Abstract
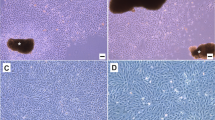
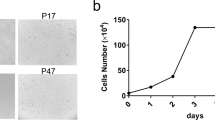
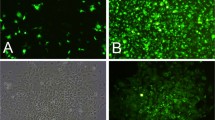
The Chinese tree shrew holds a great potential as a viable animal model in biomedical research, especially for infectious diseases and neuropsychiatric disorders. A thorough understanding of the innate immunity, which represents the first line that defends the host against viral infection, of the Chinese tree shrew, is needed. However, the progress is hindered by the lack of a proper cell line for research usage. In this study, we established a cell line that is applicable to the study of tree shrew innate immune responses against viral infections. The Chinese tree shrew primary renal cells (TSPRCs) were immortalized by simian virus 40 large T antigen (SV40LT) transduction, and the immortalized cells were termed TSR6 (tree shrew renal cell #6). TSR6 showed a similar morphology to TSPRCs and expressed the epithelial cell-specific marker cytokeratin 18 (KRT18). In addition, TSR6 could be transfected by transfection reagent and was suitable for CRISPR/Cas9-mediated gene editing. Infection of Newcastle disease virus (NDV) or herpes simplex virus 1 (HSV-1) in TSR6 induced the mRNA expression of tree shrew interferon-β (tIFNB1) and myxovirus resistance protein 1 (tMx1) in a dose- and time-dependent manner. Collectively, we successfully established a tree shrew renal cell line and demonstrated that this cell line was suitable for the study of the innate immune response to viral infections.





Similar content being viewed by others
References
Ahuja D, Saenz-Robles MT, Pipas JM (2005) SV40 large T antigen targets multiple cellular pathways to elicit cellular transformation. Oncogene 24:7729–7745. https://doi.org/10.1038/sj.onc.1209046
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124:783–801. https://doi.org/10.1016/j.cell.2006.02.015
Amako Y, Tsukiyama-Kohara K, Katsume A, Hirata Y, Sekiguchi S, Tobita Y, Hayashi Y, Hishima T, Funata N, Yonekawa H, Kohara M (2010) Pathogenesis of hepatitis C virus infection in Tupaia belangeri. J Virol 84:303–311. https://doi.org/10.1128/jvi.01448-09
Banerjee A, Rapin N, Miller M, Griebel P, Zhou Y, Munster V, Misra V (2016) Generation and characterization of Eptesicus fuscus (Big brown bat) kidney cell lines immortalized using the myotis polyomavirus large T-antigen. J Virol Methods 237:166–173. https://doi.org/10.1016/j.jviromet.2016.09.008
Borden EC, Sen GC, Uze G, Silverman RH, Ransohoff RM, Foster GR, Stark GR (2007) Interferons at age 50: past, current and future impact on biomedicine. Nat Rev Drug Discov 6:975–990. https://doi.org/10.1038/nrd2422
Chen L, Wang G, Zhu YN, Xiang H, Wang W (2016) Advances and perspectives in the application of CRISPR/Cas9 in insects. Zool Res 37:220–228. https://doi.org/10.13918/j.issn.2095-8137.2016.4.220
de Leeuw O, Peeters B (1999) Complete nucleotide sequence of Newcastle disease virus: evidence for the existence of a new genus within the subfamily Paramyxovirinae. J Gen Virol 80(Pt 1):131–136. https://doi.org/10.1099/0022-1317-80-1-131
Fan Y, Huang ZY, Cao CC, Chen CS, Chen YX, Fan DD, He J, Hou HL, Hu L, Hu XT, Jiang XT, Lai R, Lang YS, Liang B, Liao SG, Mu D, Ma YY, Niu YY, Sun XQ, Xia JQ, Xiao J, Xiong ZQ, Xu L, Yang L, Zhang Y, Zhao W, Zhao XD, Zheng YT, Zhou JM, Zhu YB, Zhang GJ, Wang J, Yao YG (2013) Genome of the Chinese tree shrew. Nat Commun 4:1426. https://doi.org/10.1038/ncomms2416
Fan Y, Yu D, Yao YG (2014) Tree shrew database (TreeshrewDB): a genomic knowledge base for the Chinese tree shrew. Sci Rep 4:7145. https://doi.org/10.1038/srep07145
Foddis R, De Rienzo A, Broccoli D, Bocchetta M, Stekala E, Rizzo P, Tosolini A, Grobelny JV, Jhanwar SC, Pass HI, Testa JR, Carbone M (2002) SV40 infection induces telomerase activity in human mesothelial cells. Oncogene 21:1434–1442. https://doi.org/10.1038/sj.onc.1205203
Hubrecht R, Kirkwood J, Hubrecht R, Kirkwood J (2010) The UFAW handbook on the care and management of laboratory and other research animals. Wiley-Blackwell, Hoboken
