growth and development in yeast. Genetics 192:73–105. https://doi.org/10.1534/genetics.111.135731
Cai L, Sutter BM, Li B, Tu BP (2011) Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol Cell 42:426–437. https://doi.org/10.1016/j.molcel.2011.05.004
De Virgilio C (2012) The essence of yeast quiescence. FEMS Microbiol Rev 36:306–339. https://doi.org/10.1111/j.1574-6976.2011.00287.x
Dechant R, Peter M (2008) Nutrient signals driving cell growth. Curr Opin Cell Biol 20:678–687. https://doi.org/10.1016/j.ceb.2008.09.009
Estruch F (2000) Stress-controlled transcription factors, stress-induced genes and stress tolerance in budding yeast. FEMS Microbiol Rev 24:469–486
Estruch F, Carlson M (1993) Two homologous zinc finger genes identified by multicopy suppression in a SNF1 protein kinase mutant of Saccharomyces cerevisiae. Mol Cell Biol 13:3872–3881
Gray JV, Petsko GA, Johnston GC, Ringe D, Singer RA, Werner-Washburne M (2004) “Sleeping beauty”: quiescence in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 68:187–206. https://doi.org/10.1128/mmbr.68.2.187-206.2004
Klosinska MM, Crutchfield CA, Bradley PH, Rabinowitz JD, Broach JR (2011) Yeast cells can access distinct quiescent states. Genes Dev 25:336–349. https://doi.org/10.1101/gad.2011311
Kuang Z, Cai L, Zhang X, Ji H, Tu BP, Boeke JD (2014) High-temporal-resolution view of transcription and chromatin states across distinct metabolic states in budding yeast. Nat Struct Mol Biol 21:854–863. https://doi.org/10.1038/nsmb.2881
Kuang Z, Pinglay S, Ji H, Boeke JD (2017) Msn2/4 regulate expression of glycolytic enzymes and control transition from quiescence to growth. eLife. https://doi.org/10.7554/eLife.29938
Kuang Z, Ji Z, Boeke JD, Ji H (2018) Dynamic motif occupancy (DynaMO) analysis identifies transcription factors and their binding sites driving dynamic biological processes. Nucleic Acids Res 46:e2. https://doi.org/10.1093/nar/gkx905
Laporte D, Lebaudy A, Sahin A, Pinson B, Ceschin J, Daignan-Fornier B, Sagot I (2011) Metabolic status rather than cell cycle signals control quiescence entry and exit. J Cell Biol 192:949–957. https://doi.org/10.1083/jcb.201009028
Martinez-Pastor MT, Marchler G, Schuller C, Marchler-Bauer A, Ruis H, Estruch F (1996) The Saccharomyces cerevisiae zinc finger proteins Msn2p and Msn4p are required for transcriptional induction through the stress response element (STRE). EMBO J 15:2227–2235
Miles S, Breeden L (2017) A common strategy for initiating the transition from proliferation to quiescence. Curr Genet 63:179–186. https://doi.org/10.1007/s00294-016-0640-0
Schmitt AP, McEntee K (1996) Msn2p, a zinc finger DNA-binding protein, is the transcriptional activator of the multistress response in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 93:5777–5782
Shi L, Tu BP (2013) Acetyl-CoA induces transcription of the key G1 cyclin CLN3 to promote entry into the cell division cycle in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 110:7318–7323. https://doi.org/10.1073/pnas.1302490110
Soontorngun N (2017) Reprogramming of nonfermentative metabolism by stress-responsive transcription factors in the yeast Saccharomyces cerevisiae. Curr Genet 63:1–7. https://doi.org/10.1007/s00294-016-0609-z
Tu BP, Kudlicki A, Rowicka M, McKnight SL (2005) Logic of the yeast metabolic cycle: temporal compartmentalization of cellular processes. Science 310:1152–1158. https://doi.org/10.1126/science.1120499
Tu BP, Mohler RE, Liu JC, Dombek KM, Young ET, Synovec RE, McKnight SL (2007) Cyclic changes in metabolic state during the life of a yeast cell. Proc Natl Acad Sci USA 104:16886–16891. https://doi.org/10.1073/pnas.0708365104
Zhang N, Cao L (2017) Starvation signals in yeast are integrated to coordinate metabolic reprogramming and stress response to ensure longevity. Curr Genet 63:839–843. https://doi.org/10.1007/s00294-017-0697-4
Acknowledgements
Supported by NIH Grant 5P50GM107632-06 to H.J. and J.D.B and R01HG006282 to H.J.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Kuang, Z., Ji, H. & Boeke, J.D. Stress response factors drive regrowth of quiescent cells. Curr Genet 64, 807–810 (2018). https://doi.org/10.1007/s00294-018-0813-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0813-0