Abstract
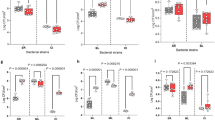
Bacterial social interaction is a potential influencing factor in determining the fate of invading pathogens in diverse environments. In this study, interactions between two representative resident species (Bacillus subtilis and Pseudomonas putida) and a leading food-borne disease causative pathogen (Vibrio parahaemolyticus) were examined. An antagonistic effect toward V. parahaemolyticus was observed for B. subtilis but not for P. putida. However, the relative richness of the pathogen remained rather high in B. subtilis co-cultures and was, unexpectedly, not sensitive to the initial inoculation ratios. Furthermore, two approaches were found to be efficient at modulating the relative richness of the pathogen. (1) The addition of trace glycerol and manganese to Luria-Bertani medium (LBGM) reduced the richness of V. parahaemolyticus in the co-culture with B. subtilis and in contrast, increased its richness in the co-culture with P. putida, although it did not affect the growth of V. parahaemolyticus by its own. (2) The relative richness of V. parahaemolyticus on semisolid medium decreased significantly as a function of an agar gradient, ranging from 0 to 2%. Furthermore, we explored the molecular basis of bacterial interaction through transcriptomic analysis. In summary, we investigated the interactions between a pathogen invader and two resident bacteria species, showing that the different influences on a pathogen by different types of interactions can be modulated by chemicals and medium fluidity.





Similar content being viewed by others
References
Flemming H-C, Wingender J, Szewzyk U, Steinberg P, Rice SA, Kjelleberg S (2016) Biofilms: an emergent form of bacterial life. Nat Rev Microbiol 14:563–575. https://doi.org/10.1038/nrmicro.2016.94
Donlan RM, Costerton JW (2002) Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 15:167–193. https://doi.org/10.1128/CMR.15.2.167-193.2002
Hall-Stoodley L, Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2:95–108. https://doi.org/10.1038/nrmicro821
Burmølle M, Ren D, Bjarnsholt T, Sørensen SJ (2014) Interactions in multispecies biofilms: do they actually matter? Trends Microbiol 22:84–91. https://doi.org/10.1016/j.tim.2013.12.004
DeLeon S, Clinton A, Fowler H, Everett J, Horswill AR, Rumbaugh KP (2014) Synergistic interactions of Pseudomonas aeruginosa and Staphylococcus aureus in an in vitro wound model. Infect Immun 82:4718–4728. https://doi.org/10.1128/IAI.02198-14
Hotterbeekx A, Kumar-Singh S, Goossens H, Malhotra-Kumar S (2017) In vivo and in vitro interactions between Pseudomonas aeruginosa and Staphylococcus spp. Front Cell Infect Microbiol 7:106. https://doi.org/10.3389/fcimb.2017.00106
Trejo-Hernández A, Andrade-Domínguez A, Hernández M, Encarnación S (2014) Interspecies competition triggers virulence and mutability in Candida albicans–Pseudomonas aeruginosa mixed biofilms. ISME J 8:1974–1988. https://doi.org/10.1038/ismej.2014.53
de Sablet T, Chassard C, Bernalier-Donadille A et al (2009) Human microbiota-secreted factors inhibit Shiga toxin synthesis by Enterohemorrhagic Escherichia coli O157:H7. Infect Immun 77:783–790. https://doi.org/10.1128/IAI.01048-08
Curtis MM, Hu Z, Klimko C, Narayanan S, Deberardinis R, Sperandio V (2014) The gut commensal Bacteroides thetaiotaomicron exacerbates enteric infection through modification of the metabolic landscape. Cell Host Microbe 16:759–769. https://doi.org/10.1016/j.chom.2014.11.005
Goswami K, Chen C, Xiaoli L, Eaton KA, Dudley EG (2015) Coculture of Escherichia coli O157:H7 with a Nonpathogenic E. Coli strain increases toxin production and virulence in a germfree mouse model. Infect Immun 83:4185–4193. https://doi.org/10.1128/IAI.00663-15
Vos M, Wolf AB, Jennings SJ, Kowalchuk GA (2013) Micro-scale determinants of bacterial diversity in soil. FEMS Microbiol Rev 37:936–954. https://doi.org/10.1111/1574-6976.12023
