Abstract
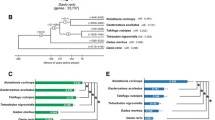
Gene expression of antifreeze proteins (AFPs) facilitates various species of fish to survive in near-freezing seawater. The marine sculpin genus Myoxocephalus (Teleostei; Perciformes) is widely distributed from Arctic to Subarctic and is known to express type I AFP. Here, we clarify gene structures and the antifreeze activity of three Myoxocephalus species and discuss their cold adaptation and their geographic movements across a global-level time scale. These fishes were collected during winter in 2014–2015 at Hokkaido and Alaska. A total of 14–16 amino acid sequences were determined as type I AFP, some of which exhibit the same amino acid sequence among species. The Subarctic species contains many amino acid sequences, and the antifreeze activities are lowest. In contrast, the Arctic species contains a few active proteins. Mitochondrial DNA analysis suggests that the Atlantic and Arctic species migrated to those oceans 7.9 and 3.0 million years ago (Mya), respectively. Since the AFP sequences of Myoxocephalus are similar to each other regardless of their difference in the timing of migration, the ancestral species of this genus may have developed the AFP gene(s) approximately 7.9 Mya, before speciation within the genus. It is speculated that these sculpins survived the cold environment in the Arctic Ocean during the Pliocene–Pleistocene by acquisition of the type I AFP in their body fluids.



Similar content being viewed by others
References
Akashi H (1994) Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics 136:927–935
Amaoka K, Nakaya K, Yabe M (2011) The pictorial book of all of the fishes of Hokkaido. The Hokkaido Shimbun Press, Sapporo
Baardsnes J, Kondejewski LH, Hodges RS, Chao H, Kay C, Davies PL (1999) New ice-binding face for type I antifreeze protein. FEBS J 463:87–91
Baardsnes J, Jelokhani-Niaraki M, Kondejewski LH, Kuiper MJ, Kay CM, Hodges RS, Davies PL (2001) Antifreeze protein from shorthorn sculpin: identification of the ice-binding surface. Protein Sci 10:2566–2576. https://doi.org/10.1101/ps.26501
Bang JK, Lee JH, Murugan RN, Lee SG, Do H, Koh HY, Shim HE, Kim HC, Kim HJ (2013) Antifreeze peptides and glycopeptides, and their derivatives: potential uses in biotechnology. Mar Drugs 11:2013–2041. https://doi.org/10.3390/md11062013
Begun DJ, Whitley P, Todd BL, Waldrip-Dail HM, Clark AG (2000) Molecular population genetics of male accessory gland proteins in Drosophila. Genetics 156:1879–1888
Benton R (2015) Multigene family evolution: perspectives from insect chemoreceptors. Trends Ecol Evol 30:590–600. https://doi.org/10.1016/j.tree.2015.07.009
Briggs JC (2003) Marine centres of origin as evolutionary engines. J Biogeogr 30:1–18
Briggs JC (2007a) Marine biogeography and ecology: invasions and introductions. J Biogeogr 34:193–198. https://doi.org/10.1111/j.1365-2699.2006.01632.x
Briggs JC (2007b) Marine longitudinal biodiversity: causes and conservation. Divers Distrib 13:544–555. https://doi.org/10.1111/j.1472-4642.2007.00362.x
Briggs JC, Bowen BW, McClain C (2013) Marine shelf habitat: biogeography and evolution. J Biogeogr 40:1023–1035. https://doi.org/10.1111/jbi.12082
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Chakrabartty A, Hew CL, Shears MA, Fletcher GL (1988) Primary structures of the alanine-rich antifreeze polypeptides from grubby sculpin, Myoxocephalus aenaeus. Can J Zool 66:403–408
Chao H, Houston ME, Hodges RS, Kay CM, Sykes BD, Loewen MC, Davies PL, Sönnichsen FD (1997) A diminished role for hydrogen bonds in antifreeze protein binding to ice. Biochemistry 36:14652–14660
Chen L, DeVries AL, Cheng CH (1997) Convergent evolution of antifreeze glycoproteins in Antarctic notothenioid fish and Arctic cod. Proc Natl Acad Sci USA 94:3817–3822
Cheng CH (1988) Evolution of the diverse antifreeze proteins. Curr Opin Genet Dev 8:715–720