Jin LF, Li JS (2016) Generation of genetically modified mice using CRISPR/Cas9 and haploid embryonic stem cell systems. Zool Res 37:205–213. https://doi.org/10.13918/j.issn.2095-8137.2016.4.205
Kayesh MEH, Kitab B, Sanada T, Hayasaka D, Morita K, Kohara M, Tsukiyama-Kohara K (2017) Susceptibility and initial immune response of Tupaia belangeri cells to dengue virus infection. Infect Genet Evol 51:203–210. https://doi.org/10.1016/j.meegid.2017.04.003
Li JP, Liao Y, Zhang Y, Wang JJ, Wang LC, Feng K, Li QH, Liu LD (2014) Experimental infection of tree shrews (Tupaia belangeri) with Coxsackie virus A16. Zool Res 35:485–491. https://doi.org/10.13918/j.issn.2095-8137.2014.6.485
Li L, Li Z, Wang E, Yang R, Xiao Y, Han H, Lang F, Li X, Xia Y, Gao F, Li Q, Fraser NW, Zhou J (2016) Herpes simplex virus 1 infection of tree shrews differs from that of mice in the severity of acute infection and viral transcription in the peripheral nervous system. J Virol 90:790–804. https://doi.org/10.1128/jvi.02258-15
Li CH, Yan LZ, Ban WZ, Tu Q, Wu Y, Wang L, Bi R, Ji S, Ma YH, Nie WH, Lv LB, Yao YG, Zhao XD, Zheng P (2017) Long-term propagation of tree shrew spermatogonial stem cells in culture and successful generation of transgenic offspring. Cell Res 27:241–252. https://doi.org/10.1038/cr.2016.156
Li R, Zanin M, Xia X, Yang Z (2018) The tree shrew as a model for infectious diseases research. J Thorac Dis 10:S2272–s2279. https://doi.org/10.21037/jtd.2017.12.121
Luo X, Li M, SU B (2016) Application of the genome editing tool CRISPR/Cas9 in non-human primates. Zool Res 37:214–219. https://doi.org/10.13918/j.issn.2095-8137.2016.4.214
Ma X, Wong AS, Tam HY, Tsui SY, Chung DL, Feng B (2018) In vivo genome editing thrives with diversified CRISPR technologies. Zool Res 39:58–71. https://doi.org/10.24272/j.issn.2095-8137.2017.012
Maqsood MI, Matin MM, Bahrami AR, Ghasroldasht MM (2013) Immortality of cell lines: challenges and advantages of establishment. Cell Biol Int 37:1038–1045. https://doi.org/10.1002/cbin.10137
McNab F, Mayer-Barber K, Sher A, Wack A, O’Garra A (2015) Type I interferons in infectious disease. Nat Rev Immunol 15:87–103. https://doi.org/10.1038/nri3787
Minin AA, Moldaver MV (2008) Intermediate vimentin filaments and their role in intracellular organelle distribution. Biochemistry (Mosc) 73:1453–1466. https://doi.org/10.1134/S0006297908130063
Mork C, van Deurs B, Petersen OW (1990) Regulation of vimentin expression in cultured human mammary epithelial cells. Differentiation 43:146–156. https://doi.org/10.1111/j.1432-0436.1990.tb00441.x
Nelson WG (1983) The 50- and 58-kdalton keratin classes as molecular markers for stratified squamous epithelia: cell culture studies. J Cell Biol 97:244–251. https://doi.org/10.1083/jcb.97.1.244
Oliveros JC, Franch M, Tabas-Madrid D, San-Leon D, Montoliu L, Cubas P, Pazos F (2016) Breaking-Cas-interactive design of guide RNAs for CRISPR-Cas experiments for ENSEMBL genomes. Nucleic Acids Res 44:W267–W271. https://doi.org/10.1093/nar/gkw407
Ramboer E, De Craene B, De Kock J, Vanhaecke T, Berx G, Rogiers V, Vinken M (2014) Strategies for immortalization of primary hepatocytes. J Hepatol 61:925–943. https://doi.org/10.1016/j.jhep.2014.05.046
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Ruan P, Yang C, Su J, Cao J, Ou C, Luo C, Tang Y, Wang Q, Yang F, Shi J, Lu X, Zhu L, Qin H, Sun W, Lao Y, Li Y (2013) Histopathological changes in the liver of tree shrew (Tupaia belangeri chinensis) persistently infected with hepatitis B virus. Virol J 10:333. https://doi.org/10.1186/1743-422x-10-333
Schneider WM, Chevillotte MD, Rice CM (2014) Interferon-stimulated genes: a complex web of host defenses. Annu Rev Immunol 32:513–545. https://doi.org/10.1146/annurev-immunol-032713-120231
Su C, Zheng C (2017) Herpes simplex virus 1 abrogates the cGAS/STING-mediated cytosolic DNA-sensing pathway via its virion host shutoff protein, UL41. J Virol 91:e02414–e02416. https://doi.org/10.1128/JVI.02414-16
