Abstract
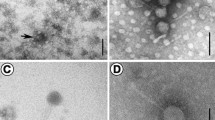
We used the double-agar layer method to isolate a novel Marinobacter marina bacteriophage, B23, from the surface water sample of the Bohai sea of China. There is some work to better understand the phage. The result of transmission electron microscopy revealed that B23 belongs to the family Siphoviridae with a head of 80 nm in diameter and a tail of 230 nm. Microbiological characterization evidenced that phage B23 is stable at the temperatures from − 25 to 60 °C, and showed vigorous vitality at pH between 4.0 and 12.0. One-step growth experiment showed that it had a longer latent period and higher lysis efficiency. Furthermore, the complete genome of B23 was sequenced and analyzed, which consists of a 35132 bp DNA with a G + C content of 59.8% and 50 putative open reading frames. The genome was divided into five parts, consisting of DNA replication and regulation, phage packaging, phage structure, host lysis and hypothetical protein.






Similar content being viewed by others
References
Motlagh AM et al (2017) Insights of phage-host interaction in hypersaline ecosystem through metagenomics analyses. Front Microbiol 8(8):352. https://doi.org/10.3389/fmicb.2017.00352
Labonté JM et al (2015) Single-cell genomics-based analysis of virus-host interactions in marine surface bacterioplankton. ISME J 9(11):2386–2399. https://doi.org/10.1038/ismej.2015.48
Jian H et al (2016) Filamentous phage sw1 is active and influences the transcriptome of the host at high-pressure and low-temperature. Environ Microbiol Rep 8(3):358. https://doi.org/10.1111/1758-2229.12388
Duhaime MB, Wichels A, Waldmann J (2011) Ecogenomics and genome landscapes of marine Pseudoalteromonas phage h105/1. ISME J 5(1):107–121. https://doi.org/10.1038/ismej.2010.94
Paez-Espino D et al (2016) Uncovering earth’s virome. Nature 536(7617):425. https://doi.org/10.1038/nature19094
Gauthier MJ et al (1992) Marinobacter hydrocarbonoclasticus gen. nov. sp. nov. a new, extremely halotolerant, hydrocarbon-degrading marine bacterium. Int J Syst Bacteriol 42(4):568. https://doi.org/10.1099/00207713-42-4-568
Ivanova EP et al (2014) Draft genome sequences of Marinobacter similis A3d10T and Marinobacter salarius R9SW1T. Genome Announc 2(3):e00442-14. https://doi.org/10.1128/genomeA.00442-14
Yuan J et al (2015) The diversity of pah-degrading bacteria in a deep-sea water column above the southwest Indian ridge. Front Microbiol 6:853. https://doi.org/10.3389/fmicb.2015.00853
Luo YJ et al (2015) Marinobacter Shengliensis sp. nov. a moderately halophilic bacterium isolated from oil-contaminated saline soil. Antonie Van Leeuwenhoek 107(4):1085–1094. https://doi.org/10.1007/s10482-015-0401-y
Bonin P et al (2015) Substrates specialization in lipid compounds and hydrocarbons of Marinobacter genus. Environ Sci Pollut Res 22(20):15347–15359. https://doi.org/10.1007/s11356-014-4009-y
Dombrowski N et al (2016) Reconstructing metabolic pathways of hydrocarbon-degrading bacteria from the deepwater horizon oil spill. Nat Microbiol 1(7):16057. https://doi.org/10.1186/1471-2164-15-483
Raddadi N et al (2017) Marinobacter sp. from marine sediments produce highly stable surface-active agents for combatting marine oil spills. Microb Cell Fact 16(1):186. https://doi.org/10.1186/s12934-017-0797-3
Rosenberg E et al (2010) The phage-driven microbial loop in petroleum bioremediation. Microb Biotechnol 3(4):467–472. https://doi.org/10.1111/j.1751-7915.2010.00182.x
Azam F (1998) Microbial control of oceanic carbon flux: the plot thickens. Science 280:694–696
Middelboe M, Chan AM, Bertelsen SK (2010) Isolation and life-cycle characterization of lytic viruses infecting heterotrophic bacteria and cyanobacteria. Man Aquat Viral Ecol 2014:149–180. https://doi.org/10.4319/mave.2010.978-0-9845591-0-7.118
Shahrbabak SS et al (2013) Isolation, characterization and complete genome sequence of Phaxi: a phage of Escherichia coli o157: h7. Microbiology 159(8):1629–1638. https://doi.org/10.1099/mic.0.063776-0
