Abstract
Purpose
The aim of this systematic review was to analyse literature on artificial intelligence (AI) and radiomics, including all medical imaging modalities, for oncological and non-oncological applications, in order to assess how far the image mining research stands from routine medical application. To do this, we applied a trial phases classification inspired from the drug development process.
Methods
Among the articles we considered for inclusion from PubMed were multimodality AI and radiomics investigations, with a validation analysis aimed at relevant clinical objectives. Quality assessment of selected papers was performed according to the QUADAS-2 criteria. We developed the phases classification criteria for image mining studies.
Results
Overall 34,626 articles were retrieved, 300 were selected applying the inclusion/exclusion criteria, and 171 high-quality papers (QUADAS-2 ≥ 7) were identified and analysed. In 27/171 (16%), 141/171 (82%), and 3/171 (2%) studies the development of an AI-based algorithm, radiomics model, and a combined radiomics/AI approach, respectively, was described. A total of 26/27(96%) and 1/27 (4%) AI studies were classified as phase II and III, respectively. Consequently, 13/141 (9%), 10/141 (7%), 111/141 (79%), and 7/141 (5%) radiomics studies were classified as phase 0, I, II, and III, respectively. All three radiomics/AI studies were categorised as phase II trials.
Conclusions
The results of the studies are promising but still not mature enough for image mining tools to be implemented in the clinical setting and be widely used. The transfer learning from the well-known drug development process, with some specific adaptations to the image mining discipline could represent the most effective way for radiomics and AI algorithms to become the standard of care tools.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
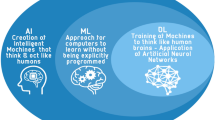
The “Artificial Intelligence (AI) winter” [1] is over. AI and radiomics approaches applied to medical images for the non-invasive characterisation of diseases (i.e., image mining) have remarkably increased in recent years. The first reports on AI and radiomics applied to medical images date back to 1963 [2] and 1973 [3], respectively, but the enthusiasm of those years broke off quite soon. Recently, increasing amounts of electronic medical data, technological improvements, and health sustainability issues resulted in a renewed interest in both AI and radiomics applications.
Image mining is claimed to have a potentially huge clinical relevance with the possibility to non-invasively diagnose, characterise and predict the outcome in almost all medical conditions. However, despite the amount of published studies, some issues including significance, goodness, and strength of the reported results are still to be addressed. Particularly, it is not clear how far image mining is from clinical practice.
Therefore, the aim of this systematic review was to analyse literature on AI and radiomics, including all medical imaging modalities, for oncological and non-oncological applications, in order to assess how far the image mining research stands from routine medical application. To do this, we applied a trial phases classification inspired from the drug development process.
Material and methods
This systematic review was performed according to the PRISMA statement [4]. The PRISMA checklist is provided in Supplemental material.
Search, eligibility criteria and study selection
The endpoint of the analysis was to assess the potential of AI applied to medical images and radiomics (i.e., image mining) to be implemented in clinics. Our search algorithm within the PubMed/MEDLINE database consisted of the combination of the following terms: “Artificial intelligence[Mesh]” OR “Radiomic” OR “Radiomics” AND/OR “Radiography[Mesh]”, OR “Ultrasonography[Mesh]” OR “Tomography[Mesh]”, OR “Image interpretation, computed-assisted[Mesh]”, OR “Multimodal imaging[Mesh]”, OR “Tomography, emission-computed[Mesh]”, OR “Photography[Mesh]”. No start date limit was used, and the search was extended until September 5th, 2018.
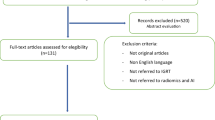
According to the scope of the review, we considered AI (i.e., algorithms that take images as inputs) and radiomics investigations aimed at relevant objectives in clinical practice: biological characterisation, risk stratification, treatment response prediction, toxicity prediction, and prognostication of a certain disease. The imaging modalities we considered were ultrasound, radiography, mammography, endoscopy, skin pictures, ocular fundus pictures, computed tomography (CT), magnetic resonance imaging (MRI), scintigraphy (either planar images, SPECT, or SPECT/CT) and positron emission tomography (PET) or PET/CT. Subsequently, we applied the following exclusion criteria: (a) articles not in the English language; (b) studies not within the field of interest; (c) guidelines, review articles and meta-analysis, editorials or letters, comments, and conference proceedings; (d) “in vitro”, phantom or animal studies; (e) case reports or small case series (≤ 10 patients); (f) studies involving healthy subjects; (g) research articles focused on methodological aspects (algorithm and/or software development and/or comparison; evaluation/comparison of method(s) for parameters optimization, segmentation and features extraction; test–retest studies); (h) testing data (not medical images) as input for AI algorithm(s); (i) radiomics studies evaluating descriptors of shape and size or image intensity histogram only (i.e., not textural features); (j) lack of validation in a clinical setting; (k) lack of conventional metrics (i.e., sensitivity, specificity, accuracy, and/or hazard ratio, and/or recall, and/or AUC, and/or C-index) for the report of validation results.
Two reviewers (MK and MS) independently performed an initial screening of the identified titles and abstracts applying the inclusion/exclusion criteria. The discrepancies were resolved by a third reviewer (LA). The decision rule for consensus was simple majority. Then, the reviewers retrieved the full-text reports of the selected abstracts and, subsequently, performed an independent second-step selection.
Quality assessment of the literature
Quality assessment of selected papers was performed according to the QUADAS-2 criteria, assessing 4 domains: (1) patient selection, (2) index test, (3) reference standard, and (4) flow and timing [5]. The signalling questions for each QUADAS-2 domain were tailored for the aim of this review as detailed in Table 1. This evaluation assigned the risk of bias to a study and ranked it as low (score = 2), high (score = 1), or indeterminate (score = 0) for each domain. We calculated the overall QUADAS-2 score as the sum of the scores. The appropriateness of statistical analysis was defined considering two aspects. First, the total number of patients analysed was considered appropriate if at least five patients/feature (after feature selection, if performed) were included in a radiomics study; while AI studies with more than 50 patients were considered as acceptable quality. The sample size criterion for radiomics studies was used adapting the conventional rule for multiple regression: the number of data points (i.e., observations or cases) should be considerably more than 5–10 times the number of variables [6]. At least ten patients per feature have been recommended in radiomics studies [7, 8]. The sample size criterion for AI studies was established assuming that at least 50 patients are needed to train and validate an algorithm, minimising the effects of overfitting and improving the quality of performance metrics, similarly to what is recommended for biomarker discovery [9]. Second, we assessed the balance in the number of patients between the subgroups (e.g. number of patients with benign vs malignant lesions in a study aimed at differential diagnosis); an imbalance of more than 2/3 was considered inappropriate.
Phases classification criteria
We developed the phases classification criteria for image mining studies, inspired by the classification applied to the clinical trials (Fig. 1). The parameters for phase categorisation included: sample size, type of study (retrospective/prospective), type of validation approach (internal/independent), and the development stage (pre-/post-marketing). Figure 1 reports the classification criteria in detail. We assigned each selected article to a phase: from 0 to IV.
Statistical analysis
A database was created for the qualitative synthesis of the papers and the studies’ results. We recorded the results obtained in the validation set only. In case of multiple aims within the same article, the primary one was considered. If image mining was applied to different imaging modalities within the same article (e.g., PET and CT), results were recorded for each one. If different approaches were tested within the same paper (e.g., models derived from random forest and support vector machine), the best in terms of diagnostic performance was analysed. The articles reporting identical or very similar sample size, research hypothesis, methodology and results sharing with almost the same authorship—“salami” publishing [10]—were excluded to avoid overlap in the study population and results redundancy. Accordingly, in case of a series of articles considered as “salami”, the one with the larger sample size was evaluated and the other(s) excluded. Papers with a QUADAS-2 ≥ 7 were included in the quantitative analysis. Descriptive statistical measures were used to summarise the data. Excel ® 2017 (Microsoft®, Redmond, WA) was used for analysis.