Davies PL (2014) Ice-binding proteins: a remarkable diversity of structures for stopping and starting ice growth. Trends Biochem Sci 39:548–555. https://doi.org/10.1016/j.tibs.2014.09.005
Desjardins M, Graham LA, Davies PL, Fletcher GL (2012) Antifreeze protein gene amplification facilitated niche exploitation and speciation in wolffish. FEBS J 279:2215–2230. https://doi.org/10.1111/j.1742-4658.2012.08605.x
DeVries AL (1982) Biological antifreeze agents in coldwater fishes. Comp Biochem Physiol A Mol Integr Physiol 73:627–640
DeVries AL (1984) Role of glycopeptides and peptides in inhibition of crystallization of water in polar fishes. Philos Trans R Soc Lond Ser B Biol Sci 304:575–588
DeVries AL, Wohlschlag DE (1969) Freezing Resistance in Some Antarctic Fishes. Science 163:1073–1075. https://doi.org/10.1126/science.163.3871.1073
Duman JG, DeVries AL (1974) The role of macromolecular antifreezes in cold water fishes. Comp Biochem Physiol A Mol Integr Physiol 52:193–199
Duman JG, DeVries AL (1976) Isolation, characterization, and physical properties of protein antifreezes from the winter flounder, Pseudopleuronectes americanus. Comp Biochem Physiol A Mol Integr Physiol 54:375–380. https://doi.org/10.1016/0305-0491(76)90260-1
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.2307/2408678
Froese R, Pauly D (2015) FishBase. World Wide Web electronic publication. http://www.fishbase.org/search.php. Accessed 13 Jun 2015
Gong Z, Ewart KV, Hu Z, Flethcer GL, Hew CL (1996) Skin antifreeze protein genes of the winter flounder, Pleuronectes americanus, encode distinct and active polypeptides without the secretory signal and prosequences. J Biol Chem 271:4106–4112. https://doi.org/10.1074/jbc.271.8.4106
Graham LA, Hobbs RS, Fletcher GL, Davies PL (2013) Helical antifreeze proteins have independently evolved in fishes on four occasions. PLoS One 8:e81285. https://doi.org/10.1371/journal.pone.0081285
Harada N, Ahagon N, Sakamoto T, Uchida M, Ikehara M, Shibata Y (2006) Rapid fluctuation of alkenone temperature in the southwestern Okhotsk Sea during the past 120 ky. Glob Planet Change 53:29–46. https://doi.org/10.1016/j.gloplacha.2006.01.010
Harrison MK, Crespi BJ (1999) Phylogenetics of cancer crabs (Crustacea: Decapoda: Brachyura). Mol Phylogen Evol 12:186–199. https://doi.org/10.1006/mpev.1998.0608
Haymet A, Ward LG, Harding MM, Knight CA (1998) Valine substituted winter flounder ‘antifreeze’: preservation of ice growth hysteresis. FEBS Lett 430:301–306
Hew CL, Fletcher GL, Ananthanarayanan VS (1980) Antifreeze proteins from the shorthorn sculpin, Myoxocephalus scorpius: isolation and characterization. Can J Biochem 58:377–383. https://doi.org/10.1139/o80-049
Hew CL, Slaughter D, Joshi SB, Fletcher GL, Ananthanarayanan VS (1984) Antifreeze polypeptides from the Newfoundland ocean pout, Macrozoarces americanus: presence of multiple and compositionally diverse components. J Comp Physiol B 155:81–88. https://doi.org/10.1007/BF00688795
Hew CL, Wang NC, Joshi S, Fletcher GL, Scott GK, Hayes PH, Buettner B, Davies PL (1988) Multiple genes provide the basis for antifreeze protein diversity and dosage in the ocean pout, Macrozoarces americanus. J Biol Chem 263:12049–12055
Kafanov A (1999) Neogene Macoma (Bivalvia, Tellinidae) migration from the Pacific to the Atlantic through the Bering Strait: taxonomic and biogeographic remarks. Boll-Soc Paleontol Ital 38:77–86
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kimura MR, Yanagimoto T, Munehara H (2007) Maternal identification of hybrid eggs in Hexagrammos spp. by means of multiplex amplified product length polymorphism of mitochondrial DNA. Aquatic Biol 1:187–194. https://doi.org/10.3354/ab00019
Kontula T, Väinölä R (2003) Relationships of palearctic and nearctic ‘glacial relict’Myoxocephalus sculpins from mitochondrial DNA data. Mol Ecol 12:3179–3184. https://doi.org/10.1046/j.1365-294X.2003.01963.x
