Abstract
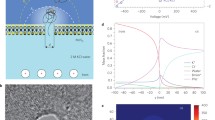
Solid-state nanopores are considered an attractive basis for single-molecule DNA sequencing. At present, one obstacle to be overcome is the improvement of their temporal resolution, with the DNA molecules remaining in the sensing volume of the nanopore for a long period of time. Here, we used a composite system of a concentration gradient of LiCl in solution and a nanofiber mesh to slow the DNA perforation speed. Compared to different alkali metal solutions with the same concentration, LiCl can extend the dwell time to 20 ms, five times longer than NaCl and KCl. Moreover, as the concentration gradient increases, the dwell time can be tuned from dozens of milliseconds to more than 100 ms. When we introduce a nanofiber mesh layer on top of the pore in the asymmetric solution, the DNA molecules get retarded by 162–185 \(\upmu \)s/nt, which is three orders of magnitude slower than the bare nanopore. At the same time, because the molecule absorption region becomes larger at the pore vicinity, the higher molecule capture rate improves the detection efficiency.





Similar content being viewed by others
References
Akahori R, Haga T et al (2014) Slowing single-stranded DNA translocation through a solid-state nanopore by decreasing the nanopore diameter. Nanotechnology 25:275501
Akahori R, Yanagi I et al (2017) Discrimination of three types of homopolymers in single-stranded DNA with solid-state nanopores through external control of the DNA motion. Sci Rep 7:9073
Alibakhshi MA, Halman JR et al (2017) Picomolar fingerprinting of nucleic acid nanoparticles using solid-state nanopores. ACS Nano 11:9701–9710
Branton D, Deamer DW et al (2008) The potential and challenges of nanopore sequencing. Nat Biotechnol 26:1146–1153
Feng J, Liu K et al (2015) Identification of single nucleotides in \(\text{ MoS }_2\) nanopores. Nat Nanotechnol 10:1070–1076
Fologea D, Uplinger J et al (2005) Slowing DNA translocation in a solid-state nanopore. Nano Lett 5:1734–1737
Garaj S, Hubbard W et al (2010) Graphene as a subnanometre trans-electrode membrane. Nature 467:190–193
He Y, Tsutsui M et al (2011) Controlling DNA translocation through gate modulation of nanopore wall surface charges. ACS Nano 5:5509–5518
He Y, Tsutsui M et al (2013) Mechanism of how salt-gradient-induced charges affect the translocation of DNA molecules through a nanopore. Biophys J 105:776–782
Hyun C, Kaur H et al (2017) A tip-attached tuning fork sensor for the control of DNA translocation through a nanopore. Rev Sci Instrum 88:025001
Iqbal SM, Bashir R (2011) Nanopores: sensing and fundamental biological interactions. In: Hall AR, Dekker C (eds) Molecular detection and force spectroscopy in solid-state nanopores with integrated optical tweezers. Springer, Boston, p 37
Knust S, Kreft D et al (2017) Measuring DNA translocation forces through \(\text{ MoS }_2\) nanopores with optical tweezers. Mater Today Proc 4:S168–S173
Kowalczyk SW, Grosberg AY et al (2011) Modelling the conductance and DNA blockade of solid-state nanopores. Nanotechnology 22:315101
Kowalczyk SW, Wells DB et al (2012) Slowing down DNA translocation through a nanopore in lithium chloride. Nano Lett 12:1038–1044
Kwok H, Briggs K et al (2014) Nanopore fabrication by controlled dielectric breakdown. PLoS One 9:e92880
Li J, Gershow M et al (2003) DNA molecules and configurations in a solid-state nanopore microscope. Nat Mater 2:611–615
Liu K, Feng J et al (2014) Automically thin molybdenum disulfide nanopores with high sensitivity for DNA translocation. ACS Nano 8:2504–2511
Liu Y, Yobas L (2016) Slowing DNA translocation in a nanofluidic field-effect transistor. ACS Nano 10:3985–3994
Mirsaidov U, Comer J et al (2010) Slowing the translocation of double-stranded DNA using a nanopore smaller than the double helix. Nanotechnology 21:395501
Muthukumar M (2010) Theory of capture rate in polymer translocation. J Chem Phys 132:195101
Nelson EM, Li H et al (2014) Direct, concurrent measurements of the forces and currents affecting DNA in a nanopore with comparable topography. ACS Nano 8:5484–5493
Nova IC, Derrington IM et al (2017) Investigating asymmetric salt profiles for nanopore DNA sequencing with biological porin MspA. PLoS One 12:e0181599
Ross PD, Scruggs RL (1964) Electrophoresis of DNA. III. The effect of several univalent electrolytes on the mobility of DNA. Biopolymers 2:231–236
Sha J, Shi H et al (2017) Salt gradient improving signal-to-noise ration in solid-state nanopore. ACS Sens 2:506–512
Squires AH, Hersey JS et al (2013) A nanopore-nanofiber mesh biosensor to control DNA translocation. J Am Chem Soc 135:16304–16307
Tajmir-Riahi HA, Messaoudi S (1992) The effects of monovalent cations Li\(^+\), Na\(^+\), K\(^+\), \(\text{ NH }_4^+\), Rb\(^+\) and Cs\(^+\) on the solid and solution structures of the nucleic acid components. Metal ion binding and sugar conformation. J Biomol Struct Dyn 10:345–365
Tang Z, Liang Z et al (2015) Gel mesh as “brake” to slow down DNA translocation through solid-state nanopores. Nanoscale 7:13207–13214
Trepagnier EH, Radenovic A et al (2007) Controlling DNA capture and propagation through artificial nanopores. Nano Lett 7:2824–2830
Tsutsui M, He Y et al (2012) Transverse electric field dragging of DNA in a nanochannel. Sci Rep 2:394
Wanunu M, Morrison W et al (2010) Electrostatic focusing of unlabelled DNA in to nanoscale pores using a salt gradient. Nat Nanotechnol 5:160–165
Waugh M, Carlsen A et al (2015) Interfacing solid-state nanopores with gel media to slow DNA translocations. Electrophoresis 36:1759–1767
Wong CTA, Muthukumar M (2007) Polymer capture by electro-osmotic flow of oppositely charged nanopores. J Chem Phys 126:164903
Zhang H, Zhao Q et al (2013) Slowing down DNA translocation through solid-state nanopores by pressure. Small 9:4112–4117
Zhao Y, Xie W et al (2018) Slowing down DNA translocation by a nanofiber meshed layer. J Phys D Appl Phys 51:045402
Zhou D, Deng Y et al (2016) Rectification of ion current determined by the nanopore geometry: experiments and modelling. Chin Phys Lett 33:108501
Zoysa RSS, Jayawardhana DA et al (2009) Slowing DNA translocation through nanopores using a solution containing organic salts. J Phys Chem B 113:13332–13336
Acknowledgements
Acknowledgements are made to National Natural Science Foundations of China (NNSFC 61701474, 61471336, 31501084, 51503206), and West Light Foundation of CAS and Chongqing Sci. & Technol. Commission (cstc2017jcyjAX0320) for support of this work.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yan, H., Zhou, D., Shi, B. et al. Slowing down DNA translocation velocity using a LiCl salt gradient and nanofiber mesh. Eur Biophys J 48, 261–266 (2019). https://doi.org/10.1007/s00249-019-01350-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-019-01350-x