Results
Search, eligibility criteria and study selection
Overall 34,626 articles were retrieved using the search algorithm. Subsequently, 33,997 papers were excluded reviewing titles and abstracts and applying the inclusion/exclusion criteria. Five papers, suspected to be a “salami” publishing, were excluded. Figure 2 summarises the research process. Table S1 reports a qualitative summary of the selected 300, including the 171 high quality articles. In recent years, a striking increase in the number of papers published on image mining occurred. In fact, especially in 2017 and 2018, 66 (22%) and 131 (44%) articles, respectively, were published. Figure 3 shows the literature trend. The vast majority (more than 80%) come from the oncology field. However, more than 50% of the studies included in the qualitative analysis have been assigned a high risk of bias (Fig. 4). Accordingly, a substantial proportion 129/300 (43%) studies have been scored as having a considerable risk of bias, mainly in the “index test” and “patient selection” domains (QUADAS-2 ≤ 6) (Fig. 5). Table S2 reports a qualitative summary of the 171 high-quality (QUADAS ≥7) papers. The temporal trend of the literature according to the phase of the study is shown in Fig. 6. Figures 7, 8, and 9 represent the graphical syntheses of the high-quality articles (QUADAS-2 ≥ 7) considered from three different points of view: the clinician, the imager, and the researcher. Quantitative synthesis is summarised in Table 2. The main results of the phase III studies are reported in Table 3.
Discussion
The present systematic review is the first assessing the potential for implementation of image mining tools in clinical practice, by means of classification of the literature in development phases. Despite the amount of literature on image mining with a validation analysis, more than 90% of studies were classified as phase 0, I or II (i.e., retrospective). Collectively, their results were uncertain in terms of significance, goodness, and strength and their generalisability weak. Even among the studies with a QUADAS-2 ≥ 7, only 4.6% were categorised as phase III studies. As it emerges from the present systematic review, the results are promising but still not mature enough for clinical implementation and widespread use of image mining tools. Nonetheless, the study quality has increased in recent years.
Because of the paucity of phase III and IV studies, we did not proceed to a meta-analysis. Therefore, no definitive conclusion can be drawn on which approach among radiomics and AI should be preferred. AI techniques, in particular convolutional neural networks, have the advantage over radiomics of not requiring tumour segmentation, feature calculation and selection. These steps are even more critical in tiny lesions that have to be submitted to radiomics processing. On the other hand, vast cohorts are crucial for a robust AI-based model development that require big efforts to be collected and analysed. Also, an unbiased reference standard, not always easy to obtain, should be chosen to ensure AI model reliability. The combined radiomics/AI strategy is at its early stages [179,180,181,182,183] and the complementary role of radiomics and AI techniques should be addressed [184]. Which is the best image mining approach is still an open question.
Similar to the other “omics” domains (e.g., proteomics), few of the image mining biomarkers reached clinical practice [9]. The translation of image mining research in the clinical arena is limited by the huge variability of the methods used for image analysis, together with the impasse to reproduce the results when tested in a different cohort of patients. Validation is a critical issue. Theoretically, the validation analysis of a successful model should provide consistent performance measures to those obtained in the training process. Thereafter, results obtained in the validation cohort should be confirmed by the test-independent validation. Finally, the proposed approach should be effective for the indication within the “real world” population of patients the model has been developed for. The validation process may be internal (e.g., cross-validation and bootstrapping) or external (using data not used for training). Typically, the internal one, used for a preliminary evaluation or for the fine-tuning of the model under development, overestimate the performance [185]. In fact, the same cohort is used twice, once to choose the filtered subset and again to build a classification model resulting in the overfitting of the algorithm to the data [9]. The external validation may be performed using three different strategies: (i) temporal (i.e., data obtained in newly recruited patients), (ii) geographic (i.e., data collected in a different institution), and (iii) split-sample (i.e., data split from the entire dataset and kept untouched for the test). External validation is crucial to verify the generalisability of the results [185]; and the random patient selection is an essential prerequisite, as well as the balance in patient characteristics. Temporal or geographic validations should be preferred to the split-sample one. Particularly, the geographic validation, which accounts for technical variability aspects (scanners, acquisition parameters and protocols) [185], is expected to be more representative of the clinical setting.
We excluded a priori from the present analysis studies testing shape and size as well as histogram-based features since our aim was to assess the “maturity” of advanced image analysis at its’ full potential, entailing textural indexes derived from up to second-order features. Image analysis based on the gray level histogram only does not provide any information about the relative position of pixels/voxels to each other within the region of interest. Therefore, these features are not able to describe whether any low/high gray levels are positioned together, or if they are distributed between high/low-value gray levels [186].
In order to develop a valid and trustworthy image mining tool the cohorts in study (training, validation and test) should be representative of the target population. This means that the sample size should be big enough to minimise the effects of overfitting, be comprehensive of the “outliers”, and, consequently, be reliable when used for the assessment of unseen patients. We proposed that at least 50 patients should be included in AI studies, as also suggested by simulated analyses [9, 187, 188]. However, especially for deep learning approaches and complex tasks, much larger populations are needed. The effect of the sample size on the model performance has been already demonstrated. When a limited dataset (1000 samples) vs the complete dataset (>100,000) was used for retinopathy classifier development, the weighted error resulted as 12.7% vs 6.6%, respectively [33]. Moreover, the sample size was not the only criterion used to score papers. Accordingly, even if the number of patients was relatively low, only well-designed studies have been scored as having low risk of bias and included in the quantitative analysis.
In the present work we arbitrarily chose 100 samples as the threshold for trial phases categorisation (II vs III ad above). This was a conservative decision. The conventional rule of the necessity to include at least ten patients per tested variable if applied for phase classification would have led to downgrading of most of the studies. Considering the fact that generally a feature reduction strategy was put in place we chose 100 as a reasonable cut-off. When planning an image mining study, both statistical recommendations (sample size and prospective design) and clinical conditions’ variability should be considered in order to develop an algorithm on a dataset that realistically represents the target patient population. The sample size calculation has been estimated in only 3 out of the 171 papers [44, 88, 125]. The use of multiple images taken from the same patient should be limited since they are prone to be similar (or almost identical) with a negative impact on the generalisability potential. In fact, even if this process increases the sample size, the multiplied number of observations (or cases) not representative of the inter-patient variability, overestimates the model performance. Accordingly, data augmentation should be properly used to avoid overfitting, keeping in mind that it cannot completely overcome the requirement of a proper sample size.
To assess the appropriateness of the statistical analysis, we considered not only the sample size but also the number of patients within the subgroups (an imbalance of more than 2/3 was considered inappropriate). Imbalanced cohorts in image mining studies may lead to constitutively biased results, which confer higher uncertainty and poor generalisability [189]. Consequently, in the developmental phase, proper study design and analysis strategy using stratification, matching, weighting, covariate adjustment, or regression should be adopted. Therefore, imbalance per se does not prevent the use of radiomic and AI-based approaches when the prevalence of a disease or an outcome is very low [190].
The applicability of the image mining framework to rare diseases is still an issue because of the limited data availability for model development. Transversal platforms for sharing and analysis of images and data, as envisaged by some research groups [191], could represent a valuable strategy for the investigations in this field.
Presently, it is unrealistic to justify a medical decision by the output provided by a neural network or a radiomics feature/signature. Little or nothing is still known on the biological significance of the image-derived parameters. Correlations with tumour grading [39, 45, 46, 94, 100, 117, 118, 138, 174], inflammatory infiltrate [131], gene expression, mutation and molecular pathways [25, 44, 47,48,49,50,51,52,53,54,55,56,57,58, 85,86,87, 101, 121, 122, 139,140,141,142, 180] have been reported. Nonetheless, more should be learned about the functioning of AI and radiomics approaches in order to solve the “black box” problem and to understand the underlying clinical and/or molecular connotation. Imagers should be able to assess the reliability of image mining approaches and to manage independently the patient (i.e., the pilot, plane and passengers during a flight). This innovative attitude, which implies the acquisition of technical and informatics skills, will contribute to remove the “black box” uncertainties, and to promote image mining towards clinical practice.