Chakravorty S et al (2007) A detailed analysis of 16 s ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J Microbiol Methods 69(2):330. https://doi.org/10.1016/j.mimet.2007.02.005
Xiu P, Liu R, Zhang D (2017) Pumilacidin-like lipopeptides derived from marine bacterium Bacillus sp. suppress the motility of Vibrio alginolyticus. Appl Environ Microbiol 31:83(12). https://doi.org/10.1128/AEM.00450-17
Jamalludeen N et al (2007) Isolation and characterization of nine bacteriophages that lyse o149 enterotoxigenic Escherichia coli. Vet Microbiol 124(1–2):47–57. https://doi.org/10.1016/j.vetmic.2007.03.028
Wang DB et al (2015) Characterization and genome sequencing of a novel bacteriophage ph101 infecting Pseudoalteromonas marina, bh101 from the Yellow sea of China. Curr Microbiol 71(5):594–600. https://doi.org/10.1007/s00284-015-0896-5
Deveau H et al (2006) Biodiversity and classification of lactococcal phages. Appl Environ Microbiol 72(6):4338. https://doi.org/10.1128/AEM.02517-05
Pajunen M, Kiljunen S, Skurnik M (2000) Bacteriophage YeO3-12, Specific for Yersinia enterocolitica Serotype O:3, Is Related to Coliphages T3 and T7 This study. J Bacteriol 182(18):5114–5120
Capra ML, Quiberoni A, Reinheimer JA (2004) Thermal and chemical resistance of Lactobacillus casei, and Lactobacillus paracasei bacteriophages. Lett Appl Microbiol 38(6):499–504. https://doi.org/10.1111/j.1472-765X.2004.01525.x
Karumidze N et al (2013) Isolation and characterisation of lytic bacteriophages of Klebsiella pneumoniae, and Klebsiella oxytoca. Curr Microbiol 66(3):251. https://doi.org/10.1007/s00284-012-0264-7
Xu Y et al (2016) Characterization, genome sequence, and analysis of Escherichia phage cicc 80001, a bacteriophage infecting an efficient l-aspartic acid producing Escherichia coli. Food Environ Virol 8(1):1–9. https://doi.org/10.1007/s12560-015-9218-0
Liu Z et al (2016) Isolation and genome sequencing of a novel Pseudoalteromonas phage ph1. Curr Microbiol 74(2):1–7. https://doi.org/10.1007/s00284-016-1175-9
Kassa T, Chhibber S (2012) Thermal treatment of the bacteriophage lysate of Klebsiella pneumoniae b5055 as a step for the purification of capsular depolymerase enzyme. J Virol Methods 179(1):135–141. https://doi.org/10.1016/j.jviromet.2011.10.011
Jun JW et al (2013) Characterization and complete genome sequence of the Shigella bacteriophage psf-1. Res Microbiol 164(10):979–986. https://doi.org/10.1016/j.resmic.2013.08.007
Brussow H, Desiere F (2001) Comparative phage genomics and the evolution of Siphoviridae: insights from dairy phages. Mol Microbiol 39(2):213–222
Eppler K et al (1991) Nucleotide sequence of the bacteriophage p22 genes required for DNA packaging. Virology 183(2):519–538
Kropinski AM et al (2013) The host-range, genomics and proteomics of Escherichia coli, o157:h7 bacteriophage rv5. Virol J 10(1):1–12. https://doi.org/10.1186/1743-422X-10-76
Gong Z et al (2017) Isolation and complete genome sequence of a novel Pseudoalteromonas, phage ph357 from the Yangtze river estuary. Curr Microbiol 74(7):1–8. https://doi.org/10.1007/s00284-017-1244-8
Acknowledgements
We are grateful to the research vessel Dong Fang Hong 2, for providing the seawater samples. The research was funded by the National Natural Science Foundation of China (Nos. 31500339 and 41076088), the National Key Basic Research Program of China (973Program, Grant No: 2013CB429704), China Postdoctoral Science Foundation (Grant Nos. 2015M570612 and 2016T90649), and Fundamental Research Funds for the Central University of Ocean University of China (Grant Nos. 201762017, 201564010 and 201512008).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhu, M., Wang, M., Jiang, Y. et al. Isolation and Complete Genome Sequence of a Novel Marinobacter Phage B23. Curr Microbiol 75, 1619–1625 (2018). https://doi.org/10.1007/s00284-018-1568-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-018-1568-z