Abstract
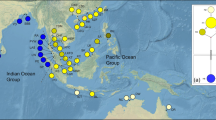
Knowledge of spatial patterns of genetic differentiation between populations is key to understanding processes in evolutionary history of biological species. Caulerpa is a genus of marine green algae, which has attracted much public attention, mainly because of the impacts of invasive species in the Mediterranean. However, very little is known about the ecological and evolutionary history of the Mediterranean native Caulerpa prolifera, a species which is currently found at sites distributed worldwide. C. prolifera provides a good model to explore the patterns of genetic diversity at different scales across the Mediterranean and Atlantic area. This study aims to investigate the biogeographical patterns of diversity and differentiation of C. prolifera in the Mediterranean, with special focus on the Mediterranean/Atlantic transition zone. We used two nuclear (ITS rDNA and the hypervariable microsatellite locus CaPr_J2) and one chloroplast (tufA) DNA markers on samples of C. prolifera from its entire range. Analyses of 51 sequences of the cpDNA tufA of C. prolifera, 87 ITS2 sequences and genotypes of 788 ramets of C. prolifera for the locus CaPr_J2 revealed three different biogeographical areas: West Atlantic, East Atlantic and a larger area representing the Mediterranean, the Mediterranean/Atlantic transition zone and a Pacific site (Bali). It was found out that the Mediterranean/Atlantic transition zone is a biogeographical boundary for C. prolifera. A lack of connectivity was revealed between Atlantic and Mediterranean types, and identical sequences found in the Mediterranean and Indo-Pacific suggest either recent gene flow along the Red Sea connection or a possible ancient Indo-Pacific origin.




Similar content being viewed by others
References
Aguirre J, Perez-Munoz AB, Sanchez-Almazo I (2006) Benthic foraminifera assemblages on the lower. Pliocene deposits of the Almeria-Nijar Basin (SE Spain). Rev Esp Micropaleontol 38(2–3):411–428
Alberto F, Massa S, Manent P, Diaz-Almela E, Arnaud-Haond S, Duarte CM, Serrão EA (2008) Genetic differentiation and secondary contact zone in the seagrass Cymodocea nodosa across the Mediterranean–Atlantic transition region. J Biogeogr 35:1270–1294
Almada VC, Oliveira RF, Gonçalves EJ, Almeida AJ, Santos RS, Wirtz P (2001) Patterns of diversity of the north-eastern Atlantic blenniid fish fauna (Pisces: Blenniidae). Glob Ecol Biogeogr 10:411–422
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Baus E, Darrock DJ, Bruford MW (2005) Gene-flow patterns in Atlantic and Mediterranean populations of the Lusitanian Sea star Asterina gibbosa. Mol Ecol 14:3373–3382
Belton GS, Prud’homme van Reine W, Huisman JM, Draisma GA, Gurgel FD (2013) Resolving phenotypic plasticity and species designation in the morphologically challenging Caulerpa racemosa–peltata complex (Chlorophyta, Caulerpaceae). J Phycol 50:32–54. doi:10.1111/jpy.12132
Benzie JAH, Price IR, Ballment E (1997) Population genetics and taxonomy of Caulerpa (Chlorophyta) from the Great Barrier Reef, Australia. J Phycol 33:491–504
Bryden HL, Candela J, Kinder TH (1994) Exchange through the Strait of Gibraltar. Prog Oceanogr 33:201–248
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Cunha AH, Varela-Álvarez E, Paulo DS, Sousa I, Serrão EA (2013) The rediscovery of Caulerpa prolifera in Ria Formosa, Portugal, 60 years after being undetected. Cah Biol Mar 54:359–364
Domingues VS, Bucciarelli G, Almada VC, Bernardi G (2005) Historical colonization and demography of the Mediterranean damselfish, Chromis chromis. Mol Ecol 14:4051–4063
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Duran S, Pascual M, Estoup A, Turon X (2004) Strong population structure in the marine sponge Crambe crambe (Poecilosclerida) as revealed by microsatellite markers. Mol Ecol 13:511–522
Durand C, Manuel M, Boudouresque CF, Meinesz A, Verlaque M, Le Parco Y (2002) Molecular data suggest a hybrid origin for the invasive Caulerpa racemosa (Caulerpales, Chlorophyta) in the Mediterranean Sea. J Evol Biol 15:122–133