Additionally, some technical barriers should be faced when considering implementing image mining tools into the every-day practice. These include a time-consuming workflow; uncertain reproducibility of results among different scanners, acquisition protocols, and image-processing approaches; regulatory issues concerning privacy and ethics; and data protection. Common efforts should be realised to accelerate the research path on these aspects, and to implement the technological infrastructure and make the tools easy to use. Privacy and ethics regulations may restrict data and image sharing for the purpose of research and every-day clinical practice. A shared strategy needs to be built up for the management of these aspects. These challenges are an opportunity to develop a reliable methodology able to provide controlled data collection and secure infrastructure, instead of gathering uncertain-quality datasets.
The need to provide reliable results has generated multiple initiatives and recommendations to achieve methodology standardisation and reproducibility [7, 185, 191,192,193]. The increasing awareness among researchers of the urgency to increase the quality of the investigations determined an increase in the number of phase III trials in the last 2 years. In the era of evidence-based medicine, rigorous research with strict rules is the only way forward to achieve clinical acceptance and become part of the “standard of care”. The research process should aim to address a clinical need through an adequate statistical strategy, prospective and multi centre studies, robust reference standards, and adequate timing. Independent validation is mandatory together with the clarification of the impact of the technical aspects on image mining models. These items imply that reproducible, strong, and, hopefully, excellent results will be achieved through an adequate research process. In this respect, a closer collaboration should be established among clinical researchers, algorithm developers and data scientists.
We foresee the transfer learning from the well-known drug development process, with some specific adaptations, to the image mining discipline as the most effective way for radiomics and AI algorithms to get into routine clinical practice and avoid a new glacial era of image mining in the next decades.
References
Bohannon J, Bohannon J. Fears of an AI pioneer. Science (80- ). 2015;349:252.
Lodwick GS, Keats TE, Dorst JP. The coding of roentgen images for computer analysis as applied to lung cancer. Radiology. 1963;81:185–200.
Haralick R, Shanmugam K, Dinstein I. Texture features for image classification. IEEE Trans Sys Man Cybern. 1973;SMC 3:610–21.
Zorzela L, Loke Y, Ioannidis J, Golder S, Santaguida P, Altman D, et al. PRISMA harms checklist: improving harms reporting in systematic reviews. BMJ. 2016;352:i157.
Whiting PF, Rutjes AWS, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: strumento per valutare la qualità degli studi di accuratezza diagnostica. Evidence. 2016;8:e1000131.
Kleinbaum DG, Kupper LL, Muller KE, editors. Applied regression analysis and other multivariable methods. Boston: PWS Kent Publishing Co; 1988.
Sollini M, Cozzi L, Antunovic L, Chiti A, Kirienko M. PET radiomics in NSCLC: state of the art and a proposal for harmonization of methodology. Sci Rep. 2017;7:358.
Gillies RJ, Kinahan PE, Hricak H. Radiomics: images are more than pictures, they are data. Radiology. 2016;278:563–77.
Hernández B, Parnell A, Pennington SR. Why have so few proteomic biomarkers “survived” validation? (sample size and independent validation considerations). Proteomics. 2014;14:1587–92.
Supak Smolcić V. Salami publication: definitions and examples. Biochem medica. 2013;23:237–41.
Tong T, Wolz R, Gao Q, Guerrero R, Hajnal JV, Rueckert D. Multiple instance learning for classification of dementia in brain MRI. Med Image Anal. 2014;18:808–18.
Acton PD, Newberg A. Artificial neural network classifier for the diagnosis of Parkinson’s disease using [99mTc]TRODAT-1 and SPECT. Phys Med Biol. 2006;51:3057–66.
Guner LA, Karabacak NI, Akdemir OU, Karagoz PS, Kocaman SA, Cengel A, et al. An open-source framework of neural networks for diagnosis of coronary artery disease from myocardial perfusion SPECT. J Nucl Cardiol. 2010;17:405–13.
Lee JH, Kim DH, Jeong SN, Choi SH. Detection and diagnosis of dental caries using a deep learning-based convolutional neural network algorithm. J Dent. 2018;77:106–11.
Lee J-H, Kim D-H, Jeong S-N, Choi S-H. Diagnosis and prediction of periodontally compromised teeth using a deep learning-based convolutional neural network algorithm. J Periodontal Implant Sci. 2018;48:114–23.
Kong X, Gong S, Su L, Howard N, Kong Y. Automatic detection of acromegaly from facial photographs using machine learning methods. EBioMedicine. 2018;27:94–102.
Wang K, Lu X, Zhou H, Gao Y, Zheng J, Tong M, et al. Deep learning radiomics of shear wave elastography significantly improved diagnostic performance for assessing liver fibrosis in chronic hepatitis B: a prospective multicentre study. Gut. 2018:1–13.
Yasaka K, Akai H, Kunimatsu A, Abe O, Kiryu S. Deep learning for staging liver fibrosis on CT: a pilot study. Eur Radiol. 2018;28:4578–85.
Komeda Y, Handa H, Watanabe T, Nomura T, Kitahashi M, Sakurai T, et al. Computer-aided diagnosis based on convolutional neural network system for colorectal polyp classification: preliminary experience. Oncology. 2017;93:30–4.
Byrne MF, Chapados N, Soudan F, Oertel C, Linares Pérez M, Kelly R, et al. Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut. 2017;68:94–100.
Shichijo S, Nomura S, Aoyama K, Nishikawa Y, Miura M, Shinagawa T, et al. Application of convolutional neural networks in the diagnosis of helicobacter pylori infection based on endoscopic images. EBioMedicine. 2017;25:106–11.
Han SS, Park GH, Lim W, Kim MS, Na JI, Park I, et al. Deep neural networks show an equivalent and often superior performance to dermatologists in onychomycosis diagnosis: automatic construction of onychomycosis datasets by region-based convolutional deep neural network. PLoS One. 2018;13:1–14.
Li W, Cao P, Zhao D, Wang J. Pulmonary nodule classification with deep convolutional neural networks on computed tomography images. Comput Math Methods Med. 2016;2016:6215085.
Poedjiastoeti W, Suebnukarn S. Application of convolutional neural network in the diagnosis of jaw tumors. Healthc Inform Res. 2018;24:236–41.
Akkus Z, Ali I, Sedlář J, Agrawal JP, Parney IF, Giannini C, et al. Predicting deletion of chromosomal arms 1p/19q in low-grade gliomas from MR images using machine intelligence. J Digit Imaging. 2017;30:469–76.
Sato M, Horie K, Hara A, Miyamoto Y, Kurihara K, Tomio K, et al. Application of deep learning to the classification of images from colposcopy. Oncol Lett. 2018;15:3518–23.
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542:115–8.
Ypsilantis PP, Siddique M, Sohn HM, Davies A, Cook G, Goh V, et al. Predicting response to neoadjuvant chemotherapy with PET imaging using convolutional neural networks. PLoS One. 2015;10:1–18.
Gargeya R, Leng T. Automated identification of diabetic retinopathy using deep learning. Ophthalmology. 2017;124:962–9.
Niemeijer M, van Ginneken B, Russell SR, Suttorp-Schulten MSA, Abràmoff MD. Automated detection and differentiation of drusen, exudates, and cotton-wool spots in digital color fundus photographs for diabetic retinopathy diagnosis. Invest Ophthalmol Vis Sci. 2007;48:2260–7.
Takahashi H, Tampo H, Arai Y, Inoue Y, Kawashima H. Applying artificial intelligence to disease staging: deep learning for improved staging of diabetic retinopathy. PLoS One. 2017;12:1–11.
Raju M, Pagidimarri V, Barreto R, Kadam A, Kasivajjala V, Aswath A. Development of a deep learning algorithm for automatic diagnosis of diabetic retinopathy. Stud Health Technol Inform. 2017;245:559–63.