van Elsas JD, Hill P, Chronakova A et al (2007) Survival of genetically marked Escherichia coli O157 : H7 in soil as affected by soil microbial community shifts. ISME J 1:204–214. https://doi.org/10.1038/ismej.2007.21
van Elsas JD, Chiurazzi M, Mallon CA, Elhottova D, Kristufek V, Salles JF (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc Natl Acad Sci 109:1159–1164. https://doi.org/10.1073/pnas.1109326109
Yao Z, Wang H, Wu L, Wu J, Brookes PC, Xu J (2014) Interaction between the microbial community and invading Escherichia coli O157:H7 in soils from vegetable fields. Appl Environ Microbiol 80:70–76. https://doi.org/10.1128/AEM.03046-13
Cooley MB, Miller WG, Mandrell RE (2003) Colonization of Arabidopsis thaliana with Salmonella enterica and Enterohemorrhagic Escherichia coli O157:H7 and competition by Enterobacter asburiae. Appl Environ Microbiol 69:4915–4926. https://doi.org/10.1128/AEM.69.8.4915-4926.2003
You Y, Rankin SC, Aceto HW, Benson CE, Toth JD, Dou Z (2006) Survival of Salmonella enterica Serovar Newport in manure and manure-amended soils. Appl Environ Microbiol 72:5777–5783. https://doi.org/10.1128/AEM.00791-06
Liu NT, Bauchan GR, Francoeur CB, Shelton DR, Lo YM, Nou X (2016) Ralstonia insidiosa serves as bridges in biofilm formation by foodborne pathogens listeria monocytogenes, Salmonella enterica, and Enterohemorrhagic Escherichia coli. Food Control 65:14–20. https://doi.org/10.1016/j.foodcont.2016.01.004
Bradford SA, Morales VL, Zhang W, Harvey RW, Packman AI, Mohanram A, Welty C (2013) Transport and fate of microbial pathogens in agricultural settings. Crit Rev Environ Sci Technol 43:775–893. https://doi.org/10.1080/10643389.2012.710449
Wang R, Huang J, Zhang W, Lin G, Lian J, Jiang L, Lin H, Wang S, Wang S (2011) Detection and identification of Vibrio parahaemolyticus by multiplex PCR and DNA–DNA hybridization on a microarray. J Genet Genomics 38:129–135. https://doi.org/10.1016/j.jgg.2011.02.002
Wang R, Zhong Y, Gu X, Yuan J, Saeed AF, Wang S (2015) The pathogenesis, detection, and prevention of Vibrio parahaemolyticus. Front Microbiol 6:144. https://doi.org/10.3389/fmicb.2015.00144
Su Y-C, Liu C (2007) Vibrio parahaemolyticus: a concern of seafood safety. Food Microbiol 24:549–558. https://doi.org/10.1016/j.fm.2007.01.005
Hara-Kudo Y, Sugiyama K, Nishibuchi M, Chowdhury A, Yatsuyanagi J, Ohtomo Y, Saito A, Nagano H, Nishina T, Nakagawa H, Konuma H, Miyahara M, Kumagai S (2003) Prevalence of pandemic thermostable direct Hemolysin-producing Vibrio parahaemolyticus O3:K6 in seafood and the coastal environment in Japan. Appl Environ Microbiol 69:3883–3891. https://doi.org/10.1128/AEM.69.7.3883-3891.2003
Li B, Ju F, Cai L, Zhang T (2015) Profile and fate of bacterial pathogens in sewage treatment plants revealed by high-throughput metagenomic approach. Environ Sci Technol 49:10492–10502. https://doi.org/10.1021/acs.est.5b02345
Li X, Harwood VJ, Nayak B, Staley C, Sadowsky MJ, Weidhaas J (2015) A novel microbial source tracking microarray for pathogen detection and fecal source identification in environmental systems. Environ Sci Technol 49:7319–7329. https://doi.org/10.1021/acs.est.5b00980
Mallon CA, Roux X, Doorn GS et al (2018) The impact of failure: unsuccessful bacterial invasions steer the soil microbial community away from the invader’s niche. ISME J 1:728–741. https://doi.org/10.1038/s41396-017-0003-y
Shemesh M, Chai Y (2013) A combination of glycerol and manganese promotes biofilm formation in Bacillus subtilis via histidine kinase KinD signaling. J Bacteriol 195:2747–2754. https://doi.org/10.1128/JB.00028-13
Smith MD, Roheim CA, Crowder LB, Halpern BS, Turnipseed M, Anderson JL, Asche F, Bourillon L, Guttormsen AG, Khan A, Liguori LA, McNevin A, O'Connor MI, Squires D, Tyedmers P, Brownstein C, Carden K, Klinger DH, Sagarin R, Selkoe KA (2010) Sustainability and global seafood. Science 327:784–786. https://doi.org/10.1126/science.1185345