Li X-M, Trinh K-Y, Hew CL (1985) Structure of an antifreeze polypeptide and its precursor from the ocean pout, Macrozoarces americanus. J Biol Chem 260:12904–12909
Low WK, Miao M, Ewart KV, Yang DSC, Fletcher GL, Hew CL (1998) Skin-type antifreeze protein from the shorthorn sculpin, Myoxocephalus scorpius: expression and characterization of a M r 9,700 recombinant protein. J Biol Chem 273:23098–23103. https://doi.org/10.1074/jbc.273.36.23098
Low WK, Lin Q, Stathakis C, Miao M, Fletcher GL, Hew CL (2001) Isolation and characterization of skin-type, type I antifreeze polypeptides from the longhorn sculpin, Myoxocephalus octodecemspinosus. J Biol Chem 276:11582–11589. https://doi.org/10.1074/jbc.M009293200
Mahatabuddin S, Hanada Y, Nishimiya Y, Miura A, Kondo H, Davies PL, Tsuda S (2017) Concentration-dependent oligomerization of an alpha-helical antifreeze polypeptide makes it hyperactive. Sci Rep 7:42501. https://doi.org/10.1038/srep42501
Marincovich LJ, Gladenkov AY (1999) Evidence for an early opening of the Bering Strait. Nature 397:149–151
Marqusee S, Baldwin RL (1987) Helix stabilization by Glu-… Lys + salt bridges in short peptides of de novo design. Proc Natl Acad Sci 84:8898–8902
Mecklenburg CW, Mecklenburg TA, Thorsteinson LK (2002) Fishes of Alaska. American Fisheries Society, Maryland
Mecklenburg CW, Stein DL, Sheiko BA, Chernova NV, Mecklenburg TA, Holladay BA (2007) Russian-American long-term census of the Arctic: Benthic fishes Trawled in the Chukchi Sea and Bering Strait, August 2004. Northwest Nat 88:168–187. https://doi.org/10.1898/1051-1733(2007)88%5b168:rlcota%5d2.0.co;2
Mecklenburg CW, Møller PR, Steinke D (2011) Biodiversity of arctic marine fishes: taxonomy and zoogeography. Mar Biodivs 41:109–140. https://doi.org/10.1007/s12526-010-0070-z
Nelson JS, Grande TC, Wilson MV (2016) Fishes of the World. Wiley, Hoboken
Nishimiya Y, Sato R, Takamichi M, Miura A, Tsuda S (2005) Co-operative effect of the isoforms of type III antifreeze protein expressed in Notched-fin eelpout, Zoarces elongatus Kner. FEBS J 272:482–492. https://doi.org/10.1111/j.1742-4658.2004.04490.x
Nishimiya Y, Mie Y, Hirano Y, Kondo H, Miura A, Tsuda S (2008) Mass preparation and technological development of an antifreeze protein. Synthesiology 1:7–14
Podlesnykh AV, Moreva IN (2014) Variability and relationships of the Far Eastern species of sculpins Myoxocephalus and Megalocottus (Cottidae) based on mtDNA markers and karyological data. Russ J Genet 50:949–956. https://doi.org/10.1134/s1022795414090117
Raymo M (1994) The initiation of Northern Hemisphere glaciation. Annu Rev Earth Pl Sci 22:353–383
Raymond JA, DeVries AL (1977) Adsorption inhibition as a mechanism of freezing resistance in polar fishes. Proc Natl Acad Sci USA 74:2589–2593
Raymond JA, Lin Y, DeVries AL (1975) Glycoprotein and protein antifreezes in two Alaskan fishes. J Exp Zool A Comp Exp Biol 193:125–130. https://doi.org/10.1002/jez.1401930112
Reisman HM, Fletcher GL, Kao MH, Shears MA (1987) Antifreeze proteins in the grubby sculpin, Myoxocephalus aenaeus and the tomcod, Microgadus tomcod: comparisons of seasonal cycles. Environ Biol Fishes 18:295–301
Schmid KJ, Aquadro CF (2001) The evolutionary analysis of “Orphans” from the Drosophila genome identifies rapidly diverging and incorrectly annotated genes. Genetics 159:589–598
Slaughter D, Fletcher GL, Ananthanarayanan VS, Hew CL (1981) Antifreeze proteins from the sea raven, Hemitripterus americanus. Further evidence for diversity among fish polypeptide antifreezes. J Biol Chem 256:2022–2026
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Stickley CE, St John K, Koç N, Jordan RW, Passchier S, Pearce RB, Kearns LE (2009) Evidence for middle Eocene Arctic sea ice from diatoms and ice-rafted debris. Nature 460:376–379