Kermany DDS, Goldbaum M, Cai W, Valentim CCS, Liang H, Baxter SL, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell. 2018;172:1122–1131.e1129.
Burlina PM, Joshi N, Pekala M, Pacheco KD, Freund DE, Bressler NM. Automated grading of age-related macular degeneration from color fundus images using deep convolutional neural networks. JAMA Ophthalmol. 2017;135:1170–6.
Olczak J, Fahlberg N, Maki A, Razavian AS, Jilert A, Stark A, et al. Artificial intelligence for analyzing orthopedic trauma radiographs: deep learning algorithms—are they on par with humans for diagnosing fractures? Acta Orthop. 2017;88:581–6.
Chung SW, Han SS, Lee JW, Oh KS, Kim NR, Yoon JP, et al. Automated detection and classification of the proximal humerus fracture by using deep learning algorithm. Acta Orthop. 2018;89:468–73.
Gonzalez G, Ash SY, Vegas-Sánchez-Ferrero G, Onieva JO, Rahaghi FN, Ross JC, et al. Disease staging and prognosis in smokers using deep learning in chest computed tomography. Am J Respir Crit Care Med. 2018;197:193–203.
Li W, Huang Y, Zhuang BW, Liu GJ, Hu HT, Li X, et al. Multiparametric ultrasomics of significant liver fibrosis: a machine learning-based analysis. Eur Radiol. 2019;29:1496–506.
Zhang X, Xu X, Tian Q, Li B, Wu Y, Yang Z, et al. Radiomics assessment of bladder cancer grade using texture features from diffusion-weighted imaging. J Magn Reson Imaging. 2017;46:1281–8.
Wu S, Zheng J, Li Y, Yu H, Shi S, Xie W, et al. A radiomics nomogram for the preoperative prediction of lymph node metastasis in bladder cancer. Clin Cancer Res. 2017;23:6904–11.
Wu S, Zheng J, Li Y, Wu Z, Shi S, Huang M, et al. Development and validation of an MRI-based radiomics signature for the preoperative prediction of lymph node metastasis in bladder cancer. EBioMedicine. 2018;34:76–84.
Zhang Z, Yang J, Ho A, Jiang W, Logan J, Wang X, et al. A predictive model for distinguishing radiation necrosis from tumor progression after gamma knife radiosurgery based on radiomic features from MR images HHS public access. Eur Radiol. 2018;28:2255–63.
Kim JY, Park JE, Jo Y, Shim WH, Nam SJ, Kim JH, et al. Incorporating diffusion- and perfusion-weighted MRI into a radiomics model improves diagnostic performance for pseudoprogression in glioblastoma patients. Neuro-Oncology. 2018:1–11.
Wei J, Yang G, Hao X, Gu D, Tan Y, Wang X, et al. A multi-sequence and habitat-based MRI radiomics signature for preoperative prediction of MGMT promoter methylation in astrocytomas with prognostic implication. Eur Radiol. 2019;29:877–88.
Coroller TP, Bi WL, Huynh E, Abedalthagafi M, Aizer AA, Greenwald NF, et al. Radiographic prediction of meningioma grade by semantic and radiomic features. PLoS One. 2017;12:1–15.
Chen W, Liu B, Peng S, Sun J, Qiao X. Computer-aided grading of gliomas combining automatic segmentation and radiomics. Int J Biomed Imaging. 2018;2018:2512037.
Rathore S, Akbari H, Rozycki M, Abdullah KG, Nasrallah MP, Binder ZA, et al. Radiomic MRI signature reveals three distinct subtypes of glioblastoma with different clinical and molecular characteristics, offering prognostic value beyond IDH1. Sci Rep. 2018;8:1–12.
Li Z-C, Bai H, Sun Q, Li Q, Liu L, Zou Y, et al. Multiregional radiomics features from multiparametric MRI for prediction of MGMT methylation status in glioblastoma multiforme: a multicentre study. Eur Radiol. 2018;28:3640–50.
Arita H, Kinoshita M, Kawaguchi A, Takahashi M, Narita Y, Terakawa Y, et al. Lesion location implemented magnetic resonance imaging radiomics for predicting IDH and TERT promoter mutations in grade II/III gliomas. Sci Rep. 2018;8:1–10.
Li Y, Liu X, Qian Z, Sun Z, Xu K, Wang K, et al. Genotype prediction of ATRX mutation in lower-grade gliomas using an MRI radiomics signature. Eur Radiol. 2018;28:2960–8.
Zhang X, Tian Q, Wang L, Liu Y, Li B, Liang Z, et al. Radiomics strategy for molecular subtype stratification of lower-grade glioma: detecting IDH and TP53 mutations based on multimodal MRI. J Magn Reson Imaging. 2018;48:916–26.
Akbari H, Bakas S, Pisapia JM, Nasrallah MP, Rozycki M, Martinez-Lage M, et al. In vivo evaluation of EGFRvIII mutation in primary glioblastoma patients via complex multiparametric MRI signature. Neuro-Oncology. 2018;20:1068–79.
Lu CF, Hsu FT, Hsieh KLC, Kao YCJ, Cheng SJ, Hsu JBK, et al. Machine learning–based radiomics for molecular subtyping of gliomas. Clin Cancer Res. 2018;24:4429–36.
Han Y, Xie Z, Zang Y, Zhang S, Gu D, Zhou M, et al. Non-invasive genotype prediction of chromosome 1p/19q co-deletion by development and validation of an MRI-based radiomics signature in lower-grade gliomas. J Neuro-Oncol. 2018;140:297–306.
Li Y, Liu X, Xu K, Qian Z, Wang K, Fan X, et al. MRI features can predict EGFR expression in lower grade gliomas: a voxel-based radiomic analysis. Eur Radiol. 2018;28:356–62.
García-Gómez JM, Luts J, Julià-Sapé M, Krooshof P, Tortajada S, Robledo JV, et al. Multiproject-multicenter evaluation of automatic brain tumor classification by magnetic resonance spectroscopy. MAGMA. 2009;22:5–18.
Li Y, Qian Z, Xu K, Wang K, Fan X, Li S, et al. Radiomic features predict Ki-67 expression level and survival in lower grade gliomas. J Neuro-Oncol. 2017;135:317–24.
Li Y, Qian Z, Xu K, Wang K, Fan X, Li S, et al. MRI features predict p53 status in lower-grade gliomas via a machine-learning approach. NeuroImage Clin. 2018;17:306–11.
Ortiz-Ramón R, Larroza A, Ruiz-España S, Arana E, Moratal D. Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur Radiol. 2018;28:4514–23.
Suh HB, Choi YS, Bae S, Ahn SS, Chang JH, Kang SG, et al. Primary central nervous system lymphoma and atypical glioblastoma: differentiation using radiomics approach. Eur Radiol. 2018;28:3832–9.
Chen Y, Li Z, Wu G, Yu J, Wang Y, Lv X, et al. Primary central nervous system lymphoma and glioblastoma differentiation based on conventional magnetic resonance imaging by high-throughput SIFT features. Int J Neurosci. 2018;128:608–18.
Kang D, Park JE, Kim YH, Kim JH, Oh JY, Kim J, et al. Diffusion radiomics as a diagnostic modal for atypical manifestation of primary central nervous system lymphoma: development and multicenter external validation. Neuro-Oncology. 2018;20:1251–61.
Yu J, Shi Z, Lian Y, Li Z, Liu T, Gao Y, et al. Noninvasive IDH1 mutation estimation based on a quantitative radiomics approach for grade II glioma. Eur Radiol. 2017;27:3509–22.
Liu Z, Wang Y, Liu X, Du Y, Tang Z, Wang K, et al. Radiomics analysis allows for precise prediction of epilepsy in patients with low-grade gliomas. NeuroImage Clin. 2018;19:271–8.
Kickingereder P, Götz M, Muschelli J, Wick A, Neuberger U, Shinohara RT, et al. Large-scale Radiomic profiling of recurrent glioblastoma identifies an imaging predictor for stratifying anti-Angiogenic treatment response. Clin Cancer Res. 2016;22:5765–71.