Tsukiyama-Kohara K, Kohara M (2014) Tupaia belangeri as an experimental animal model for viral infection. Exp Anim 63:367–374. https://doi.org/10.1538/expanim.14-0007
Wang J, Hu G, Lin Z, He L, Xu L, Zhang Y (2014) Characteristic and functional analysis of a newly established porcine small intestinal epithelial cell line. PLoS One 9:e110916. https://doi.org/10.1371/journal.pone.0110916
Xiao J, Liu R, Chen CS (2017) Tree shrew (Tupaia belangeri) as a novel laboratory disease animal model. Zool Res 38:127–137. https://doi.org/10.24272/j.issn.2095-8137.2017.033
Xu L, Chen S-Y, Nie W-H, Jiang X-L, Yao Y-G (2012) Evaluating the phylogenetic position of Chinese tree shrew (Tupaia belangeri chinensis) based on complete mitochondrial genome: implication for using tree shrew as an alternative experimental animal to primates in biomedical research. J Genet Genomics 39:131–137. https://doi.org/10.1016/j.jgg.2012.02.003
Xu L, Fan Y, Jiang XL, Yao YG (2013a) Molecular evidence on the phylogenetic position of tree shrews. Zool Res 34:70–76. https://doi.org/10.3724/sp.j.1141.2013.02070
Xu L, Zhang Y, Liang B, Lu LB, Chen CS, Chen YB, Zhou JM, Yao YG (2013b) Tree shrews under the spot light: emerging model of human diseases. Zool Res 34:59–69. https://doi.org/10.3724/sp.j.1141.2013.02059
Xu L, Yu D, Peng L, Fan Y, Chen J, Zheng YT, Wang C, Yao YG (2015) Characterization of a MAVS ortholog from the Chinese tree shrew (Tupaia belangeri chinensis). Dev Comp Immunol 52:58–68. https://doi.org/10.1016/j.dci.2015.04.014
Xu L, Yu D, Fan Y, Peng L, Wu Y, Yao YG (2016) Loss of RIG-I leads to a functional replacement with MDA5 in the Chinese tree shrew. Proc Natl Acad Sci U S A 113:10950–10955. https://doi.org/10.1073/pnas.1604939113
Xu L, Peng L, Gu T, Yu D, Yao Y-G (2019) The 3′UTR of human MAVS mRNA contains multiple regulatory elements for the control of protein expression and subcellular localization. Biochim Biophys Acta 1862:47–57. https://doi.org/10.1016/j.bbagrm.2018.10.017
Yang ZF, Zhao J, Zhu YT, Wang YT, Liu R, Zhao SS, Li RF, Yang CG, Li JQ, Zhong NS (2013) The tree shrew provides a useful alternative model for the study of influenza H1N1 virus. Virol J 10:111. https://doi.org/10.1186/1743-422x-10-111
Yao YG (2017) Creating animal models, why not use the Chinese tree shrew (Tupaia belangeri chinensis)? Zool Res 38:118–126. https://doi.org/10.24272/j.issn.2095-8137.2017.032
Yao YL, Yu D, Xu L, Fan Y, Wu Y, Gu T, Chen J, Lv LB, Yao YG (2019) Molecular characterization of the 2′,5′-oligoadenylate synthetase family in the Chinese tree shrew (Tupaia belangeri chinensis). Cytokine. https://doi.org/10.1016/j.cyto.2018.11.009
Yu D, Xu L, Liu XH, Fan Y, Lu LB, Yao YG (2014) Diverse interleukin-7 mRNA transcripts in Chinese tree shrew (Tupaia belangeri chinensis). PLoS One 9:e99859. https://doi.org/10.1371/journal.pone.0099859
Yu D, Wu Y, Xu L, Fan Y, Peng L, Xu M, Yao YG (2016) Identification and characterization of toll-like receptors (TLRs) in the Chinese tree shrew (Tupaia belangeri chinensis). Dev Comp Immunol 60:127–138. https://doi.org/10.1016/j.dci.2016.02.025
Zheng YT, Yao YG, Xu L (2014) Basic biology and disease models of tree shrews. Yunnan Science and Technology Press, Kunming
Funding
This study was supported by the National Natural Science Foundation of China (U1402224 to Y-GY), Yunnan Province (2018FB046 to DY), and the Chinese Academy of Sciences (CAS zsys-02 to Y-GY).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All animal studies with tree shrew were approved by the Institutional Animal Care and Use Committee of Kunming Institute of Zoology, Chinese Academy of Sciences (Approval No: SYDW20110315001).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gu, T., Yu, D., Li, Y. et al. Establishment and characterization of an immortalized renal cell line of the Chinese tree shrew (Tupaia belangeri chinesis). Appl Microbiol Biotechnol 103, 2171–2180 (2019). https://doi.org/10.1007/s00253-019-09615-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09615-3