Famà P, Olsen JL, Stam WT, Procaccini G (2000) High levels of intra- and inter-individual polymorphism in the rDNA ITS1 of Caulerpa racemosa (Chlorophyta). Eur J Phycol 35:349–356
Famà P, Wysor B, Kooistra WHCF, Zuccarello GC (2002) Molecular phylogeny of the genus Caulerpa (Caulerpales, Chlorophyta) inferred from chloroplast tufA gene. J Phycol 38:1040–1050
Fuertes Aguilar J, Rossello JA, Nieto Feliner G (1999) Molecular evidence for the compilospecies model of reticulate evolution in Armeria (Plumbaginaceae). Syst Biol 48:735–775
Galil BS (2009) Taking stock: inventory of alien species in the Mediterranean Sea. Biol Invasions 11:359–372
Galtier N, Gouy M, Gautier C (1996) SeaView and Phylo_win, two graphic tools for sequence alignment and molecular phylogeny. Comput Appl Biosci 12:543–548
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Guiry MD, Guiry GM (2014) AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org. Accessed 05 Nov 2014
Hrbek T, Meyer A (2003) Closing of the Tethys Sea and the phylogeny of Eurasian killifishes (Cyprinodontiformes: Cyprinodontidae). J Evol Biol 16(1):17–36
Jongma D, Campo D, Dattolo E, D’Esposito D, Duchi D, Grewe P, Huisman J, Verlaque M, Yokes M, Procaccini G (2013) Identity and origin of a slender Caulerpa taxifolia strain introduced into the Mediterranean Sea. Bot Mar 56(1):27–39
Jousson O, Pawlowski J, Zaninetti L, Meinesz A, Boudouresque CF (1998) Molecular evidence for the aquariologic origin of the green alga Caulerpa taxifolia invading the Mediterranean Sea. Mar Ecol Prog Ser 172:275–280
Jousson O, Pawlowski J, Zaninetti L, Zechman FW, Dini F, Di Guiseppe G, Woodfield R, Millar A, Meinesz A (2000) Invasive alga reaches California. Nature 408:9
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lehman R, Manhart JR (1997) A preliminary comparison of restriction fragment patterns in the genus Caulerpa (Chlorophyta) and the unique structure of the chloroplast genome of Caulerpa sertularioides. J Phycol 33:1055–1062
Lowe CD, Martin LE, Montagnesa DJS, Watts PC (2012) A legacy of contrasting spatial genetic structure on either side of the Atlantic–Mediterranean transition zone in a marine protest. Proc Natl Acad Sci USA 109(51):20998–21003
Masucci AP, Arnaud-Haond S, Eguiluz VM, Hernandez-Garcia E, Serrao EA (2012) Genetic flow directionality and geographical segregation in a Cymodocea nodosa genetic diversity network. EPJ Data Sci 1:11
Matschiner M, Salzburger W (2009) TANDEM: integrating automated allele binning into genetics and genomics workflows. Bioinformatics 25:1982–1983
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794
Meusnier I, Olsen JL, Stam WT, Destombe C, Valero M (2001) Phylogenetic analyses of Caulerpa taxifolia (Chlorophyta) and of its associated bacterial microflora provide clues to the origin of the Mediterranean introduction. Mol Ecol 10(4):931–946
Meusnier I, Valero M, Destombe C, Godé C, Desmarais E, Stam WT, Olsen JL (2002) Polymerase chain reaction-single strand conformation polymorphism analyses of nuclear and chloroplast DNA provide evidence for recombination, multiple introductions and nascent speciation in the Caulerpa taxifolia complex. Mol Ecol 11:2317–2325
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Parilla G, Kinder TH (1992) The physical oceanography of the Alborán Sea. Rep Meteorol Oceanogr 40:143–184
Patarnello T, Filipa AM, Volckaert MJ, Castilho R (2007) Pillars of Hercules: is the Atlantic–Mediterranean transition a phylogeographical break? Mol Ecol 16:4426–4444
Pérès JM (1967) The Mediterranean benthos. Oceanogr Mar Biol Annu Rev 5:449–553
Petit RJ, Duminil J, Fineschi S, Hampe A, Salvini D, Vendramin GG (2005) Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14:689–701
Por FD (2009) Tethys returns to the Mediterranean: success and limits of tropical re-colonization. In: Krupp F, Musselman JL, Kotb MMA, Weidig I (eds) Environment, biodiversity and conservation in the Middle East. Proceedings of the first middle eastern biodiversity congress, Aqaba, Jordan, 20–23 October 2008. BioRisk 3:5–19. doi:10.3897/biorisk.3.30