Kickingereder P, Burth S, Wick A, Götz M, Eidel O, Schlemmer H-P, et al. Radiomic profiling of glioblastoma: identifying an imaging predictor of patient survival with improved performance over established clinical and radiologic risk models. Radiology. 2016;280:880–9.
Ingrisch M, Schneider MJ, Nörenberg D, De Figueiredo GN, Maier-Hein K, Suchorska B, et al. Radiomic analysis reveals prognostic information in T1-weighted baseline magnetic resonance imaging in patients with glioblastoma. Investig Radiol. 2017;52:360–6.
Li Q, Bai H, Chen Y, Sun Q, Liu L, Zhou S, et al. A fully-automatic multiparametric radiomics model: towards reproducible and prognostic imaging signature for prediction of overall survival in glioblastoma multiforme. Sci Rep. 2017;7:1–9.
Beig N, Patel J, Prasanna P, Hill V, Gupta A, Correa R, et al. Radiogenomic analysis of hypoxia pathway is predictive of overall survival in glioblastoma. Sci Rep. 2018;8:7.
Kickingereder P, Neuberger U, Bonekamp D, Piechotta PL, Götz M, Wick A, et al. Radiomic subtyping improves disease stratification beyond key molecular, clinical, and standard imaging characteristics in patients with glioblastoma. Neuro-Oncology. 2018;20:848–57.
Grossmann P, Narayan V, Chang K, Rahman R, Abrey L, Reardon DA, et al. Quantitative imaging biomarkers for risk stratification of patients with recurrent glioblastoma treated with bevacizumab. Neuro-Oncology. 2017;19:1688–97.
Chaddad A, Sabri S, Niazi T, Abdulkarim B. Prediction of survival with multi-scale radiomic analysis in glioblastoma patients. Med Biol Eng Comput. 2018;56:2287–300.
Bickelhaupt S, Jaeger PF, Laun FB, Lederer W, Daniel H, Kuder TA, et al. Radiomics based on adapted diffusion kurtosis imaging helps to clarify most mammographic findings suspicious for cancer. Radiology. 2018;287:170273.
Lim WK, Er MJ. Classification of mammographic masses using generalized dynamic fuzzy neural networks. Med Phys. 2004;31:1288–95.
Sapate SG, Mahajan A, Talbar SN, Sable N, Desai S, Thakur M. Radiomics based detection and characterization of suspicious lesions on full field digital mammograms. Comput Methods Prog Biomed. 2018;163:1–20.
Chen W, Giger ML, Lan L, Bick U. Computerized interpretation of breast MRI: investigation of enhancement-variance dynamics. Med Phys. 2004;31:1076–82.
Sahiner B, Chan HP, Roubidoux MA, Helvie MA, Hadjiiski LM, Ramachandran A, et al. Computerized characterization of breast masses on three-dimensional ultrasound volumes. Med Phys. 2004;31:744–54.
Nandi RJ, Nandi AK, Rangayyan R, Scutt D. Genetic programming and feature selection for classification of breast masses in mammograms. Conf Proc IEEE Eng Med Biol Soc. 2006;44:3021–4.
Cui J, Sahiner B, Chan HP, Nees A, Paramagul C, Hadjiiski LM, et al. A new automated method for the segmentation and characterization of breast masses on ultrasound images. Med Phys. 2009;36:1553–65.
Moon WK, Shen YW, Huang CS, Chiang LR, Chang RF. Computer-aided diagnosis for the classification of breast masses in automated whole breast ultrasound images. Ultrasound Med Biol. 2011;37:539–48.
Ramos-Pollán R, Guevara-López MA, Suárez-Ortega C, Díaz-Herrero G, Franco-Valiente JM, Rubio-Del-Solar M, et al. Discovering mammography-based machine learning classifiers for breast cancer diagnosis. J Med Syst. 2012;36:2259–69.
Cai H, Peng Y, Ou C, Chen M, Li L. Diagnosis of breast masses from dynamic contrast-enhanced and diffusion-weighted MR: a machine learning approach. PLoS One. 2014;9:e87387.
Dash R, Roy S, Beura S, Majhi B. Classification of mammogram using two-dimensional discrete orthonormal S-transform for breast cancer detection. Healthc Technol Lett. 2015;2:46–51.
Cai H, Liu L, Peng Y, Wu Y, Li L. Diagnostic assessment by dynamic contrast-enhanced and diffusion-weighted magnetic resonance in differentiation of breast lesions under different imaging protocols. BMC Cancer. 2014;14:1–12.
Ma W, Ji Y, Qi L, Guo X, Jian X, Liu P. Breast cancer Ki67 expression prediction by DCE-MRI radiomics features. Clin Radiol. 2018;73:909.e1–5.
Li H, Zhu Y, Burnside ES, Huang E, Drukker K, Hoadley KA, et al. Quantitative MRI radiomics in the prediction of molecular classifications of breast cancer subtypes in the TCGA/TCIA data set. npj Breast Cancer. 2016;2:16012.
Liang C, Cheng Z, Huang Y, He L, Chen X. An MRI-based radiomics classifier for preoperative prediction of Ki-67 status in breast cancer. Acad Radiol. 2018;25:1111–7.
Dong Y, Feng Q, Yang W, Lu Z, Deng C, Zhang L, et al. Preoperative prediction of sentinel lymph node metastasis in breast cancer based on radiomics of T2-weighted fat-suppression and diffusion-weighted MRI. Eur Radiol. 2018;28:582–91.
Liu C, Ding J, Spuhler K, Gao Y, Serrano Sosa M, Moriarty M, et al. Preoperative prediction of sentinel lymph node metastasis in breast cancer by radiomic signatures from dynamic contrast-enhanced MRI. J Magn Reson Imaging. 2019;49:131–40.
Braman NM, Etesami M, Prasanna P, Dubchuk C, Gilmore H, Tiwari P, et al. Intratumoral and peritumoral radiomics for the pretreatment prediction of pathological complete response to neoadjuvant chemotherapy based on breast DCE-MRI. Breast Cancer Res. 2017;19:1–14.
Fan M, Wu G, Cheng H, Zhang J, Shao G, Li L. Radiomic analysis of DCE-MRI for prediction of response to neoadjuvant chemotherapy in breast cancer patients. Eur J Radiol. 2017;94:140–7.
Park H, Lim Y, Ko ES, Cho H-H, Lee JE, Han B-K, et al. Radiomics signature on magnetic resonance imaging: association with disease-free survival in patients with invasive breast cancer. Clin Cancer Res. 2018;24:4705–14.
Huang SY, Franc BL, Harnish RJ, Liu G, Mitra D, Copeland TP, et al. Exploration of PET and MRI radiomic features for decoding breast cancer phenotypes and prognosis. NPJ Breast Cancer. 2018;4:24.
Liu Y, Zhang Y, Cheng R, Liu S, Qu F, Yin X, et al. Radiomics analysis of apparent diffusion coefficient in cervical cancer: a preliminary study on histological grade evaluation. J Magn Reson Imaging. 2019;49:1–11.
De Bernardi E, Buda A, Guerra L, Vicini D, Elisei F, Landoni C, et al. Radiomics of the primary tumour as a tool to improve 18F-FDG-PET sensitivity in detecting nodal metastases in endometrial cancer. EJNMMI Res. 2018;8:1–9.
Reuzé S, Orlhac F, Chargari C, Nioche C, Limkin E, Riet F, et al. Prediction of cervical cancer recurrence using textural features extracted from 18F-FDG PET images acquired with different scanners. Oncotarget. 2017;8:43169–79.
Schernberg A, Reuze S, Orlhac F, Buvat I, Dercle L, Sun R, et al. A score combining baseline neutrophilia and primary tumor SUVpeakmeasured from FDG PET is associated with outcome in locally advanced cervical cancer. Eur J Nucl Med Mol Imaging. 2018;45:187–95.