Sheehan MC, Burke TA, Navas-Acien A, Breysse PN, McGready J, Fox MA (2014) Global methylmercury exposure from seafood consumption and risk of developmental neurotoxicity: a systematic review. Bull World Health Organ 92:254–269F. https://doi.org/10.2471/BLT.12.116152
Nielsen AT, Tolker-Nielsen T, Barken KB, Molin S (2000) Role of commensal relationships on the spatial structure of a surface-attached microbial consortium. Environ Microbiol 2:59–68. https://doi.org/10.1046/j.1462-2920.2000.00084.x
Garbeva P, Silby MW, Raaijmakers JM, Levy SB, Boer W (2011) Transcriptional and antagonistic responses of Pseudomonas fluorescens Pf0-1 to phylogenetically different bacterial competitors. ISME J 5:973–985. https://doi.org/10.1038/ismej.2010.196
Blin K, Wolf T, Chevrette MG, Lu X, Schwalen CJ, Kautsar SA, Suarez Duran HG, de los Santos ELC, Kim HU, Nave M, Dickschat JS, Mitchell DA, Shelest E, Breitling R, Takano E, Lee SY, Weber T, Medema MH (2017) antiSMASH 4.0—improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res 45:W36–W41. https://doi.org/10.1093/nar/gkx319
Weller DM (2007) Pseudomonas biocontrol agents of Soilborne pathogens: looking back over 30 years. Phytopathology 97:250–256. https://doi.org/10.1094/PHYTO-97-2-0250
Wall DH, Nielsen UN, Six J (2015) Soil biodiversity and human health. Nature 528:69–76. https://doi.org/10.1038/nature15744
Kim W, Levy SB, Foster KR (2016) Rapid radiation in bacteria leads to a division of labour. Nat Commun 7:10508. https://doi.org/10.1038/ncomms10508
Oliveira NM, Martinez-Garcia E, Xavier J, Durham WM, Kolter R, Kim W, Foster KR (2015) Biofilm formation as a response to ecological competition. PLoS Biol 13:e1002191. https://doi.org/10.1371/journal.pbio.1002191
Ren D, Madsen JS, Sørensen SJ, Burmølle M (2015) High prevalence of biofilm synergy among bacterial soil isolates in cocultures indicates bacterial interspecific cooperation. ISME J 9:81–89. https://doi.org/10.1038/ismej.2014.96
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Hall T (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Ren D, Madsen JS, de la Cruz-Perera CI et al (2014) High-throughput screening of multispecies biofilm formation and quantitative PCR-based assessment of individual species proportions, useful for exploring interspecific bacterial interactions. Microb Ecol 68:146–154. https://doi.org/10.1007/s00248-013-0315-z
Zhong S, Joung J-G, Zheng Y, Chen YR, Liu B, Shao Y, Xiang JZ, Fei Z, Giovannoni JJ (2011) High-throughput Illumina strand-specific RNA sequencing library preparation. Cold Spring Harb Protoc 2011:pdb.prot5652:940–949. https://doi.org/10.1101/pdb.prot5652
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS J Integr Biol 16:284–287. https://doi.org/10.1089/omi.2011.0118
Acknowledgements
This work was supported by the National Key Research Program of China (2016YFD0800206) and National Natural Science Foundation of China (41522106).
Author information
Authors and Affiliations
Contributions
P.C. conceived the study. C.H.G. and M.Z. performed experiments. C.H.G., Y.W., Q.H. and P.C. analyzed and interpreted the data. C.H.G. and P. C. wrote the paper with the help of all authors.
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Supplementary Figure S1
(PDF 200 kb)
Supplementary Figure S2
(PDF 42.5 kb)
Supplementary Figure S3
(PDF 185 kb)
Supplementary Figure S4
(PDF 242 kb)
Supplementary Figure S5
(PDF 342 kb)
Supplementary Table S1
(XLSX 49.4 kb)
Supplementary Table S2
(XLSX 11.4 kb)
Supplementary Figure S6
(PDF 464 kb)
Rights and permissions
About this article
Cite this article
Gao, CH., Zhang, M., Wu, Y. et al. Divergent Influence to a Pathogen Invader by Resident Bacteria with Different Social Interactions. Microb Ecol 77, 76–86 (2019). https://doi.org/10.1007/s00248-018-1207-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1207-z