Posada D, Crandall KA (1998) Model test testing the model of DNA substitution. Bioinformatics 14:817–818
Quijada A, Liston A, Robinson W, Alvarez-Buylla E (1997) The ribosomal ITS region as a marker to detect hybridization in pines. Mol Ecol 6:995–996
Sauvage T, Payri CE, Draisma SGA, Prud’homme van Reine WF, Verbruggen H, Belton GS, Gurgel CFD, Gabriel D, Sherwood AR, Fredericq S (2013) Molecular diversity of the Caulerpa racemosa–Caulerpa peltata complex (Caulerpaceae, Bryopsidales) in New Caledonia, with new Australasian records for C. racemosa var. cylindracea. Phycologia 52:6–13
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474
Schaffelke B, Murphy N, Uthicke S (2002) Using genetic techniques to investigate the sources of the invasive alga Caulerpa taxifolia in three new locations in Australia. Mar Pollut Bull 44:204–221
Stam WF, Olsen JL, Zaleski SF, Murray SN, Brown KR, Walters LW (2006) A forensic and phylogenetic survey of Caulerpa species (Caulerpales, chlorophyta) from the Florida Coast, local aquarium shops, and E-commerce: establishing a proactive baseline for early detection. J Phycol 42:113–1124
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tintore J, Violette PE, Blade I, Cruzado A (1988) A study of an intense density front in the eastern Alboran sea: the Almeria–Oran front. J Phys Oceanogr 18:1384–1397
Varela-Álvarez E, Glenn TC, Serrão EA, Duarte CM, Martínez-Daranas B, Valero M, Marba N (2011) Dinucleotide microsatellite markers in the genus Caulerpa. J Appl Phycol 23:715–719
Varela-Álvarez E, Gómez Garreta A, Rull Lluch J, Salvador Soler N, Serrao EA, Ribera Siguan MA (2012) Mediterranean species of Caulerpa are polyploid with smaller genomes in the invasive ones. PLoS ONE 7(10):e47728
Verlaque M, Durand C, Huisman JM, Boudouresque CF, Le Parco Y (2003) On the identity and origin of the Mediterranean invasive Caulerpa racemosa (Caulerpales, Chlorophyta). Eur J Phycol 38:325–339
Voisin M, Engel CR, Viard F (2005) Differential shuffling of native genetic diversity across introduced regions in a brown alga: aquaculture versus maritime traffic effects. Proc Natl Acad Sci USA 102:5432–5437
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR-protocols a guide to methods and applications. Academic press, Orlando, Florida, pp 315–322
Zuljevic A, Antolic B, Nikolic V, Despalatovic M, Cvitkovic I (2012) Absence of successful sexual reproduction of Caulerpa racemosa var. cylindracea in the Adriatic Sea. Phycologia 51:283–286
Acknowledgments
We thank all the people and Institutions that collected plants and/or helped with the sampling collection: Liam Cronin, Ligia Collado-Vides, Gísela Dionisio, Elvira Álvarez, Rocío Santiago, Elena Díaz-Almela, Ricardo Bermejo, Fernando Ojeda Copete, Alexandra Cunha, Pablo Manent, Estibaliz Berecibar, Antonia Ribera-Siguan, Carolina Peña-Martín, Pablo Sanchez Jerez, Felio Lozano, Jacinto Barquín and Francisco Valdés; Natural Park of s’Albufera des Grau (Government of the Balearic Islands: http://www.balearsnatura.com/parc-natural-de-s-albufera-des-grau/equipamiento.html) and the Marine Reserve of Tabarca (Alicante). We also thank for the sequencing work Marta Valente and Cristina Paulino. Financial support for this study was provided by several research projects: CAULERPA_GENETICS (PTDC/MAR/70921/2006) by FCT (Portuguese Science Foundation) co-funded by FEDER and a postdoctoral fellowship (SFRH/BPD/17206/2004) from FCT, Portugal, co-funded by FSE, both to EVA, EXCL/AAG-GLO/0661/2012 to ES, CAULEXPAN (Spanish Ministry of Education, REN2002-00701) to NM, MedVeg (EU contract no. Q5RS-2001-02456), PRADERAS by the Foundation BBVA (Banco Bilbao Vizcaya Argentaria) to CD.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Reusch.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Varela-Álvarez, E., Balau, A.C., Marbà, N. et al. Genetic diversity and biogeographical patterns of Caulerpa prolifera across the Mediterranean and Mediterranean/Atlantic transition zone. Mar Biol 162, 557–569 (2015). https://doi.org/10.1007/s00227-014-2605-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-014-2605-5