Altazi BA, Fernandez DC, Zhang GG, Hawkins S, Naqvi SM, Kim Y, et al. Investigating multi-radiomic models for enhancing prediction power of cervical cancer treatment outcomes. Phys Medica. 2018;46:180–8.
Lucia F, Visvikis D, Desseroit M-C, Miranda O, Malhaire J-P, Robin P, et al. Prediction of outcome using pretreatment 18F-FDG PET/CT and MRI radiomics in locally advanced cervical cancer treated with chemoradiotherapy. Eur J Nucl Med Mol Imaging. 2018;45:768–86.
Huang X, Cheng Z, Huang Y, Liang CC, He L, Ma Z, et al. CT-based radiomics signature to discriminate high-grade from low- grade colorectal adenocarcinoma. Acad Radiol RA. 2018;25:1285–97.
Yang L, Dong D, Fang M, Zhu Y, Zang Y, Liu Z, et al. Can CT-based radiomics signature predict KRAS/NRAS/BRAF mutations in colorectal cancer? Eur Radiol. 2018;28:2058–67.
Liang C, Huan Y, He L, Chen X, Ma Z, Dong D, et al. The development and validation of a CT-based radiomics signature for the preoperative discrimination of stage I-II and stage III-IV colorectal cancer. Oncotarget. 2016;7:31401–12.
Huang YQ, Liang CH, He L, Tian J, Liang CS, Chen X, et al. Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer. J Clin Oncol. 2016;34:2157–64.
Carvalho S, Leijenaar RTH, Troost EGC, van Timmeren JE, Oberije C, van Elmpt W, et al. 18F-fluorodeoxyglucose positron-emission tomography (FDG-PET)-radiomics of metastatic lymph nodes and primary tumor in non-small cell lung cancer (NSCLC) - a prospective externally validated study. PLoS One. 2018;13:e0192859.
Cusumano D, Dinapoli N, Boldrini L, Chiloiro G, Gatta R, Masciocchi C, et al. Fractal-based radiomic approach to predict complete pathological response after chemo-radiotherapy in rectal cancer. Radiol Med. 2018;123:286–95.
Liu Z, Zhang XY, Shi YJ, Wang L, Zhu HT, Tang Z, et al. Radiomics analysis for evaluation of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Clin Cancer Res. 2017;23:7253–62.
Bibault JE, Giraud P, Durdux C, Taieb J, Berger A, Coriat R, et al. Deep learning and radiomics predict complete response after neo-adjuvant chemoradiation for locally advanced rectal cancer. Sci Rep. 2018;8:1–8.
Cui Y, Yang X, Shi Z, Yang Z, Du X, Zhao Z, et al. Radiomics analysis of multiparametric MRI for prediction of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Eur Radiol. 2019;29:1211–20.
Meng Y, Zhang Y, Dong D, Li C, Liang X, Zhang C, et al. Novel radiomic signature as a prognostic biomarker for locally advanced rectal cancer. J Magn Reson Imaging. 2018;48:605–14.
Tan X, Ma Z, Yan L, Ye W, Liu Z, Liang C. Radiomics nomogram outperforms size criteria in discriminating lymph node metastasis in resectable esophageal squamous cell carcinoma. Eur Radiol. 2019;29:392–400.
Shen C, Liu Z, Wang Z, Guo J, Zhang H, Wang Y, et al. Building CT radiomics based nomogram for preoperative esophageal cancer patients lymph node metastasis prediction. Transl Oncol. 2018;11:815–24.
Qu J, Shen C, Qin J, Wang Z, Liu Z, Guo J, et al. The MR radiomic signature can predict preoperative lymph node metastasis in patients with esophageal cancer. Eur Radiol. 2019;29:906–14.
Hou Z, Li S, Ren W, Liu J, Yan J, Wan S. Radiomic analysis in T2W and SPAIR T2W MRI: predict treatment response to chemoradiotherapy in esophageal squamous cell carcinoma. J Thorac Dis. 2018;10:2256–67.
Moyo K, Porter C, Chilima B, Mwenda R, Kabue M, Zungu L, et al. Quantifying local tumor morphological changes with Jacobian map for prediction of pathologic tumor response to chemo-radiotherapy in locally advanced esophageal cancer. Phys Med Biol. 2016;4:1–19.
Ypsilantis P-P, Siddique M, Sohn H-M, Davies A, Cook G, Goh V, et al. Predicting response to neoadjuvant chemotherapy with PET imaging using convolutional neural networks. PLoS One. 2015;10:e0137036.
Larue RTHM, Klaassen R, Jochems A, Leijenaar RTH, Hulshof MCCM, van Berge Henegouwen MI, et al. Pre-treatment CT radiomics to predict 3-year overall survival following chemoradiotherapy of esophageal cancer. Acta Oncol (Madr). 2018;57:1475–81.
Chen T, Ning Z, Xu L, Feng X, Han S, Roth HR, et al. Radiomics nomogram for predicting the malignant potential of gastrointestinal stromal tumours preoperatively. Eur Radiol. 2019;29:1074–82.
Bektas CT, Kocak B, Yardimci AH, Turkcanoglu MH, Yucetas U, Koca SB, et al. Clear cell renal cell carcinoma: machine learning-based quantitative computed tomography texture analysis for prediction of Fuhrman nuclear grade. Eur Radiol. 2019;29:1153–63.
Peng J, Zhang J, Zhang Q, Xu Y, Zhou J, Liu L. A radiomics nomogram for preoperative prediction of microvascular invasion risk in hepatitis b virus-related hepatocellular carcinoma. Diagn Interv Radiol. 2018;24:121–7.
Zhang J, Wang Y, Dong Y, Wang Y. Computer-aided diagnosis of cervical lymph nodes on ultrasonography. Comput Biol Med. 2008;38:234–43.
Bogowicz M, Riesterer O, Ikenberg K, Stieb S, Moch H, Studer G, et al. Computed tomography radiomics predicts HPV status and local tumor control after definitive Radiochemotherapy in head and neck squamous cell carcinoma. Int J Radiat Oncol Biol Phys. 2017;99:921–8.
Leijenaar RT, Bogowicz M, Jochems A, Hoebers FJ, Wesseling FW, Huang SH, et al. Development and validation of a radiomic signature to predict HPV (p16) status from standard CT imaging: a multicenter study. Br J Radiol. 2018;91:20170498.
Al Ajmi E, Forghani B, Reinhold C, Bayat M, Forghani R. Spectral multi-energy CT texture analysis with machine learning for tissue classification: an investigation using classification of benign parotid tumours as a testing paradigm. Eur Radiol. 2018;28:2604–11.
Wang G, He L, Yuan C, Huang Y, Liu Z, Liang C. Pretreatment MR imaging radiomics signatures for response prediction to induction chemotherapy in patients with nasopharyngeal carcinoma. Eur J Radiol. 2018;98:100–6.
Zhang B, Tian J, Dong D, Gu D, Dong Y, Zhang L, et al. Radiomics features of multiparametric MRI as novel prognostic factors in advanced nasopharyngeal carcinoma. Clin Cancer Res. 2017;23:4259–69.
Leijenaar RTH, Carvalho S, Hoebers FJP, Aerts HJWL, van Elmpt WJC, Huang SH, et al. External validation of a prognostic CT-based radiomic signature in oropharyngeal squamous cell carcinoma. Acta Oncol. 2015;54:1423–9.
Bogowicz M, Riesterer O, Stark LS, Studer G, Unkelbach J, Guckenberger M, et al. Comparison of PET and CT radiomics for prediction of local tumor control in head and neck squamous cell carcinoma. Acta Oncol (Madr). 2017;56:1531–6.
Vallières M, Kay-Rivest E, Perrin LJ, Liem X, Furstoss C, Aerts HJWL, et al. Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer. Sci Rep. 2017;7:1–14.