Takamichi M, Nishimiya Y, Miura A, Tsuda S (2007) Effect of annealing time of an ice crystal on the activity of type III antifreeze protein. FEBS J 274:6469–6476. https://doi.org/10.1111/j.1742-4658.2007.06164.x
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tanabe AS (2011) Kakusan4 and Aminosan: two programs for comparing nonpartitioned, proportional and separate models for combined molecular phylogenetic analyses of multilocus sequence data. Mol Ecol Resour 11:914–921. https://doi.org/10.1111/j.1755-0998.2011.03021.x
Tsuda S, Miura A (2005) Antifreeze proteins originating in fishes, Japan. US Patent 2005/0019856 A1, application no. 10/496,104
Tyshenko MG, Doucet D, Walker VK (2005) Analysis of antifreeze proteins within spruce budworm sister species. Insect Mol Biol 14:319–326. https://doi.org/10.1111/j.1365-2583.2005.00562.x
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PD (2005) DNA barcoding Australia’s fish species. Philos Trans R Soc Lond B Biol Sci 360:1847–1857. https://doi.org/10.1098/rstb.2005.1716
Wierzbicki A, Taylor MS, Knight CA, Madura JD, Harrington JP, Sikes CS (1996) Analysis of shorthorn sculpin antifreeze protein stereospecific binding to (2–1 0) faces of ice. Biophys J 71:8–18
Wierzbicki A, Knight CA, Salter EA, Henderson CN, Madura JD (2008) Role of nonpolar amino acid functional groups in the surface orientation-dependent adsorption of natural and synthetic antifreeze peptides on ice. Cryst Growth Des 8:3420–3429
Yamashita Y, Miura R, Takemoto Y, Tsuda S, Kawahara H, Obata H (2003) Type II antifreeze protein from mid-latitude freshwater fish, Japanese smelt (Hypomesus nipponensis). Biosci Biotechnol Biochem 67:461–466
Yamazaki A, Markevich A, Munehara H (2013) Molecular phylogeny and zoogeography of marine sculpins in the genus Gymnocanthus (Teleostei; Cottidae) based on mitochondrial DNA sequences. Mar Biol 160:2581–2589. https://doi.org/10.1007/s00227-013-2250-4
Yokoyama R, Goto A (2005) Evolutionary history of freshwater sculpins, genus Cottus (Teleostei; Cottidae) and related taxa, as inferred from mitochondrial DNA phylogeny. Mol Phylogen Evol 36:654–668. https://doi.org/10.1016/j.ympev.2005.06.004
Zhang W, Laursen RA (1998) Structure-function relationships in a type I antifreeze polypeptide the role of threonine methyl and hydroxyl groups in antifreeze activity. J Biol Chem 273:34806–34812
Acknowledgements
We appreciate associate Prof. H. Kondo and Dr. Y. Hanada for their useful advice on antifreeze protein analyses. We also thank Prof. Y. Koya, assistant Prof. S. Awata, and Mr. N. Sato, the owner of the dive shop “GruntSculpin”, the Alaska Department of Fish and Game, the Mac Enterprise, and graduate students of Usujiri Fisheries Station for sampling assistance. Lastly, we appreciate two anonymous reviewers for their useful advice.
Funding
This study was funded by Hokkaido University Grant for Research Activities Abroad, Research Fellow of Japan Society for the Promotion of Science (JSPS Research Fellow), and a Grant-in-Aid for Scientific Research from JSPS (Grant-in-Aid 26-1530, 15K13760, 25304011, 26292098).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and institutional guidelines for the use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted. Collecting permits were provided by the Alaska Department of Fish and Game. The fishes sampled for the study were acquired by fishing and scuba diving.
Additional information
Responsible Editor: C. Eizaguirre.
Reviewed by Undisclosed experts.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yamazaki, A., Nishimiya, Y., Tsuda, S. et al. Gene expression of antifreeze protein in relation to historical distributions of Myoxocephalus fish species. Mar Biol 165, 181 (2018). https://doi.org/10.1007/s00227-018-3440-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-018-3440-x