Ouyang F-S, Guo B-L, Zhang B, Dong Y-H, Zhang L, Mo X-K, et al. Exploration and validation of radiomics signature as an independent prognostic biomarker in stage III-IVb nasopharyngeal carcinoma. Oncotarget. 2017;8:74869–79.
van Dijk LV, Thor M, Steenbakkers RJHM, Apte A, Zhai TT, Borra R, et al. Parotid gland fat related magnetic resonance image biomarkers improve prediction of late radiation-induced xerostomia. Radiother Oncol. 2018;128:459–66.
Sun R, Limkin EJ, Vakalopoulou M, Dercle L, Champiat S, Han SR, et al. A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol. 2018;19:1180–91.
Colen RR, Fujii T, Bilen MA, Kotrotsou A, Abrol S, Hess KR, et al. Radiomics to predict immunotherapy-induced pneumonitis: proof of concept. Investig New Drugs. 2018;36:601–7.
Alilou M, Beig N, Orooji M, Rajiah P, Velcheti V, Clinic C, et al. An integrated segmentation and shape-based classification scheme for distinguishing adenocarcinomas from granulomas on lung CT. Med Phys. 2017;44:3556–69.
Peikert T, Duan F, Rajagopalan S, Karwoski RA, Clay R, Robb RA, et al. Novel high-resolution computed tomography-based radiomic classifier for screen-identified pulmonary nodules in the National Lung Screening Trial. PLoS One. 2018;13:1–15.
Tu SJ, Wang CW, Pan KT, Wu YC, Wu CT. Localized thin-section CT with radiomics feature extraction and machine learning to classify early-detected pulmonary nodules from lung cancer screening. Phys Med Biol. 2018;63:065005.
Carter BW, Godoy MC, Erasmus JJ. Predicting malignant nodules from screening CTs. J Thorac Oncol. 2016;11:2045–7.
Orooji M, Alilou M, Rakshit S, Beig N, Khorrami MH, Rajiah P, et al. Combination of computer extracted shape and texture features enables discrimination of granulomas from adenocarcinoma on chest computed tomography. J Med Imaging. 2018;5:1.
Chen X, Fang M, Dong D, Wei X, Liu L, Xu X, et al. A radiomics signature in preoperative predicting degree of tumor differentiation in patients with non–small cell lung cancer. Acad Radiol. 2018;25:1548–55.
Yuan M, Zhang Y-D, Pu X-H, Zhong Y, Li H, Wu J-F, et al. Comparison of a radiomic biomarker with volumetric analysis for decoding tumour phenotypes of lung adenocarcinoma with different disease-specific survival. Eur Radiol. 2017;27(11):4857–65.
Zhang L, Chen B, Liu X. Quantitative biomarkers for prediction of epidermal growth factor receptor mutation in non- small cell lung cancer. Transl Oncol. 2018;11:94–101.
Grossmann P, Stringfield O, El-Hachem N, Bui MM, Rios Velazquez E, Parmar C, et al. Defining the biological basis of radiomic phenotypes in lung cancer. Elife. 2017;6:e23421.
Rios Velazquez E, Parmar C, Liu Y, Coroller TP, Cruz G, Stringfield O, et al. Somatic mutations drive distinct imaging phenotypes in lung cancer. Cancer Res. 2017;77:3922–30.
She Y, Zhang L, Zhu H, Dai C, Xie D, Xie H, et al. The predictive value of CT-based radiomics in differentiating indolent from invasive lung adenocarcinoma in patients with pulmonary nodules. Eur Radiol. 2018;28:5121–8.
Wu W, Parmar C, Grossmann P, Quackenbush J, Lambin P, Bussink J, et al. Exploratory study to identify radiomics classifiers for lung cancer histology. Front Oncol. 2016;6:1–11.
Kirienko M, Cozzi L, Rossi A, Voulaz E, Antunovic L, Fogliata A, et al. Ability of FDG PET and CT radiomics features to differentiate between primary and metastatic lung lesions. Eur J Nucl Med Mol Imaging. 2018;45:1649–60.
Raniery J, Junior F, Koenigkam-santos M, Enrique F, Cipriano G, Todorovic A, et al. Radiomics-based features for pattern recognition of lung cancer histopathology and metastases. Comput Methods Prog Biomed. 2018;159:23–30.
Song SH, Park H, Lee G, Lee HY, Sohn I, Kim HS, et al. Imaging phenotyping using radiomics to predict micropapillary pattern within lung adenocarcinoma. J Thorac Oncol. 2017;12:624–32.
Fan L, Fang MJ, Li ZB, Tu WT, Wang SP, Chen WF. et al, Radiomics signature: a biomarker for the preoperative discrimination of lung invasive adenocarcinoma manifesting as a ground-glass nodule. Eur Radiol. 2018:1–9.
Zhu X, Dong D, Chen Z, Fang M, Zhang L, Song J, et al. Radiomic signature as a diagnostic factor for histologic subtype classification of non-small cell lung cancer. Eur Radiol. 2018;28:2772–8.
Ma Y, Feng W, Wu Z, Liu M, Zhang F, Liang Z, et al. Intra-tumoural heterogeneity characterization through texture and colour analysis for differentiation of non-small cell lung carcinoma subtypes. Phys Med Biol. 2018;63:165018.
Xue X, Yang Y, Huang Q, Cui F, Lian Y, Zhang S, et al. Use of a radiomics model to predict tumor invasiveness of pulmonary adenocarcinomas appearing as pulmonary ground-glass nodules. Biomed Res Int. 2018;2018:6803971.
Yuan M, Liu JY, Zhang T, Zhang YD, Li H, Yu TF. Prognostic impact of the findings on thin-section computed tomography in stage i lung adenocarcinoma with visceral pleural invasion. Sci Rep. 2018;8:1–9.
Liu Y, Kim J, Balagurunathan Y, Hawkins S, Stringfield O, Schabath MB, et al. Prediction of pathological nodal involvement by CT-based radiomic features of the primary tumor in patients with clinically node-negative peripheral lung adenocarcinomas. Med Phys. 2018;45:2518–26.
Yang X, Pan X, Liu H, Gao D, He J, Liang W, et al. A new approach to predict lymph node metastasis in solid lung adenocarcinoma: a radiomics nomogram. J Thorac Dis. 2018;10:S807–19.
Coroller TTP, Grossmann P, Hou Y, Rios Velazquez E, Leijenaar RTH, Hermann G, et al. CT-based radiomic signature predicts distant metastasis in lung. Radiother Oncol. 2015;114:345–50.
Coroller TP, Agrawal V, Huynh E, Narayan V, Lee SW, Mak RH, et al. Radiomic-based pathological response prediction from primary tumors and lymph nodes in NSCLC. J Thorac Oncol. 2017;12:467–76.
Huynh E, Coroller TP, Narayan V, Agrawal V, Hou Y, Romano J, et al. CT-based radiomic analysis of stereotactic body radiation therapy patients with lung cancer. Int J Radiat. 2016;120:258–66.
Huang Y, Liu Z, He L, Chen X, Pan D, Ma Z, et al. Radiomics signature: a potential biomarker for the prediction of. Radiology. 2016;281:1–11.
Yu W, Tang C, Hobbs BP, Li X, Koay EJ, Wistuba II, et al. Development and validation of a predictive radiomics model for clinical outcomes in stage I non-small cell lung cancer. Int J Radiat Oncol Biol Phys. 2018;102:1090–7.
Virginia BM, Laura F, Silvia R, Roberto F, Francesco F, Eva H, et al. Prognostic value of histogram analysis in advanced non-small cell lung cancer: a radiomic study. Oncotarget. 2018;9:1906–14.
Arshad MA, Thornton A, Lu H, Tam H, Wallitt K, Rodgers N, et al. Discovery of pre-therapy 2-deoxy-2-18F-fluoro-D-glucose positron emission tomography-based radiomics classifiers of survival outcome in non-small-cell lung cancer patients. Eur J Nucl Med Mol Imaging. 2018;46:455–66.
Tang C, Hobbs B, Amer A, Li X, Behrens C, Canales JR, et al. Development of an immune-pathology informed radiomics model for non-small cell lung cancer. Sci Rep. 2018;8:1–9.
Carvalho S, Leijenaar RTH, Troost EGC, Van Timmeren JE, Oberije C, Van Elmpt W, et al. 18FDG-PET-radiomics of metastatic lymph nodes and primary tumour in NSCLC - a prospective externally validated study. PLoS One. 2018;13:e0192859.
Mattonen SA, Palma DA, Johnson C, Louie AV, Landis M, Rodrigues G, et al. Detection of local cancer recurrence after stereotactic ablative radiation therapy for lung cancer: physician performance versus radiomic assessment. Int J Radiat Oncol Biol Phys. 2016;94:1121–8.
van Timmeren JE, Leijenaar RTH, van Elmpt W, Reymen B, Oberije C, Monshouwer R, et al. Survival prediction of non-small cell lung cancer patients using radiomics analyses of cone-beam CT images. Radiother Oncol. 2017;123:363–9.
Chaddad A, Desrosiers C, Toews M, Abdulkarim B. Predicting survival time of lung cancer patients using radiomic analysis. Oncotarget. 2017;8:104393–407.
Kirienko M, Cozzi L, Antunovic L, Lozza L, Fogliata A, Voulaz E, et al. Prediction of disease-free survival by the PET/CT radiomic signature in non-small cell lung cancer patients undergoing surgery. Eur J Nucl Med Mol Imaging. 2018;45:207–17.
Krafft SP, Rao A, Stingo F, Briere TM, Court LE, Liao Z, et al. The utility of quantitative CT radiomics features for improved prediction of radiation pneumonitis. Med Phys. 2018;45:5317–24.
Guo J, Liu Z, Shen C, Li Z, Yan F, Tian J, et al. MR-based radiomics signature in differentiating ocular adnexal lymphoma from idiopathic orbital inflammation. Eur Radiol. 2018;28:3872–81.
Lucidarme O, Akakpo JP, Granberg S, Sideri M, Levavi H, Schneider A, et al. A new computer-aided diagnostic tool for non-invasive characterisation of malignant ovarian masses: results of a multicentre validation study. Eur Radiol. 2010;20:1822–30.
Chakraborty J, Midya A, Gazit L, Attiyeh M, Langdon-Embry L, Allen PJ, et al. CT radiomics to predict high-risk intraductal papillary mucinous neoplasms of the pancreas. Med Phys. 2018;45:5019–29.
Bonekamp D, Kohl S, Wiesenfarth M, Schelb P, Radtke JP, Götz M, et al. Radiomic machine learning for characterization of prostate lesions with MRI: comparison to ADC values. Radiology. 2018;289:128–37. https://doi.org/10.1148/radiol.2018173064.
Shiradkar R, Ghose S, Jambor I, Taimen P, Ettala O, Purysko AS, et al. Radiomic features from pretreatment biparametric MRI predict prostate cancer biochemical recurrence: preliminary findings. J Magn Reson Imaging. 2018;48:1626–36.
Corino VDA, Montin E, Messina A, Casali PG, Gronchi A, Marchianò A, et al. Radiomic analysis of soft tissues sarcomas can distinguish intermediate from high-grade lesions. J Magn Reson Imaging. 2018;47:829–40.
Wu Y, Xu L, Yang P, Lin N, Huang X, Pan W, et al. Survival prediction in high-grade osteosarcoma using radiomics of diagnostic computed tomography. EBioMedicine. 2018;34:27–34.
Iyatomi H, Oka H, Celebi ME, Hashimoto M, Hagiwara M, Tanaka M, et al. An improved internet-based melanoma screening system with dermatologist-like tumor area extraction algorithm. Comput Med Imaging Graph. 2008;32:566–79.
Liu YY, Ishikawa H, Chen M, Wollstein G, Duker JS, Fujimoto JG, et al. Computerized macular pathology diagnosis in spectral domain optical coherence tomography scans based on multiscale texture and shape features. Investig Ophthalmol Vis Sci. 2011;52:8316–22.
Bodduluri S, Newell JD, Hoffman EA, Reinhardt JM. Registration-based lung mechanical analysis of chronic obstructive pulmonary disease (COPD) using a supervised machine learning framework. Acad Radiol. 2013;20:527–36.
Bocchi L, Nori J. Shape analysis of microcalcifications using radon transform. Med Eng Phys. 2007;29:691–8.
Li Z, Wang Y, Yu J, Guo Y, Cao W. Deep learning based radiomics (DLR) and its usage in noninvasive IDH1 prediction for low grade glioma. Sci Rep. 2017;7:1–11.
Lao J, Chen Y, Li ZC, Li Q, Zhang J, Liu J, et al. A deep learning-based radiomics model for prediction of survival in glioblastoma multiforme. Sci Rep. 2017;7:1–8.
Ning Z, Luo J, Li Y, Han S, Feng Q, Xu Y, et al. Pattern classification for gastrointestinal stromal tumors by integration of radiomics and deep convolutional features. IEEE J Biomed Heal Informatics. 2018:1–1.
Huynh BQ, Li H, Giger ML. Digital mammographic tumor classification using transfer learning from deep convolutional neural networks. J Med Imaging. 2016;3:034501.
Kirienko M, Sollini M, Silvestri G, Mognetti S, Voulaz E, Antunovic L, et al. Convolutional neural networks promising in lung cancer T-parameter assessment on baseline FDG-PET/CT. Contrast Media Mol Imaging. 2018;2018:1–6.
Park SH, Han K. Methodologic guide for evaluating clinical performance and effect of artificial intelligence technology for medical diagnosis and prediction. Radiology. 2018;286:800–9.
Srinivasan GN, Shobha G. Statistical texture analysis. Proc WORLD Acad Sci Eng Technol. 2008;36:1264–9.
Way TW, Sahiner B, Hadjiiski LM, Chan H-P. Effect of finite sample size on feature selection and classification: a simulation study. Med Phys. 2010;37:907–20.
Chan H-P, Sahiner B, Wagner RF, Petrick N. Classifier design for computer-aided diagnosis: effects of finite sample size on the mean performance of classical and neural network classifiers. Med Phys. 1999;26:2654–68.
Mazurowski MA, Habas PA, Zurada JM, Lo JY, Baker JA, Tourassi GD. Training neural network classifiers for medical decision making: the effects of imbalanced datasets on classification performance. Neural Netw. 2008;21:427–36.
Brazauskas R, Logan BR. Observational studies: matching or regression? Biol Blood Marrow Transplant. 2016;22:557–63.
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017;14:749–62.
Vallieres M, Zwanenburg A, Badic B, Cheze-Le Rest C, Visvikis D, Hatt M. Responsible radiomics research for faster clinical translation. J Nucl Med. 2018;59(2):189–93.
Zwanenburg A, Leger S, Vallières M, Löck S. Initiative for the IBS. Image biomarker standardisation initiative. arXiv:1612.07003.
Acknowledgments
We thank Prof. Wim Oyen for advice and critical discussion. We thank Eng. Luca Antiga for inspiring talks and collaboration. We thank Dr. Marra for her support in setting the literature search strategy.
Funding
M. Kirienko's PhD scholarship is supported by the AIRC grant (IG-2016-18585).
Author information
Authors and Affiliations
Contributions
MS and MK planned the study, performed article search and selection, analyzed the data, and drafted the manuscript. LA gave support during study search and selection. AC conceptualized the study, coordinated the study workflow, and critically reviewed the manuscript. All the authors reviewed the final version of the text.
Corresponding author
Ethics declarations
Ethics
This manuscript does not involve any study on animals or humans.
Conflict of interest
The authors declare no conflict of interest related to the present work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Advanced Image Analyses (Radiomics and Artificial Intelligence).
Electronic supplementary material
ESM 1
(DOCX 31 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Sollini, M., Antunovic, L., Chiti, A. et al. Towards clinical application of image mining: a systematic review on artificial intelligence and radiomics. Eur J Nucl Med Mol Imaging 46, 2656–2672 (2019). https://doi.org/10.1007/s00259-019-04372-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-019-04372-x